Manual
User Manual:
Open the PDF directly: View PDF ![]() .
.
Page Count: 20
- Introduction
- How to use SomaticSeq.Wrapper.sh
- The step-by-step SomaticSeq Workflow
- To run the dockerized somatic mutation callers
- Use BAMSurgeon to create training data
- Release Notes
- Version 1.0
- Version 1.1
- Version 1.2
- Version 2.0
- Version 2.0.2
- Version 2.1.2
- Version 2.2
- Version 2.2.1
- Version 2.2.2
- Version 2.2.3
- Version 2.2.4
- Version 2.2.5
- Version 2.3.0
- Version 2.3.1
- Version 2.3.2
- Version 2.4.0
- Version 2.4.1
- Version 2.5.0
- Version 2.5.1
- Version 2.5.2
- Version 2.6.0
- Version 2.6.1
- Version 2.7.0
- Version 2.7.1
- Version 2.7.2
- Version 2.8.0
- Contact Us
SomaticSeq Documentation
Li Tai Fang / li_tai.fang@roche.com
July 8, 2018
1 Introduction
SomaticSeq is a exible post-somatic-mutation-calling algorithm for improved accuracy. We have incorpo-
rated 10+ somatic mutation caller(s). Any combinatin of them can be used to obtain a combined call set,
and then SomaticSeq uses machine learning (adaptive boosting) to distinguish true mutations from false
positives from that call set. The mutation callers we have incorporated are MuTect/Indelocator/MuTect2,
VarScan2, JointSNVMix, SomaticSniper, VarDict, MuSE, LoFreq, Scalpel, Strelka, and TNscope. You may
incorporate some or all of those callers into your own pipeline with SomaticSeq.
The manuscript, An ensemble approach to accurately detect somatic mutations using Somat-
icSeq, was published in Genome Biology 2015, 16:197. The SomaticSeq project is located at
https://github.com/bioinform/somaticseq. There have been some major improvements in SomaticSeq since
that Genome Biology publication in 2015.
SomaticSeq.Wrapper.sh is a bash script that calls a series of algorithms to combine the output of the
somatic mutation caller(s). Then, depending on the input les and R scripts that are fed to Somatic-
Seq.Wrapper.sh, it will either 1) train the call set into a classier, 2) predict high-condence somatic muta-
tions from the call set based on a pre-dened classier, or 3) simply label the calls (i.e., PASS, LowQual, or
REJECT) based on majority vote of the tools.
1.1 Dependencies
• Python 3, plus regex, pysam, numpy, and scipy libraries. All the .py scripts are written in Python 3.
• R, plus the ada package in R.
• BEDTools (if inclusion and/or an exclusion region les are supplied)
• Optional: dbSNP in VCF format (if you want to use dbSNP membership as a part of the training).
• At least one of MuTect/Indelocator/MuTect2, VarScan2, JointSNVMix2, SomaticSniper, VarDict,
MuSE, LoFreq, Scalpel, Strelka2 and/or TNscope. Those are the tools we have incorporated in So-
maticSeq. If there are other somatic tools that may be good addition to our list, please make the
suggestion to us.
1.2 Docker images
SomaticSeq and most somatic mutation callers we have incorporated are dockerized.
• SomaticSeq: https://hub.docker.com/r/lethalfang/somaticseq
• MuTect2: https://hub.docker.com/r/broadinstitute/gatk
• VarScan2: https://hub.docker.com/r/djordjeklisic/sbg-varscan2
• JointSNVMix2: https://hub.docker.com/r/lethalfang/jointsnvmix2
• SomaticSniper: https://hub.docker.com/r/lethalfang/somaticsniper
1

• VarDict: https://hub.docker.com/r/lethalfang/vardictjava
• MuSE: https://hub.docker.com/r/marghoob/muse
• LoFreq: https://hub.docker.com/r/marghoob/lofreq
• Scalpel: https://hub.docker.com/r/lethalfang/scalpel
• Strelka2: https://hub.docker.com/r/lethalfang/strelka
2 How to use SomaticSeq.Wrapper.sh
The SomaticSeq.Wrapper.sh is a wrapper script that calls a series of programs and procedures after you
have run your individual somatic mutation callers. Section 4 will teach you how to run those mutation callers
that have been dockerized. It also includes ways to create semi-simulated training data that can be used to
create SomaticSeq classiers. In the next section, we will describe the workow in this wrapper script in
detail, so you may not be dependent on this wrapper script. You can either modify this wrapper script or
create your own workow using whatever workow language you want.
2.1 SomaticSeq Training Mode
To create SomaticSeq classiers, you need VCF les containing true positive SNVs and INDELs. There is
also an option to include a list of regions to include and/or exclude from this exercise. The exclusion or
inclusion regions can be VCF or BED les. An inclusion region may be subset of the call sets where you have
validated their true/false mutation status, so that only those regions will be used for training. An exclusion
region can be regions where the “truth” is ambigious. All the variants in the truth VCF les are assumed
to be true positives. Every mutation call not in the truth VCF les is assumed to be false positives (as long
as the genomic coordiante is in inclusion region and not in exclusion region if those regions are provided).
All the output VCF les from individual callers are optional. Those VCF les can be bgzipped if they
have .vcf.gz extensions. It is imperative that you will use the same parameter for prediction as you do for
training.
1# An example : f o r tr a in i ng , tr ut h f i l e and t he c o r r e c t R s c r i p t ar e r eq u i re d .
3SomaticSeq . Wrapper . sh \
−−mutect2 MuTect2/ v ar ia nt s . v cf \
5−−varscan−snv VarScan2/ var ia nt s . snp . v cf \
−−varscan−i n d e l VarScan2/ v a r i a n ts . i n d e l . v c f \
7−−jsm JointSNVMix2/ v ar ian ts . snp . vcf \
−−s ni pe r SomaticSniper/ v ar ia nt s . snp . vc f \
9−−v ar di ct VarDict/ v ar ia nts . v cf \
−−muse MuSE/ v ar ia nt s . snp . vc f \
11 −−lofreq−snv LoFreq/ v ar ia nt s . snp . vc f \
−−lofreq−i n d e l LoFreq/ v a r i a n ts . i n d e l . v c f \
13 −− s c a l p e l S ca lp e l / v ar ia nt s . i n d e l . v cf \
−−strelka−snv St re lk a / va ria nt s . snv . v cf \
15 −−strelka−i n d e l S t re l k a / v a r i a nt s . i n d e l . vc f \
−−tnscope TNscope . v cf . gz \
17 −−normal−bam matched_normal .bam \
−−tumor−bam tumor .bam \
19 −−ada−r−s c r i p t $somaticseq / r _s cr ip ts /ada_model_builder_ntChange .R \
−−genome−r e f e r e n c e GRCh38 . f a \
21 −−cosmic cosmic .GRCh38. v cf \
−−dbsnp dbSNP.GRCh38. vcf \
23 −−exclusion−regio n bl ac kL is t . bed \
−−inclusion−region highConfidenceRegions .bed
25 −−truth−snv t r u e P o s i t i v e s . snv . v c f \
−−truth−i n d e l t r u e P o s i t i v e s . i n d e l . v c f \
27 −−output−d i r $OUTPUT_DIR
2

SomaticSeq.Wrapper.sh supports any combination of the somatic mutation callers we have incorporated
into the workow. SomaticSeq will run based on the output VCFs you have provided. It will train for SNV
and/or INDEL if you provide the truePositives.snv.vcf and/or truePositives.indel.vcf le(s) as well as the
proper R script (ada_model_builder_ntChange.R). Otherwise, it will fall back to the simple caller consensus
mode.
2.2 SomaticSeq Prediction Mode
Make sure the classiers (.RData les) and the proper R script (ada_model_predictor.R) are supplied,
Without either of them, it will fall back to the simple caller consensus mode.
1# The ∗. RData f i l e s a re t r ai n e d c l a s s i f i e r from the t r a i n i n g mode .
