Light Sheet Software Guide
User Manual:
Open the PDF directly: View PDF ![]() .
.
Page Count: 10

Light Sheet Software Guide
by Dawnis Chow, Xinyue Cui, and Eileen Troconis
Updated 09/26/2018
Introduction
This is a guide to the software written by Dawnis for use in light sheet data acquisition and
processing. This document explains both the procedures involved in using the programs and
details of what each program does and how it works. Comments are welcome.
The light sheet pipeline can be broken down into several stages. There are a number of
software programs (primarily in Matlab) that have been developed for each step.
Stages of the Experiment and Analysis Pipeline
I.) Running the Experiment
II.) Conversion of stacks to fluorescence time series
* Compression
* Registration
* Segmentation
III.) Validation
IV.) Analytic Approaches
Software Description.
I. Running the Experiment
Programs to allow stimulation and physiological recording during light sheet experiments was
created using Labview. The software was designed to run using the Behavior Laptop connected
to the NI USB-6221 board with appropriate inputs, depending on desired experimental
configuration.
For piezo-electric stimulation experiments, I recommend using the
LSExpt_Variable_Timing_Pulse_GO+RAND LabView file. (It is located under Dropbox/Labview/
LightSheet). This program relies on the generation of a stimulus amplitude and timing text file
from a Matlab script (GenerateRandomStimulusFile.m). The subsequent stimulus file is saved
under Dropbox\Light Sheet\StimulusFile\GoRandomStimulus.txt. Previous stimulus files that
have the same name are moved to a new file that have the creation date of that file appended to
prevent the loss of information of previously used stimulation patterns.

Another useful LabView program is the BuzzPiezoelectridDevice program. This allows testing of
the piezo stimulation device to see if everything is wired correctly before the experiment.
Experiment Procedure
To perform the experiment, in the LSExpt Lab View program, the user must specify the frames
per stack and the amount of time to acquire signal data (for physiological experiments). The
program automatically reads the stimulus file called GoRandomStimulus.txt. The LabView
program is started before acquisition is started on the light sheet computer. Once acquisition
starts on the computer, the current frame count should increment if things are working correctly.
II. Conversion of Stacks to Useable Data
1. Data Compression and Registration
After the collection of data with the light sheet microscope, image stacks are written to a
folder containing the ‘.stack’ files along with meta-data including ‘ch0.xml’,
‘stack_dimensions.txt’, and ‘stack_frequency.txt’. The primary goal of this stage of
analysis is to convert the .stack files into either the more compressed ‘.klb’ format (10x
smaller) or ‘.tif’ format. At the same time, the conversion software allows cropping of
regions that do not contain data, and also simultaneously performs a rough 2D (in X and
Y) rigid registration of stack volumes over time.
The getConversionParamteres.m helper script in Matlab saves user-defined parameters
into each folder to allow batch-conversion of multiple experiments at once. Once all user
settings have defined per experiment, specify the appropriate folders under
list_of_directories in the script Batch_parLightSheet_Convert_Stacks.m and run.
Note: There the possibility of performing a mutual-information based registration which
takes much longer. Anecdotally, I’ve seen that this can reduce the amount of jitter in time
series over just using the 2D method. Whether or not to use this should be decided a
priori for each paper’s worth of experiments.
Procedure:
On the light sheet computer do the following:
2.
-Open MATLAB
-Open script: getConversionParameters.m, set the user parameters as:
oOutputformat = ‘.klb’
oregisterConcurrently = true
oCrop_and_mask_image = true
ocomputeTimeSeriesMaxProjection = true
ocomputeTimeSeriesMeanProjection = true