SomaticSeq . Wrapper . sh \
3−−mutect2 MuTect2/ v ar ia nt s . v cf \
−−varscan−snv VarScan2/ v ar ia nt s . snp . vc f \
5−−varscan−i n d e l VarScan2/ v a r i a n ts . i n d e l . v c f \
−−jsm JointSNVMix2/ v ar ian ts . snp . vcf \
7−−s ni pe r SomaticSniper/ v ar ia nt s . snp . vc f \
−−v ar di ct VarDict/ v ar ia nts . v cf \
9−−muse MuSE/ v ar ia nt s . snp . vc f \
−−lofreq−snv LoFreq/ v ar ia nt s . snp . vc f \
11 −−lofreq−i n d e l LoFreq/ v a r i a n ts . i n d e l . v c f \
−− s c a l p e l S ca lp e l / v ar ia nt s . i n d e l . v cf \
13 −−strelka−snv St re lk a / va ria nt s . snv . v cf \
−−strelka−i n d e l S t re l k a / v a r i a nt s . i n d e l . vc f \
15 −−tnscope TNscope . v cf . gz \
−−normal−bam matched_normal .bam \
17 −−tumor−bam tumor .bam \
−−ada−r−s c r i p t ada_model_predictor .R \
19 −−genome−r e f e r e n c e human_b37 . f a s t a \
−−genome−r e f e r e n c e GRCh38 . f a \
21 −−cosmic cosmic .GRCh38. v cf \
−−dbsnp dbSNP.GRCh38. vcf \
23 −−exclusion−regio n bl ac kL is t . bed \
−−inclusion−region highConfidenceRegions .bed
25 −− classifier−snv sSNV . C l a s s i f i e r . RData \
−− classifier−in de l sINDEL. C l a s s i f i e r . RData \
27 −−pass−thres hol d 0. 5 \
−−lowqual−th res hold 0.1 \
29 −−output−d i r $OUTPUT_DIR
2.3 Consensus Mode
Same as the commands previously, but not including the R script or the ground truth les. Without those
information, SomaticSeq will forgo machine learning, and fall back into a simple majority vote.
3 The step-by-step SomaticSeq Workow
We’ll describe the workow here, so you may modify the workow and/or create your own workow (better
optimized for your own usage), instead of using SomaticSeq.Wrapper.sh we have included in the repo.
3.1 Combine the call sets
We use utilities/getUniqueVcfPositions.py and vcfsorter.pl to combine the VCF les from dierent callers.
For each caller output, intermediate VCF le(s) were modied separate the SNVs and INDELs calls, and
also remove some REJECT calls to reduce le sizes.
3

1. Modify (original) MuTect and/or Indelocator output VCF les. Since MuTect’s output VCF do not
always put the tumor and normal samples in the same columns, the script will determine that infor-
mation based on either the BAM les (the header has sample name information), or based on the
sample information that you tell it, and then determine which column belongs to the normal, and
which column belongs to the tumor.
# Modify MuTect and In del oca tor ’ s output VCF based on BAM f i l e s
2modify_MuTect . py −i n f i l e input . v cf −o u t f i l e output . v cf −nbam normal .bam −tbam tumor .bam
4# Based on the sample name you supply :
modify_MuTect . py −i n f i l e input . v cf −o u t f i l e output . v cf −nsm NormalSampleName −tsm
TumorSampleName
2. For MuTect2, this script will split multi-allelic records into one variant per line in the VCF le. This
is to make thing easier for the SSeq_merged.vcf2tsv.py script later.
1# Based on the sample name you supply :
modify_MuTect2 . py −i n f i l e MuTect2 . F i l t e r e d . v cf −snv mutect . snp . v cf −i n d e l mutect . i n d e l . v c f
3. Modify VarScan’s output VCF les to be rigorously concordant to VCF format standard, and to attach
the tag ’VarScan2’ to somatic calls.
# Do i t f o r both the SNV and i n d e l
2modify_VJSD . py −method VarScan2 −i n f i l e input . v cf −o u t f i l e output . vcf
4. JointSNVMix2 does not output VCF les. In our own workow, we convert its out-
put into a basic VCF le with an 2 awk one-liners, which you may see at utilities/dock-
ered_pipelines/mutation_callers/submit_JointSNVMix2.sh.
# To avoid t ext f i l e s on the or der o f ter abyt es , t h i s awk one−l i n e r keeps e n t r i e s where th e
r ef e re n c e i s not ”N” , and the somatic p r o b a b i l i t i e s ar e at l e a s t 0 . 9 5.
2awk −F” \ t ” ’NR!=1 && $4!=”N” && $10+$11>=0.95’
4# This awk one−l i n e r c onve rts the text f i l e i nt o a b as ic VCF f i l e
awk −F” \ t ” ’{ p ri nt $1 ”\ t ” $2 ”\ t . \ t ” $3 ”\ t ” $4 ”\ t . \ t . \tAAAB=” $10 ” ;AABB=” $11 ”\tRD:AD\
t ” $5 ” : ” $6 ” \ t ” $7 ” : ” $8 } ’
6
## The a ct ua l commands we ’ ve used i n our workflow :
8echo −e ’##f i l e f o r m a t=VCFv4. 1 ’ > u nsorted . vc f
echo −e ’##INFO=<ID=AAAB, Number=1,Type=Float , D es cr ip ti o n=”P r ob a bi l i ty o f J oi nt Genotype AA
in Normal and AB in Tumor”>’ >> unsorted . vc f
10 echo −e ’##INFO=<ID=AABB, Number=1,Type=Float , D es c ri p ti on=”P r o ba bi l it y o f J o in t Genotype AA
in Normal and BB in Tumor”>’ >> unsorted . v cf
echo −e ’##FORMAT=<ID=RD, Number=1,Type=I nt eg er , D e sc r ip t io n=”Depth of r e f er e nc e −supporting
bases ( reads1 ) ”>’ >> unsorted . v cf
12 echo −e ’##FORMAT=<ID=AD, Number=1,Type=In t eg er , De sc ri p ti on=”Depth o f v ari an t−supporting
bases ( reads2 ) ”>’ >> unsorted . v cf
echo −e ’#CHROM\tPOS\tID\tREF\tALT\tQUAL\tFILTER\tINFO\tFORMAT\tNORMAL\tTUMOR’ >> unsorted .
vcf
14
python $PATH/TO/jsm . py c l a s s i f y joint_snv_mix_two genome .GRCh37. fa normal .bam tumor . bam
tr ai ned . parameter . cf g /dev/ stdout | \
16 awk −F” \ t ” ’NR!=1 && $4!=”N” && $10+$11 >=0.95 ’ | \
awk −F” \ t ” ’{ p ri nt $1 ”\ t ” $2 ”\ t . \ t ” $3 ”\ t ” $4 ”\ t . \ t . \tAAAB=” $10 ” ;AABB=” $11 ”\tRD:AD\
t ” $5 ” : ” $6 ” \ t ” $7 ” : ” $8 } ’ >> unsor ted . v cf
4
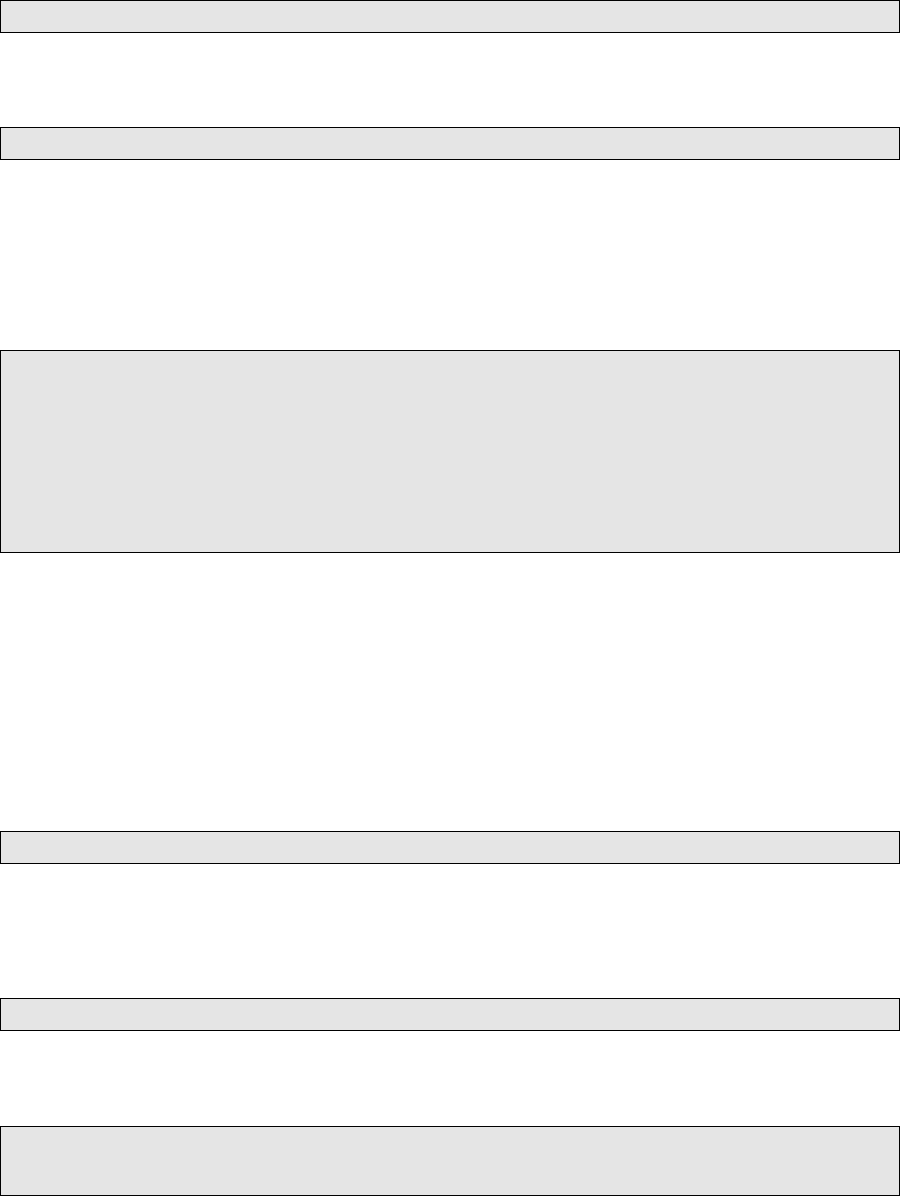
After that, you’ll also want to sort the VCF le.