-Run
-A new window will open. Find and select the folder that contains the stacks
-A new window called StackCropperGUI will open
oFit the fish inside the red square by using the arrow keys and the commands
written on the window. The amount that the square moves with each click can
be set in the %size scaling box.
oMove the bar under the histogram to reduce the black background as much
as possible. Check the ‘show masking’ box to preview.
oLoad all stacks and make sure they all look ok.
oAccept
-A new window will open. Select the same folder as before to save all output.
-A new window called ‘Stack2DRegistrationGUI’ will open.
oChoose a slice that contains components of varying brightness. The program
will use this slice to compare across stacks by assigning green and red spots
to the corresponding slices of the two stacks being compared at a time. If the
slices from the two stacks are perfectly aligned, the overlay image will appear
black and white (no green or red spots).
oIn Flattening Method, select ‘Mean of selected frames’
oChoose an appropriate slice in the box under Flattening Method
oBy clicking on arrow keys a yellow square will appear on the slice. Move it to
an area that contains components of varying brightness. The dimensions of
the square may be modified, but enlarging it increases processing time.
oFixed stack # should be the middle stack. Compare a few stacks to middle
stack by typing in Stack # box.
oPreview Registration
oIf everything looks acceptable, click Register!
oRepeat this process for each desired experimental folder.
-Put the appropriate file paths in Batch_parLightSheet_Convert_Stacks as specified.
-Files are generated in the previously selected folder. Copy these into a drive and
take them to the IMARIS computer: tS_maxproj and tS_meanProj (in timeProj
folder), Background_0, ch0, conversionLog, Stack dimensions, Stack_frequency.
- Note: the script ‘timeseries_viewer’ allows visualization of slices across time for
closer visual inspection
*Possibly consider re-moving stacks where there are significant movement artifacts
before computing max projections.