1modify_VJSD . py −method JointSNVMix2 −i n f i l e input . v cf −o u t f i l e output . v cf
5. Modify SomaticSniper’s output:
1modify_VJSD . py −method SomaticSniper −i n f i l e input . vcf −o u t f i l e output . vcf
6. VarDict has both SNV and indel, plus some other variants in the same VCF le. Our script will
create two les, one for SNV and one for indel, while everything else is ignored for now. By default,
LikelySomatic/StrongSomatic and PASS calls will be labeled VarDict. However, in our SomaticSeq
paper, based on our experience in DREAM Challenge, we implemented two custom lters to relax the
VarDict tagging criteria.
1# Def ault VarDict taggi ng c r i t e r i a , only PASS ( and Li kel y or Strong Somatic ) :
modify_VJSD . py −method VarDict −i n f i l e int pu t . vcf −o u t f i l e output . vcf
3
# When running VarDict , i f var2vcf_paired . pl i s used to gener at e the VCF f i l e , you may rel ax
the ta gging c r i t e r i a with −f i l t e r pa ir ed
5modify_VJSD . py −method VarDict −i n f i l e int pu t . vcf −o u t f i l e output . vcf −f i l t e r paire d
7# When running VarDict , i f var2vcf_somatic . pl i s used to g enerat e the VCF f i l e , you may
rel ax the tagging c r i t e r i a with −f i l t e r somatic
modify_VJSD . py −
method VarDict −i n f i l e intpu t . vcf −o u t f i l e output . vcf −f i l t e r somatic
In the SomaticSeq paper, -lter somatic was used because var2vcf_somatic.pl was used to generate
VarDict’s VCF les. In the SomaticSeq.Wrapper.sh script, however, -lter paired is used because
VarDict authors have since recommended var2vcf_paired.pl script to create the VCF les. While there
are some dierences (dierent stringencies in some lters) in what VarDict labels as PASS between
the somatic.pl and paired.pl scripts, the dierence is miniscule after applying our custom lter (which
relaxes the lter, resulting in a dierence about 5 calls out of 15,000).
The output les will be snp.output.vcf and indel.output.vcf.
7. MuSE was not a part of our analysis in the SomaticSeq paper. We have implemented it later.
modify_VJSD . py −method MuSE −i n f i l e input . v cf −o u t f i l e output . vcf
8. LoFreq and Scalpel do not require modication. LoFreq has no sample columns anyway.
9. Add “GT” eld to sample columns to make it compatible with GATK CombineVariants.
1modify_Strelka .py −i n f i l e somatic . snvs . vc f . gz −o u t f i l e s t ra lka . snv . vc f
10. Finally, with the VCF les modied, you need combine them: one for SNV and one for indel separately.
1# Combine the VCF f i l e s f or SNV. Any or a l l of the VCF f i l e s may be pres ent .
u t i l i t i e s / getUniqueVcfPos itio ns . py −v cf s mutect . vc f varscan . snp . vc f jo intsnvmix . v cf snp .
va rd ic t . vc f muse . vc f −out CombineVariants . snp . vcf
5

3.2 Apply inclusion and exclusion regions
This step may be needed for model training. The workow in SomaticSeq.Wrapper.sh allows for inclusion
and exclusion region. An inclusion region means we will only use calls inside these regions. An exclusion
region means we do not care about calls inside this region. DREAM Challenge had exclusion regions, e.g.,
blacklisted regions, etc.
# In the DREAM_Stage_3 dire ct ory , we have include d an e xc lu si on re gio n BED f i l e as an example
2# This command us es BEDtools to r i d o f a l l c a l l s i n t he e x cl u s i on r e g io n
intersectBed −header −a BINA_somatic . snp . v cf −b i gn ore . bed −v > somatic . snp . pr oces sed . v cf
4intersectBed −header −a BINA_somatic . i n d e l . v c f −b i gnor e . bed −v > soma tic . i n d e l . pr o ce s se d . v c f
6# A l te r n at i v el y ( or both ) , t h i s command us es BEDtools to keep only c a l l s i n the i n c l u s i o n r eg io n
intersectBed −header −a BINA_somatic . snp . v cf −b i n c l u s i o n . bed > som ati c . snp . p ro ce ss ed . v cf
8intersectBed −header −a BINA_somatic . i n d e l . v c f −b i n c l u s i o n . bed > som ati c . i n d e l . p ro ce ss ed . v cf
3.3 Convert the VCF le into TSV le
This script works for all VCF les. It extracts information from BAM les, as well as some individual
callers’ output VCF les. If the ground truth VCF le is included, a called variant will be annotated as a
true positive, and everything will be annotated as a false positive.
1# SNV
SSeq_merged . vcf2ts v . py −r e f genome .GRCh37. fa −myvcf somatic . snp . proc es sed . v cf −truth Ground . trut h
. snp . vc f −mutect MuTect/ va ri an ts . snp . vc f . gz −varscan VarScan2/ var ia nt s . snp . v cf −jsm JSM2/
va ri an ts . vc f −s ni pe r SomaticSniper/ v ar ia nt s . vc f −va rd ic t VarDict/snp . v ar ia nt s . v cf −muse MuSE/
va ri an ts . vc f −l o f r e q LoFreq/ v ar ia nt s . snp . v cf −s t r e l k a St r el k a / v a ri an ts . snp . v c f −dedup −tbam
tumor .bam −
nbam normal .bam −o u t f i l e Ensemble . sSNV. t sv
That was for SNV, and indel is almost the same thing. After version 2.1, we have replaced all information
from SAMtools and HaplotypeCaller with information directly from the BAM les. The accuracy dierences
are negligible with signicant improvement in usability and resource requirement.
# INDEL:
2SSeq_merged . vcf2ts v . py −r e f genome .GRCh37. fa −myvcf som atic . i n d e l . p r oc e ss e d . v c f −truth Ground .
t ru th . i n d e l . v cf −varscan VarScan2/ variants . snp . v cf −v a rd i c t VarDict / i n d e l . v a r i a nt s . v c f −
l o f r e q LoFreq/ v ar ia nt s . i n d el . v cf −s c a l p e l S c al pe l / v ar ia n ts . i n d e l . v cf −s t r e l k a S t re lk a /
v a r ia n t s . i n d e l . v cf −tbam tumor .bam −nbam normal .bam −dedup −o u t f i l e Ensemble . sINDEL . tsv
At the end of this, Ensemble.sSNV.tsv and Ensemble.sINDEL.tsv are created.
All the options for SSeq_merged.vcf2tsv.py are listed here. They can also be displayed by running
SSeq_merged.vcf2tsv.py --help.
2−myvcf Input VCF f i l e of the merged c a l l s [REQUIRED]
−r e f Genome r e f e r e n c e f a / f a s t a f i l e [REQUIRED]
4−nbam BAM f i l e o f the matched normal sample [REQUIRED]
−tbam BAM f i l e o f the tumor sample [REQUIRED]
6−r e f Genome r e f e r e n c e f a / f a s t q f i l e [REQUIRED]
−t ru th Ground tr u th VCF f i l e . Every o th er p o s i t i o n i s a Fa ls e P o s i ti v e .
8−dbsnp dbSNP VCF f i l e
−cosmic COSMIC VCF f i l e
10 −mutect VCF f i l e from e i t h e r MuTect2 , MuTect, or I n de lo c at o r
−sn ip er VCF f i l e from SomaticSniper
12 −varscan VCF f i l e from VarScan2
−jsm VCF f i l e from Bina ’ s workflow that con tai ns JointSNVMix2
14 −va rd ic t VCF f i l e that con tai ns only SNV or only INDEL from VarDict
−muse VCF f i l e from MuSE
16 −l o f r e q VCF f i l e from LoFreq
6
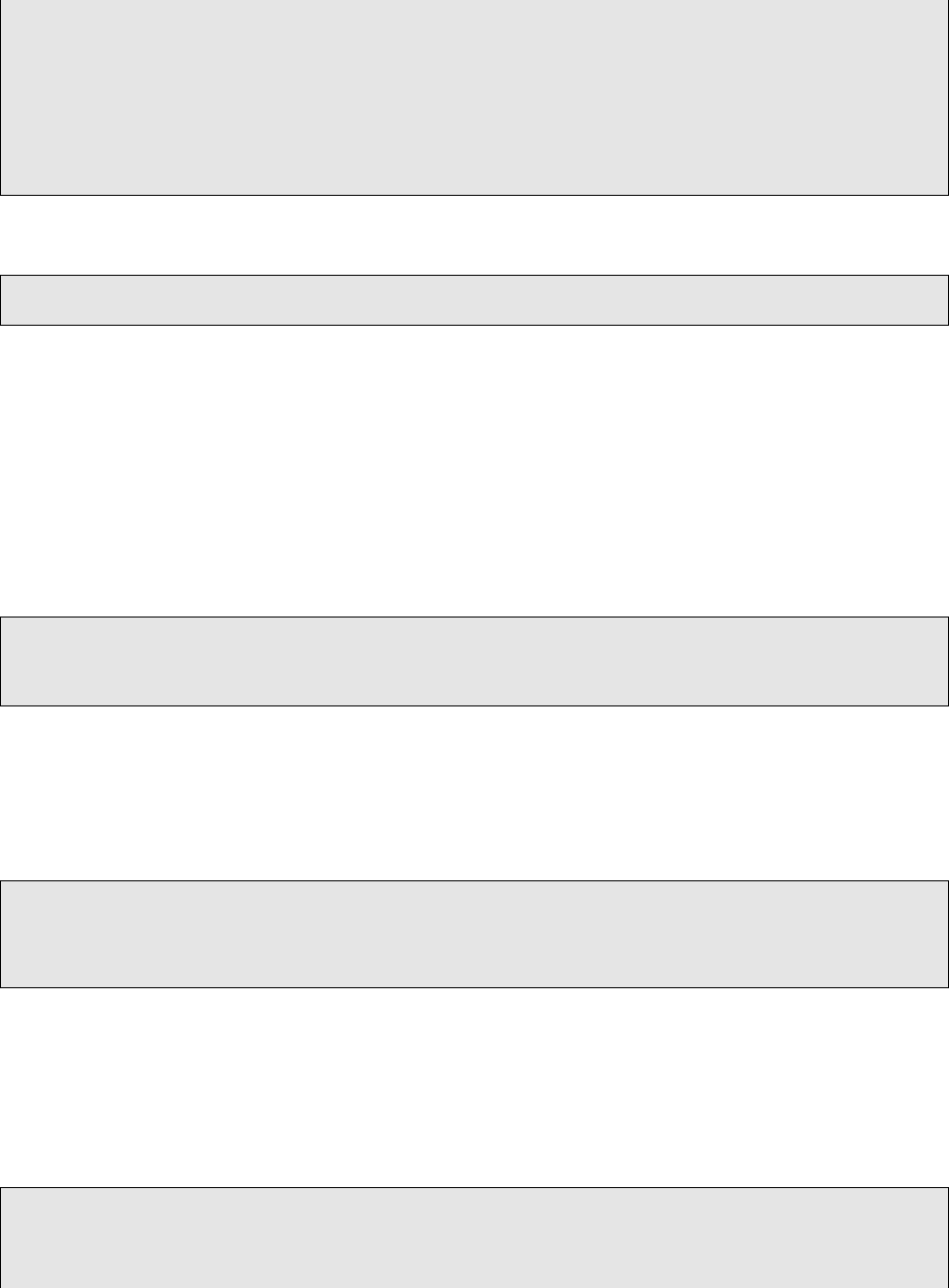
−s c a l p e l VCF f i l e from S ca l pe l
18 −s t r e l k a VCF f i l e from S tr el k a
−dedup A f l a g to c on si d er only primary reads
20 −minMQ Minimum mapping q ual it y for reads to be c ons idere d ( Default = 1)
−minBQ Minimum base q ua li ty f o r reads to be c ons idere d ( Defau lt = 5)
22 −mi nc al le r Minimum number o f caller classification f o r a c a l l to be co ns id ered ( Use 0. 5 to
co ns ide r some LowQual c a l l s . Default = 0) .
−s c a l e The op ti on s a re phred , f r a ct i o n , o r None , t o conv ert numbers to Phred s c a l e o r
f r a c t i o n a l s c a l e . ( d e fa ul t = None , i . e . , no co nv er si on )
24 −o u t f i l e Output TSV f i l e name
Note: Do not worry if Python throws a warning like this.
1RuntimeWarning : i n v a l i d val ue enco unte red i n double_scalars
z = ( s −expected ) / np . sqr t ( n1∗n2∗(n1+n2+1)/ 12. 0)
This is to tell you that scipy was attempting some statistical test with empty data. That’s usually due
to the fact that normal BAM le has no variant reads at that given position. That is why lots of values are
NaN for the normal.
3.4 Model Training or Mutation Prediction
You can use Ensemble.sSNV.tsv and Ensemble.sINDEL.tsv les either for model training (provided that their
mutation status is annotated with 0 or 1) or mutation prediction. This is done with stochastic boosting
algorithm we have implemented in R.
Model training:
# Training :
2r_ sc ri pt s /ada_model_builder_ntChange .R Ensemble . sSNV. t sv Consistent_Mates Inconsistent_Mates
r_ sc ri pt s /ada_model_builder_ntChange .R Ensemble . sINDEL . t sv Strelka_QSS Strelka_TQSS
Consistent_Mates Inconsistent_Mates
Ensemble.sSNV.tsv.Classier.RData and Ensemble.sINDEL.tsv.Classier.RData will be created from
model training. The arguments after Ensemble.sSNV.tsv and Ensemble.sINDEL.tsv tells the builder script
to ignore those features in training. These features do not improve accuracy in our data sets (mostly WGS
data, but they may help other data sets)
Mutation prediction:
1# Mutation pr ed i ct io n :
r_ sc ri pt /ada_model_predictor .R Ensemble .sSNV. tsv . C l a s s i f i e r . RData Ensemble . sSNV. t sv Trained .
sSNV . tsv
3r_ sc ri pt /ada_model_predictor .R Ensemble .sINDEL . t sv . C l a s s i f i e r . RData Ensemble . sINDEL . tsv Trained .
sINDEL . ts v
After mutation prediction, if you feel like it, you may convert Trained.sSNV.tsv and Trained.sINDEL.tsv
into VCF les. Use -tools to list ONLY the individual tools used to have appropriately annotated VCF
les. Accepted tools are MuTect2/MuTect/Indelocator, VarScan2, JointSNVMix2, SomaticSniper, VarDict,
MuSE, LoFreq, Scalpel, Strelka, and/or TNscope. To list a tool without having run it, the VCF will be
annotated as if the tool was run but did not identify that position as a somatic variant, which is probably
undesireable.