2. Cell Segmentation in Imaris
The goal of this part of the pipeline is to generate spots in Imaris for each nuclear-
labelled cell in the time series. To do this, I would recommend only moving files that are
absolutely necessary between computers. This includes the max/mean projection from
the previous step (.tif files) and the meta-data files (ch0.xml, stack_frequency.txt, and
stack_dimensions.txt).
The segmentation is done in three steps. In the first step, software from the Ahrens lab
(from Kawashima et al., 2016) finds candidate nuclei in each plane. These candidates
are communicated to Imaris to generate spots. In the second step, filtering operations
are applied in Imaris to keep user-specified spots. Lastly, the final segmentation is
exported into a file format that can be read by software on the light sheet computer to
extract fluorescence time series for each cell.
Note: The program currently checks within each slice for overlapping neurons, but
doesn’t check for this in adjacent slices at the moment.
Procedure:
-Move time projection files and meta-data to the Imaris computer
-Open IMARIS (make sure that only one window is open using Task Monitor)
-In IMARIS, open tS_meanProj (or maxProj, whichever one you think best reflects
your cell distribution)
-Make sure that on IMARIS, go into 3D view and also: Edit -> Image Properties ->
Geometry(or use ctrl+I) -> Voxel size -> x = 0.41, y = 0.41, z = 5
-Open the MATLAB script ‘segment_HuCH2BGCamP_ims.m’(File location(at
01Jun2018): C:\Users\Joe\Dropbox\CellAnalysisUtilities\LightSheet\segmentation)
oAdjust brightness threshold(br_threshold), contrast threshold (cont_threshold)
and cell radius (cell_rad) for cell detection. Usually I use br_threshold=100,
cont_threshold=7, cellrad=3. Please adjust according to your needs.
oRun
oPlease enter segmentation channel: 1
oIf the program starts counting numbers -> good sign! Each number is a slice
where the program is finding cells
-In IMARIS
oSelect ‘autosegment’
oFilter icon -> Add, In Filter Type , select in popmenu:
▪Intensity Mean Ch=1 -> 500 (recommended by Dawnis(Cui
sometimes use 100 to get largest coverage of neurons))
▪Number of voxels -> 50 to 80 (recommended by Dawnis)
▪Position Z -> get rid of most superficial layer
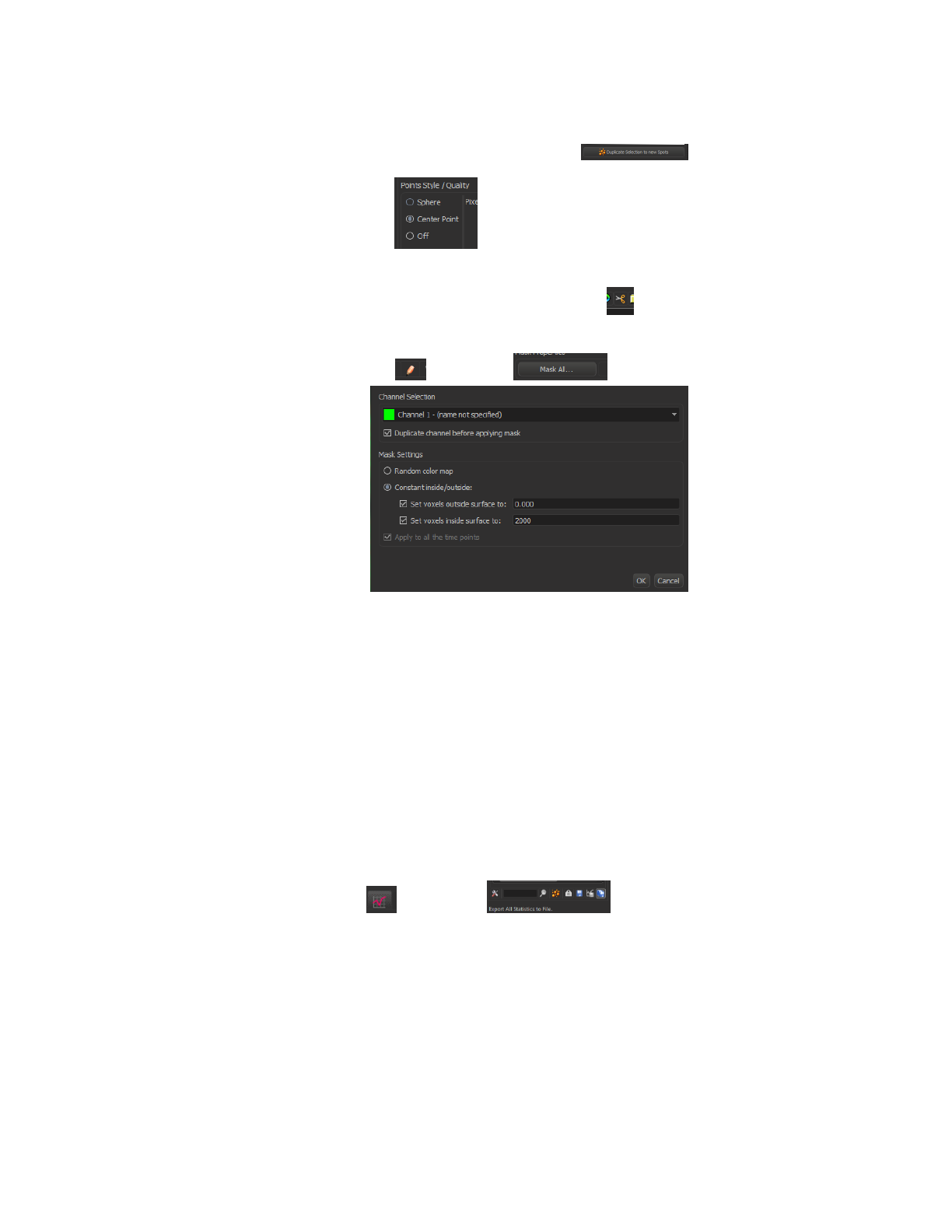
oDuplicate selection to new Spots using this (at bottom left)
oUse ‘Center Point’
oCheck the cell detection quality: (Cui comment: Usually I will just add two
clipping plane to get a slice of brain (using this ) and check the distribution
of center points in that slice, but there are other ways to do that…)
▪Pencil icon -> Mask all -> Set outside to 0 and inside
to 2000
▪Edit -> Image Properties -> make Channel 1 (max projection data)
green and Channel 2 (cells you just masked) magenta, check if they
match. If not, you can discard current segment spots, and adjust
brightness threshold (br_threshold), contrast threshold
(cont_threshold) and cell radius (cell_rad) in the
‘segment_HuCH2BGCamP_ims.m’, run that program again until
satisfied. Or adjust your Imaris fluo mean threshold or etc.
oFile ->Export(or directly press ctrl+E) -> save .ims file in Imaris K drive
-Then we have two ways to extract cell position info:
oWay1: Click on the ‘autoSegmentation Selection[“Position…..’ spots, click
Statistics icon ( ), then click (the rightest one) export All
Statistics to file. Then open the saved statistics xls file, use save as… to
convert it to xlsx. file. Copy that xlsx to your portable hard drive.
oWay2: In Matlab, first make sure you’re in the right directory which you want
to put the position .mat file in, then open script ‘imSegmentation2matfile’ (File
location(at 01Jun2018): C:
\Users\Joe\Dropbox\CellAnalysisUtilities\LightSheet)
▪backTrackData = false
▪Run