1# Pr obabi li ty above 0. 7 l ab el e d PASS (−p ass 0 . 7 ) , and between 0 . 1 and 0. 7 l a b e l e d LowQual (−low
0 . 1 ) :
# Use −a l l to incl ud e REJECT c a l l s in the VCF f i l e
3# Use −phred to c on ve rt p r o b a b i l i t y v a lu es ( between 0 t o 1) i n to Phred s c a l e i n t he QUAL column
in the VCF f i l e
7
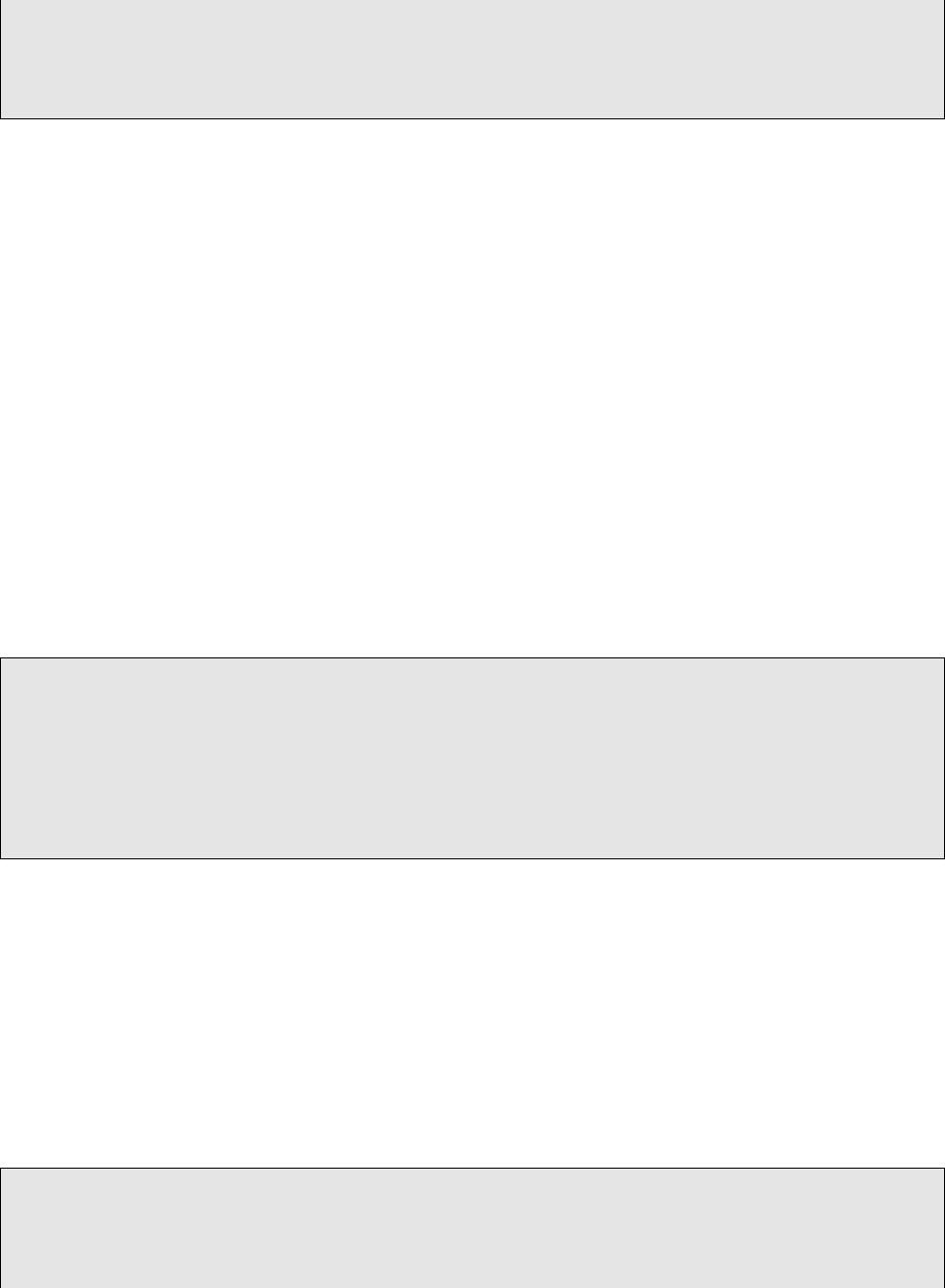
5SSeq_tsv2vcf .py −tsv Trained . sSNV. tsv −vc f Trained .sSNV . v cf −pass 0. 7 −low 0. 1 −t o o l s MuTect2
VarScan2 JointSNVMix2 SomaticSniper VarDict MuSE LoFreq S tr el ka −a l l −phred
7SSeq_tsv2vcf .py −ts v Tra ined . sINDEL . t s v −v cf Trained . sINDEL. v cf −pass 0 .7 −low 0 .1 −t o o l s MuTect2
VarScan2 VarDict LoFreq Sca lp el Strelk a −a l l −phred
4 To run the dockerized somatic mutation callers
For your convenience, we have created a couple of scripts that can generate run script for the dockerized
somatic mutation callers.
4.1 Location
• somaticseq/utilities/dockered_pipelines/
4.2 Requirements
• Have internet connection, and able to pull and run docker images from docker.io
• Have cluster management system such as Sun Grid Engine, so that the ”qsub” command is valid
4.3 Example commands
4.3.1 Single-threaded Jobs
This is best suited for whole exome sequencing or less.
1# Example command to submit the run s c r i p t s f o r each o f the f ol l ow in g somatic mutation c a l l e r s
$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/ submit_callers_singleThread . sh \
3−−normal−bam /ABSOLUTE/PATH/TO/normal_sample .bam \
−−tumor−bam /ABSOLUTE/PATH/TO/tumor_sample .bam \
5−−human−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38. f a \
−−output−d i r /ABSOLUTE/PATH/TO/RESULTS \
7−−dbsnp /ABSOLUTE/PATH/TO/dbSNP.GRCh38. vc f \
−−somaticseq−d i r /ABSOLUTE/PATH/TO/ SomaticSeq \
9−−ac tion echo \
−−mutect2 −−somaticsniper −−v ar di ct −−muse −− lofreq −− scalpel −− strelka −−somaticseq
The command shown above will create scripts for MuTect2, SomaticSniper, VarDict, MuSE, LoFreq, Scalpel,
and Strelka. Then, it will create the SomaticSeq script that merges those 7 callers. This command defaults
to majority-vote consensus.
Since it’s –aciton echo, it will echo the mutation caller scripts locations, but these scripts will not be
run. If you do –action qsub instead, then those mutation caller scripts will be qsub’ed. You’ll still need to
mantually run/submit the SomaticSeq script after all the caller jobs are done.
4.3.2 Multi-threaded Jobs
This is best suited for whole genome sequencing. This is same as above, except it will create 36 equal-size
regions in 36 bed les, and parallelize the jobs into 36 regions.
# Submitting mutation c a l l e r j obs by s p l i t t i n g each job in to 36 even re gi ons .
2$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/ submit_callers_multiThreads . sh \
−−normal−bam /ABSOLUTE/PATH/TO/normal_sample .bam \
4−−tumor−bam /ABSOLUTE/PATH/TO/tumor_sample .bam \
−−human−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38. f a \
6−−output−d i r /ABSOLUTE/PATH/TO/RESULTS \
8

−−dbsnp /ABSOLUTE/PATH/TO/dbSNP.GRCh38. vc f \
8−−threads 36 \
−−ac tion echo \
10 −−mutect2 −−somaticsniper −−v ar di ct −−muse −− lofreq −− scalpel −− strelka −−somaticseq
4.3.3 SomaticSeq Training
Two classiers will be created (*.RData les), one for SNV and one for INDEL.
# Submitting mutation c a l l e r j obs by s p l i t t i n g each job in to 36 even re gi ons .
2$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/ submit_callers_singleThread . sh \
−−normal−bam /ABSOLUTE/PATH/TO/normal_sample .bam \
4−−tumor−bam /ABSOLUTE/PATH/TO/tumor_sample .bam \
−−truth−snv /ABSOLUTE/PATH/TO/snvTruth . v cf \
6−−truth−i n d e l /ABSOLUTE/PATH/TO/ i nd el Tr uth . v c f \
−−human−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38. f a \
8−−output−d i r /ABSOLUTE/PATH/TO/RESULTS \
−−dbsnp /ABSOLUTE/PATH/TO/dbSNP.GRCh38. vc f \
10 −−somaticseq−d i r /ABSOLUTE/PATH/TO/ SomaticSeq \
−−ac tion echo \
12 −−mutect2 −−somaticsniper −−v ar di ct −−muse −− lofreq −− scalpel −− strelka −−somaticseq −−somaticseq
−train
Notice the command includes –truth-snv and –truth-indel, and invokes somaticseq-train.
For multi-threaded job, you should not invoke somaticseq-train. Instead, you should combine all the
Ensemble.sSNV.tsv and Ensemble.sINDEL.tsv les (separately), and then train on the combined les.
4.3.4 SomaticSeq Prediction
# Submitting mutation c a l l e r j obs by s p l i t t i n g each job in to 36 even re gi ons .
2$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/ submit_callers_singleThread . sh \
−−normal−bam /ABSOLUTE/PATH/TO/normal_sample .bam \
4−−tumor−bam /ABSOLUTE/PATH/TO/tumor_sample .bam \
−− classifier−snv /ABSOLUTE/PATH/TO/Ensemble .sSNV. tsv . ntChange . C l a s s i f i e r . RData \
6−− classifier−in de l /ABSOLUTE/PATH/TO/Ensemble . sINDEL . ts v . ntChange . C l a s s i f i e r . RData \
−−human−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38 . f a \
8−−output−d i r /ABSOLUTE/PATH/TO/RESULTS \
−−dbsnp /ABSOLUTE/PATH/TO/dbSNP.GRCh38. vc f \
10 −−somaticseq−d i r /ABSOLUTE/PATH/TO/SomaticSeq \
−−ac tion echo \
12 −−mutect2 −−somaticsniper −−v ar di ct −−muse −− lofreq −− scalpel −− strelka −−somaticseq
Notice the command includes –classier-snv and –classier-indel.
4.3.5 Parameters
−−normal−bam /ABSOLUTE/PATH/TO/normal_sample .bam ( Required )
2−−tumor−bam /ABSOLUTE/PATH/TO/tumor_sample .bam ( Required )
−−human−r e f e r e n c e /ABSOLUTE/PATH/TO/ human_reference . f a ( Required )
4−−dbsnp /ABSOLUTE/PATH/TO/dbsnp . vc f ( Required f o r MuSE and LoFreq )
−−cosmic /ABSOLUTE/PATH/TO/cosmic . v cf ( Optional )
6−− s e l e c t o r /ABSOLUTE/PATH/TO/Capture_region . bed ( Optional . Will assume whole
genome from the . f a i f i l e without i t . )
−−ex clude /ABSOLUTE/PATH/TO/ B la ck li st _r eg io n . bed ( Optional )
8−−min−a f ( O pti onal . The minimum VAF c u t o f f for VarDict and VarScan2 .
Defa ult s are 0 .10 f o r VarScan2 and 0.05 f o r VarDict ) .
−−ac tion qsub ( Optional : the command pre ced ing the .cmd s c r i p t s . Defa ult i s
echo)
9

10 −−threads 36 ( Optional f o r multiThreads and i n v a l i d f o r singleThread : evenly
s p l i t the genome i nt o 36 BED f i l e s . Default = 12) .
−−mutect2 ( Optional f l a g to invoke MuTect2)
12 −−vars can2 ( Optional f l a g to i nvo ke VarScan2 )
−−jo ints nvmix2 ( O ptional f l a g to invoke JointSNVMix2)
14 −−so ma ti cs nip er ( O ptional f l a g to invoke Somat icSniper )
−−v ar di ct ( O ptional f l a g to invoke VarDict )
16 −−muse ( O ptional f l a g to invoke MuSE)
−− l o f r e q ( O ptional f l a g to invoke LoFreq )
18 −− s c a l p e l ( O ptiona l f l a g to invo ke S ca lp e l )
−− s t r e l k a ( O ptional f l a g to i nvo ke S tr el ka )
20 −−so maticseq ( O ptional f l a g to invoke SomaticSeq . This s c r i p t always be echo ’ ed ,
as i t should not be submitted until a l l the c a l l e r s above complete ) .
−−output−d i r /ABSOLUTE/PATH/TO/OUTPUT_DIRECTORY ( R equired )
22 −−somaticseq−d i r SomaticSeq_Output_Directory ( O pti onal . The d i r e c t o r y name o f t he
SomaticSeq output . De fault = SomaticSeq ) .
−−somaticseq−tra i n ( Optional f l a g to invoke SomaticSeq to produce c l a s s i f i e r s i f
ground truth VCF f i l e s are provided . Only recommended in s in g le Th re ad mode , b ecau se o t he rw i se
it ’ s be tt er to combine the output TSV f i l e s f i r s t , and then train classifiers .)
24 −−somaticseq−actio n ( Optional . What to do with the somaticseq . cmd. Default i s echo .
Only do ”qsub” i f you have al re ad y completed a l l the mutation c a l l e r s , but want to run
SomaticSeq at a d i f f e r e n t s e tt i n g . )
−− classifier−snv Trained_sSNV_Classifier . RData ( Opti onal i f the re i s a c l a s s i f e r you
want to use )
26 −− classifier−i n d e l Trained_s INDEL_C lassifier . RData ( Opt ion al i f the re i s a c l a s s i f e r
you want to use )
−−truth−snv sSNV_ground_truth . vcf ( Optional i f the re i s a ground truth , and
everything e l s e w i l l be la be le d f a l s e positive)
28 −−truth−i n d e l sINDEL_ground_truth . v c f ( O pti onal i f there i s a ground truth , and
everything e l s e w i l l be la be le d f a l s e positive)
−−exome ( O ptional f l a g f o r S tr el ka )
30 −−scalpel−two−pass ( Optional parameter for Sc alp el . Default = f a l s e . )
−−mutect2−arguments ( Extra parameters to pass onto Mutect2 , e . g . , −−mutect2−arguments
’−− initial_tumor_lod 3 .0 −−log_somatic_prior −5.0 −−min_base_quality_score 20 ’)
32 −−mutect2−filter−arguments ( Extra parameters to pass onto F ilt erM ute ctC all s )
−−varscan−arguments ( Extra parameters to p ass onto VarScan2 )
34 −−varscan−pileup−arguments ( Extra par amete rs to pa ss onto sa mt oo ls mpileup th at c r e a t e s p i l eu p
files f o r VarScan )
−−jsm−train−arguments ( Extra par ame ter s t o pass onto JointSNVMix2 ’ s t r a i n command)
36 −−jsm−classify−arguments ( Extra parameters to pass onto JointSNVMix2 ’ s c l a s s i f y command)
−−somaticsniper−arguments ( Extra parameters to pass onto SomaticSniper )
38 −−var dict−arguments ( Extra parameters to pass onto VarDict )
−−muse−arguments ( Extra parameters to pass onto MuSE)
40 −−lofreq−arguments ( Extra para meter s to p ass onto LoFreq )
−−scalpel−discovery−arguments ( Extra parameters to pass onto S cal pel ’ s di sco ver y command)
42 −−scalpel−export−arguments ( Extra parameters to pass onto S cal pel ’ s export command)
−−strelka−config−arguments ( Extra parameters to pass onto St relk a ’ s c on f ig command)
44 −−strelka−run−arguments ( Extra parameters to pass onto S trek la ’ s run command)
−−somaticseq−arguments ( Extra parameters to pass onto SomaticSeq . Wrapper . sh )
4.3.6 What does the single-threaded command do
• For each ag such as --mutect2, --jointsnvmix2, ...., --strelka, a run script ending with .cmd will
be created in /ABSOLUTE/PATH/TO/RESULTS/logs. By default, these .cmd scripts will only be
created, and their le path will be printed on screen. However, if you do “--action qsub”, then these
scripts will be submitted via the qsub command. The default action is “echo.”
–Each of these .cmd script correspond to a mutation caller you specied. They all use docker
images.
–We may improve their functionalities in the future to allow more tunable parameters. For the
initial releases, POC and reproducibility take precedence.
10

• If you do “--somaticseq,” the somaticseq script will be created in /ABSOLUTE/PATH/TO/RESULT-
S/SomaticSeq/logs. However, it will not be submitted until you manually do so after each of these
mutation callers is nished running.
–In the future, we may create more sophisticated solution that will automatically solves these
dependencies. For the initial release, we’ll focus on stability and reproducibility.
• Due to the way those run scripts are written, the Sun Grid Engine’s standard error log will record the
time the task completes (i.e., Done at 2017/10/30 29:03:02), and it will only do so when the task is
completed with an exit code of 0. It can be a quick way to check if a task is done, by looking at the
nal line of the standard error log le.
4.3.7 What does the multi-threaded command do
It’s very similar to the single-threaded WES solution, except the job will be split evenly based on genomic
lengths.
• If you specied “--threads 36,” then 36 BED les will be created. Each BED le represents 1/36
of the total base pairs in the human genome (obtained from the .fa.fai le, but only including 1,
2, 3, ..., MT, or chr1, chr2, ..., chrM contigs). They are named 1.bed, 2.bed, ..., 36.bed, and will
be created into /ABSOLUTE/PATH/TO/RESULTS/1, /ABSOLUTE/PATH/TO/RESULTS/2, ...,
/ABSOLUTE/PATH/TO/RESULTS/36. You may, of course, specify any number. The default is 12.
• For each mutation callers you specify (with the exception of SomaticSniper), a script will be created
into /ABSOLUTE/PATH/TO/RESULTS/1/logs, /ABSOLUTE/PATH/TO/RESULTS/2/logs, etc.,
with partial BAM input. Again, they will be automatically submitted if you do “--action qsub.”
• Because SomaticSniper does not support partial BAM input (one would have to manually split the
BAMs in order to parallelize SomaticSniper this way), the above mentioned procedure is not applied
to SomaticSniper. Instead, a single-threaded script will be created (and potentially qsub’ed) into
/ABSOLUTE/PATH/TO/RESULTS/logs.
–However, because SomaticSniper is by far the fastest tool there, single-thread is doable even for
WGS. Even single-threaded SomaticSniper will likely nish before parallelized Scalpel. When I
benchmarked the DREAM Challenge Stage 3 by splitting it into 120 regions, Scalpel took 10
hours and 10 minutes to complete 1/120 of the data. SomaticSniper took a little under 5 hours
for the whole thing.