▪Please type the name of the spots objection to use: (Note: type in the
name of the spots object you want to extract position from, which
usually should the name of be this one. Press F2
and then Cltr+C to copy the name of this spots object and past it here
to tell the matlab program)
▪Creates file called ‘imsSegmentationData.mat’ in whatever directory
MATLAB is. Move file along with ‘autoSegmentData.mat’ file to drive.
3. Obtaining Fluorescence Time Series (FTS)
In this step, the cell segmentation information is transferred back to the light sheet
computer to extract a fluorescence time series for each segmented neuron.
Procedure: On the light sheet computer
-In MATLAB
oScript: ‘extract_lightsheet_time_series_to_file’
▪useExcelFile = true (if you used Way1, which uses excel file)
useExcelFile = false (if you used Way2, which didn’t use excel)
▪Run
▪A new window will open. Select ‘processed’ folder
▪Enter 1 to select ‘.klb’
▪Enter
▪A new window will open. Select ‘imsSegmentationData’ in drive
•Note: Or, if useExcelFile=true, you’ll need to choose ch0.xml
file and the .xlsx file (the one just generated from Imaris) in this
step.
▪This takes some time…
▪The saved fluorescence time series file will appear in the .klb
directory. This is the file needed to perform analysis on data.
▪Note: voxels are only considered a part of the volume if the center of
the voxel is within the ellipse defined by the spot.
IV. Validation Approaches: Older Approaches (independent of data_explorer_gui)
The main visually-based validation approaches are to use timeseriesviewerGUI or to
draw ROIs onto slice movies using FIJI. Both methods need to use timeseriesviwerGUI
first to generate the time slice movies.
On the light sheet computer:
•Generate of tsView files:
•In MATLAB run ‘timeseries_viewer_GUI.m’ (Should be at
Custom_Matlab_Scripts\timeseries_viewer_GUI), add to path if asked.

•Click ‘Load’, check if ‘.klb’ or ‘.tiff’ is selected, load the corresponding file folder
that contains .klb or .tiff data(usually should be the ‘processed’ folder).
•Click Generate All Slice Movies, wait. This could take ~10s/timeframe. After the
program is done, we’ll have our time series of each plane in a newly created
folder called ‘tsView’.
•You can transfer any/all of the planes you want to check onto external storage.
Drawing ROIs in timeseries movies
The information for spot positions/diameters is in the fluorescence_time_series.mat file
included in the variables spas and spRaddiiXYZ. You also need the relevant tsView (.tif)
files. This procedure generates ROIs in FIJI for each specified neuron passed into the
command line argument of genFIJI_ROIs (a MATLAB function).
Procedure: On the Imaris computer
•Code: genFIJI_ROIs, drawRois.py. Copy these in same folder as everything else that
you need.
•In MATLAB, load your fluroescence_time_series data extracted from raw image. (use
load(‘filepath/filename’);)
•In MATLAB, run: genFIJI_ROIs(spPos, [1:size(spPos,1)]', spRadiiXYZ)
oSubstitute highlighted with indices of cells of interest. This will be the index of the
cells within both spPos and spRadiiXYZ. Best way to do this is to create spPos2
and spRadiiXYZ2 with cells of interest and then make index be 1:9 or wtv, but the
entire set.
•This generates a folder called spotROIs
•Open ImageJ
•Open movie for the slice of interest, which should be a .klb file located in processed/
tsView folder
•Open 'drawRois.py' by dragging that .py file onto FIJI menu.
•Run
•Choose the position file (in folder spotROIs) of your plane of interest
•In ROI manager, make sure that it's empty initially and tick show all
•Saves ROI data as a zip
•Make sure ROI manager is clear before you load the ROIs for another plane.
data_exploration_gui
This GUI allows visualization of the spatial and time series information associated with
extracted fluorescence time series, and provides a number of options manipulating sets
of cells and interfacing with Imaris. Additionally, it allows simultaneous visualization
of calcium time series and slice movies for validation.
On installed systems, this GUI will be located in the folder data_explorer_gui (for
example, on Imaris 2 it is located in ~\Dropbox\data_explorer_gui\GUI) and is also on
the path. The current version of the GUI is data_explorer_gui.m and its associated .fig
file. If the path in MATLAB is correctly configured, typically the guy can be run simply by
typing data_explorer_gui in the command window.