–After SomaticSniper nishes, the result VCF les will be split into each of the /ABSOLUTE/-
PATH/TO/RESULTS/1, /ABSOLUTE/PATH/TO/RESULTS/2, etc.
• JointSNVMix2 also does not support partial BAM input. Unlike SomaticSniper, it’s slow and takes
massive amount of memory. It’s not a good idea to run JointSNVMix2 on a WGS data. The only way
to do so is to manually split the BAM les and run each separately. We may do so in the future, but
JointSNVMix2 is a 5-year old that’s no longer being supported, so we probably won’t bother.
• Like the single-threaded case, a SomaticSeq run script will also be created for each partition like
/ABSOLUTE/PATH/TO/RESULTS/1/SomaticSeq/logs, but will not be submitted until you do so
manually.
–For simplicity, you may wait until all the mutation calling is done, then run a command like
1fi n d /ABSOLUTE/PATH/TO/RESULTS −name ’somaticseq∗.cmd’ −exec qsub {} \ ;
11

5 Use BAMSurgeon to create training data
For your convenience, we have created a couple of wrapper scripts that can generate the run script to create
training data using BAMSurgon at somaticseq/utilities/dockered_pipelines/bamSimulator. Descriptions
and example commands can be found in the README there.
This pipeline is used to spike in in silico somatic mutations into existing BAM les in order to create a
training set for somatic mutations.
After the in silico data are generated, you can use the somatic mutation pipeline on the training data to
generate the SomaticSeq classiers.
Classiers built on training data work if the training data is similar to the data you want to predict.
Ideally, the training data are sequenced on the same platform, same sample prep, and similar depth of
coverage as the data of interest.
This method is based on BAMSurgeon, slightly modied into our own fork for some speedups.
The proper citation for BAMSurgeon is Ewing AD, Houlahan KE, Hu Y, et al. Combining tumor genome
simulation with crowdsourcing to benchmark somatic single-nucleotide-variant detection. Nat Methods.
2015;12(7):623-30.
5.1 Requirements
• Have internet connection, and able to pull and run docker images from docker.io
• Have cluster management system such as Sun Grid Engine, so that the ”qsub” command is valid
5.2 Three scenario to simulate somatic mutations
Which scenario to use depend on the data sets available to you.
5.2.1 When you have sequencing replicates of normal samples
This is our approach to dene high-condence somatic mutations in SEQC2 consortium’s cancer reference
samples, presented here.
In this case, in silico mutations will be spiked into Replicate_002.bam. Since Replicate_002.bam and
Replicate_001.bam are otherwise the same sample, any mutations detected that you did not spike in are
false positives. The following command is a single-thread example.
1$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/bamSimulator/BamSimulator_singleThread . sh \
−−genome−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38. f a \
3−−tumor−bam−in /ABSOLUTE/PATH/TO/Replicate_001 . bam \
−−normal−bam−i n /ABSOLUTE/PATH/TO/ Replicate_002 . bam \
5−−tumor−bam−out syntheticTumor .bam \
−−normal−bam−out syntheticNormal .bam \
7−− split−proportion 0.5 \
−−num−snvs 20000 \
9−−num−i n d e l s 8000 \
−−min−vaf 0 .0 \
11 −−max−vaf 1 .0 \
−− left−beta 2 \
13 −−right−beta 5 \
−−min−variant−reads 2 \
15 −−output−d i r /ABSOLUTE/PATH/TO/ t r a i n i n g S e t \
−−ac tion qsub
BamSimulator_*.sh creates semi-simulated tumor-normal pairs out of your input tumor-normal pairs.
The ”ground truth” of the somatic mutations will be synthetic_snvs.vcf, synthetic_indels.vcf in the output
directory.
For multi-thread job (WGS), use BamSimulator_multiThreads.sh instead. See below for additional
options and parameters.
A schematic of the BAMSurgeon simulation procedure
12

5.2.2 This example mimicks DREAM Challenge
DREAM Somatic Mutation Calling Challenge was an international competition to nd algorithms that gave
the most accurate performances.
In that case, a high-coverage BAM le is randomly split into two. One of which is designated normal,
and the other one is designated tumor where mutations will be spiked in. Like the previous example, any
mutations found between the designated tumor and designated normal are false positive, since not only are
they from the same sample, but also from the same sequencing run. This example will not capture false
positives as a result of run-to-run biases if they exist in your sequencing data. It will, however, still capture
artefacts related to sequencing errors, sampling errors, mapping errors, etc.
$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/bamSimulator/BamSimulator_multiThreads . sh \
2−−genome−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38 . f a −−tumor−bam−in /ABSOLUTE/PATH/TO/
highCoverageGenome .bam −−tumor−bam−out syntheticTumor .bam −−normal−bam−out syntheticNormal .
bam −− split−proportion 0.5 −−num−snvs 10000 −−num−i n d e l s 8000 −−num−sv s 1500 −−min−vaf 0 .0
−−max−vaf 1 .0 −− left−beta 2 −−right−beta 5 −−min−variant−reads 2 −−output−d i r /ABSOLUTE/PATH/
TO/trainingSet −−threads 24 −−actio n qsub −− split−bam −−indel−realign −−merge−output−bams
The –split-bem will randomly split the high coverage BAM le into two BAM les, one of which is
designated normal and the other one designated tumor for mutation spike in. The –indel-realign is an option
that will perform GATK Joint Indel Realignment on the two BAM les. You may or may not invoke it
depending on your real data sets. The –merge-output-bams creates another script that will merge the BAM
and VCF les region-by-region. It will need to be run manually after all the spike in is done.
A schematic of the DREAM Challenge simulation procedure
5.2.3 Merge and then split the input tumor and normal BAM les
$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/bamSimulator/BamSimulator_multiThreads . sh \
2−−genome−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38 . f a −−tumor−bam−in /ABSOLUTE/PATH/TO/Tumor_Sample .bam
−−normal−bam−i n /ABSOLUTE/PATH/TO/Normal_Sample .bam −−tumor−bam−out syntheticTumor .bam −−
normal−bam−out syntheticNormal .bam −− split−proportion 0.5 −−num−snvs 30000 −−num−indels
10000 −−num−sv s 1500 −−min−vaf 0.0 −−max−vaf 1 .0 −− left−beta 2 −−right−beta 5 −−min−variant−
13

reads 2 −−output−d i r /ABSOLUTE/PATH/TO/ t r a i n i n g S e t −−threads 24 −−actio n qsub −−merge−bam −−
split−bam −− indel−realign −−merge−output−bams
The –merge-bam will merge the normal and tumor BAM les into a single BAM le. Then, –split-bem
will randomly split the merged BAM le into two BAM les. One of which is designated normal, and one
of which is designated tumor. Synthetic mutations will then be spiked into the designated tumor to create
”real” mutations. This is the approach described in our 2017 AACR Abstract.
A schematic of the simulation procedure
5.3 Parameters and Options
−−genome−r e f e r e n c e /ABSOLUTE/PATH/TO/human_reference . f a ( Required )
2−− s e l e c t o r /ABSOLUTE/PATH/TO/ cap tur e_r egion . bed (BED f i l e to l i m i t where mutation sp ike
in w i l l be attempted )
−−tumor−bam−in Input BAM f i l e ( Required )
4−−normal−bam−i n Input BAM f i l e ( Optional , but re qu ire d i f you want to merge i t with the tumor
input )
−−tumor−bam−out Output BAM f i l e f o r the de si gn at ed tumor a f t e r BAMSurgeon mutation s pi ke in
6−−normal−bam−out Output BAM f i l e f o r the desi gnated normal i f −− split−bam i s chosen
−− split−p ro po rt io n The f a c t i o n o f t o t a l rea ds d es gi na te d to the normal . ( Defaut = 0 . 5 )
8−−num−snvs Number of SNVs to sp ike in to the desi gn at ed tumor
−−num−i n d e l s Number o f INDELs to s pi k e i nt o the d es ig na te d tumor
10 −−num−sv s Number o f SVs to s pik e i nt o the desi gn at ed tumor ( Default = 0)
−−min−depth Minimum depth where spi ke i n can take pla ce
12 −−max−depth Maximum depth where spi ke in can take place
−−min−vaf Minimum VAF to sim ul at e
14 −−max−vaf Maximum VAF to simulate
−− left−beta L ef t beta o f beta d i s t r i b u t i o n f o r VAF
16 −−right−beta Right beta o f be ta d i s t r i b u t i o n f o r VAF
−−min−variant−reads Minimum number of variant−supporting reads f o r a s u c c e s s f u l s p ik e in
18 −−output−d i r Output d i r e c t o r y
−−merge−bam Flag to merge the tumor and normal bam f i l e input
20 −− split−bam Flag to s p l i t BAM f i l e f o r tumor and normal
−−clean−bam Flag to go through the BAM f i l e and remove read s where more than 2 i d e n t i c a l
read names are present , or rea ds where i t s read l en gt h and CIGAR s t r i n g do not match . This
was necessary f o r some BAM f i l e s downloaded from TCGA. However , a proper pair−end BAM f i l e
should not have the same read name appea ring more than t wi ce . Use t h i s only when n ec es sa ry a s
i t f i r s t s o r t s BAM f i l e by qname , go es through th e c le an i ng procedure , then re−s o r t by
coordinates .