A Brief Tour
Navigation Window
Here, spatial data associated with your data set appears. The left window plots the
whole dataset in 3D, while the right window plots coronal slices, highlighting neurons of
interest in your selected ROI. Plot type has three modes: All, ROI, and Validation. The
default mode, ‘all’, allows general exploration of the data via coronal slices. Neurons in
this slice window are plowable via mouse or slider selection. In ROI mode, the
background neurons may be displayed somewhat more sparsely, and all highlighted
ROI groups will be plotted (selected under All ROIs listbox). In validation mode, the user
can explore time slice movies by loading slice movies (see button under Tools) and
loading a .klb Z-Stack.
Time Series Window
In general, calcium time series appear in the top window here. The second window is
currently unused, but can be adopted for displaying averaged results or stimuli. The plot
types changes whether raw or delta f/f is displayed. (Delta f/f must be calculated using
the Compute dFF button if it is not included in your fluorescence time series .mat file). If
multiple members of an ROI are selected, they can be plotted simultaneously using the
PlotFTS button.
ROI Window
Here, groups of neurons can be managed. The default group added upon load contains
all neurons in the data set. A new group containing no neurons can be created by
Navigation Panel

pressing the ‘a’ key with the ROI Master (left listbox) selected. ROIs can also be
imported from analysis code or from Imaris (see Imaris menu).
ROIs can be imported or exported to file using the Export or Load button, or to FIJI
ROIs using the Gen. Fiji ROIs button.
Tools Window
A variety of custom tools appear here.
*Generate Time Slice Movies: Generates tsView*.tif files for viewing time slice movies
from a .klb stack directory.
*Load Slice Movie: loads one of the tsView files for viewing in validation mode.
*Compute dFF: Uses a dFF converter written by Dawnis that provides a number of
options for filtering and windowing.
Correlation Window
Cui will explain this part.
Imaris Menu
This allows interface with a data set in Imaris. Use the connect menu item to connect to
an Imaris instance. Imaris visibility toggles the visibility of the Imaris window and is
useful to know whether or not the connection was successful. The main function of
connecting to Imaris is to import and export sets of spots. These must match positions
in the data set precisely to be recognized as valid. (Meaning that the positions must be
on a proper scale of 0.41 microns per pixel in X and Y and 5 microns per Z).
Loading Data and Manipulating Plots
Upon opening the GUI, the first task is to load a fluorescence_time_series.mat file into
memory. This is accomplished by going to the DataSet menu item and selecting Load
Dataset. If all goes well, neuronal locations will be marked with points colored by
maximum intensity in the Navigation window. The slider can be moved to change the
coronal slice in the slice window. The previous two windows are plotted in 3D, and the
view can be changed using the rotation tool in the toolbar underneath the menu.
Toolbar: hand (moves figure), data_tip (can select data points and display
information), rotation for 3D figures.
In general, the toolbar buttons can be used on any axis with a plot.
Manipulating ROIs
Create a new ROI by pressing a in the ROI Master listbox (left). Neurons can be added
manually by updating the circled neuron in the Navigation slice window, clicking on the
ROI members box, and pressing ‘a’. In addition, valid array expressions can be passed
in through the textbox, (for example [5:5:1000] will select every fifth neuron between 5
and 1000) in the input box in the ROI window (highlighted yellow).

ROI Master Keyboard Commands
‘a’: Create an empty ROI set.
‘delete’: delete highlighted ROI set.
’n’: rename highlighted ROI set.
ROIListbox Keyboard Commands
‘a’: add currently selected cell to ROI
‘delete’: delete currently selected cell(s).
‘k’: keep currently selected cell(s) and delete
everything else.
‘p’/‘q’: plot currently selected members.
‘f’: find the selected ROI member(s) in space
and plot
Validation Mode
To use the validation mode view, first load a
slice movie (and remember which one it is!)
using the button under Tools. The Matlab command window will let you know when it is
done loading (typically 30 to 90 seconds). Then, switch to Validation mode under Plot
Type. Use the Load Z Stack button to visualize neurons in each slice. Any neurons that
are selected under ROI members will be circled in the left Navigation window. Select the
appropriate slice in your data set using the slider. Then, hit throw ROIs to highlight the
appropriate members of each ROI for visualization on the movie. (Note, this will note
update until the slider is moved).
Note that the color axis scaling of the top windows can be controlled by inputing
appropriate min. and max. under the color axis prompt.
Correlation Analysis
To be added by Cui
Possible Future Features and Work
-Re-factor all correlation and cluster analysis to Tool panel button(s)
-Export current z-stack to new volume in Imaris
-Event triggered analysis (button in Tool panel)
-Select all ROI members in coronal slice