22 −−indel−r e a l i g n Conduct GATK J oi nt I n de l Realignment on the two output BAM f i l e s . In st ea d o f
syntheticNormal .bam and syntheticTumor . bam, the f i n a l BAM f i l e s w i l l be syntheticNormal .
Join tReal igned . bam and syntheticTumor . JointR ealigne d .bam.
−−se ed Random se ed . Pi ck any i n t e g e r for r e p r o d u c i b i l i t y pu rpose s .
14

24 −−thr ea ds S p l i t the BAM f i l e s e ve nly in N regio ns , then p roce ss each ( p ai r ) o f sub−BAM
files in parallel .
−−ac tion The command pre ce din g the run s c r i p t c re at ed i nt o /ABSOLUTE/PATH/TO/
BamSurgeoned_SAMPLES/ l o g s . ”qsub” i s to submit the s c r i p t in SGE system . Defaul t = echo
5.3.1 –merge-bam / –split-bam / –indel-realign
If you have sequenced replicate normal, that’s the best data set for training. You can use one of the normal
as normal, and designate the other normal (of the same sample) as tumor. Use –indel-realign to invoke
GATK IndelRealign.
When you have a normal that’s roughly 2X the coverage as your data of choice, you can split that into
two halves. One designated as normal, and the other one designated as tumor. That DREAM Challenge’s
approach. Use –split-bam –indel-realign options.
Another approach is to merge the tumor and normal data, and then randomly split them as described
above. When you merge the tumor and normal, the real tumor mutations are relegated as germline or
noise, so they are considered false positives, because they are supposed to be evenly split into the designated
normal. To take this approach, use –merge-bam –split-bam –indel-realign options.
Don’t use –indel-realign if you do not use indel realignment in your alignment pipeline.
In some BAM les, there are reads where read lengths and CIGAR strings don’t match. Spike in will
fail in these cases, and you’ll need to invoke –clean-bam to get rid of these problematic reads.
You can control and visualize the shape of target VAF distribution with python command:
1import s ci p y . s t a t s a s s t a t s
import numpy as np
3import m at p lo t l ib . p yp lo t as p l t
5le ft Beta , rig th Beta = 2 ,5
minAF, maxAF = 0 ,1
7x = np . l i n s p a c e ( 0 , 1 , 1 01 )
y = s t a t s . beta . pdf ( x , l ef tB et a , r igthBeta , l o c = minAF, s c a l e = minAF + maxAF)
9_ = p l t . p lo t ( x , y )
5.4 To create SomaticSeq classiers
After the mutation simulation jobs are completed, you may create classiers with the training data with the
following command:
See our somatic mutation pipeline for more details.
1$PATH/TO/ somaticseq / u t i l i t i e s / doc ke re d_ pi pelines/ submit_callers_multiThreads . sh \
−−output−d i r /ABSOLUTE/PATH/TO/ t r a i n i n g S e t / s om at ic Mu ta ti on Pi pe li ne \
3−−normal−bam /ABSOLUTE/PATH/TO/ t ra in in gSe t /syntheticNormal .bam \
−−tumor−bam /ABSOLUTE/PATH/TO/ t ra in in gSe t /syntheticTumor . bam \
5−−human−r e f e r e n c e /ABSOLUTE/PATH/TO/GRCh38. f a \
−−dbsnp /ABSOLUTE/PATH/TO/dbSNP.GRCh38. vc f \
7−−thread 24 \
−−truth−snv /ABSOLUTE/PATH/TO/ t r a i n in g S e t / s yn th et ic _s nv s . v cf \
9−−truth−i n de l /ABSOLUTE/PATH/TO/ t r a in i ng S et / s y nt he ti c _i nd el s . l e f t A l i g n . v cf \
−−ac tion echo \
11 −−mutect2 −−somaticsniper −−v ar di ct −−muse −− lofreq −− strelka −−somaticseq
6 Release Notes
Make sure training and prediction use the same SomaticSeq version, or at least make sure the dierent minor
version changes do not change the results signicantly.
15
6.1 Version 1.0
Version used to generate data in the manuscript and Stage 5 of the ICGC-TCGA DREAM Somatic Mutation
Challenge, where SomaticSeq’s results were #1 for INDEL and #2 for SNV.
In the original manuscript, VarDict’s var2vcf_somatic.pl script was used to generate VarDict VCFs,
and subsequently “-lter somatic” was used for SSeq_merged.vcf2tsv.py. Since then (including DREAM
Challenge Stage 5), VarDict recommends var2vcf_paired.pl over var2vcf_somatic.pl, and subsequently “-
lter paired” was used for SSeq_merged.vcf2tsv.py. The dierence in SomaticSeq results, however, is pretty
much negligible.
6.2 Version 1.1
Automated the SomaticSeq.Wrapper.sh script for both training and prediction mode. No change to any
algorithm.
6.3 Version 1.2
Have implemented the following improvement, mostly for indels:
• SSeq_merged.vcf2tsv.py can now accept pileup les to extract read depth and DP4 (reference forward,
reference reverse, alternate forward, and alternate reverse) information (mainly for indels). Previously,
that information can only be extracted from SAMtools VCF. Since the SAMtools or HaplotypeCaller
generated VCFs hardly contain any indel information, this option improves the indel model. The
SomaticSeq.Wrapper.sh script is modied accordingly.
• Extract mapping quality (MQ) from VarDict output if this information cannot be found in SAMtools
VCF (also mostly benets the indel model).
• Indel length now positive for insertions and negative for deletions, instead of using the absolute value
previously.
6.4 Version 2.0
• Removed dependencies for SAMtools and HaplotypeCaller during feature extraction.
SSeq_merged.vcf2tsv.py extracts those information (plus more) directly from BAM les.
• Allow not only VCF le, but also BED le or a list of chromosome coordinate as input format for
SSeq_merged.vcf2tsv.py, i.e., use -mybed or -mypos instead of -myvcf.
• Instead of a separate step to annotate ground truth, that can be done directly by
SSeq_merged.vcf2tsv.py by supplying the ground truth VCF via -truth.
• SSeq_merged.vcf2tsv.py can annotate dbSNP and COSMIC information directly if BED le or a list
of chromosome coordinates are used as input in lieu of an annotated VCF le.
• Consolidated feature sets, e.g., removed some redunda Fixed a bug: if JointSNVMix2 is not included,
the values should be “NaN” instead of 0’s. This is to keep consistency with how we handle all other
callersnt feature sets coming from dierent resources.
6.5 Version 2.0.2
• Incorporated LoFreq.
• Used getopt to replace getopts in the SomaticSeq.Wrapper.sh script to allow long options.
16
6.6 Version 2.1.2
• Properly handle cases when multiple ALT’s are calls in the same position. The VCF les can either
contain multiple calls in the ALT column (i.e., A,G), or have multiple lines corresponding to the same
position (one line for each variant call). Some functions were signicantly re-written to allow this.
• Incorporated Scalpel.
• Deprecated HaplotypeCaller and SAMTools dependencies completely as far as feature generation is
concerned.
• The Wrapper script removed SnpSift/SnpE dependencies. Those information can be directly obtained
during the SSeq_merged.vcf2tsv.py step. Also removed some additional legacy steps that has become
useless since v2 (i.e., score_Somatic.Variants.py). Added a step to check the correctness of the input.
The v2.1 and 2.1.1 had some typos in the wrapper script, so only describing v2.1.2 here.
6.7 Version 2.2
• Added MuTect2 support.
6.8 Version 2.2.1
• InDel_3bp now stands for indel counts within 3 bps of the variant site, instead of exactly 3 bps from
the variant site as it was previously (likewise for InDel_2bp).
• Collapse MQ0 (mapping quality of 0) reads supporting reference/variant reads into a single metric of
MQ0 reads (i.e., tBAM_MQ0 and nBAM_MQ0). From experience, the number of MQ0 reads is at
least equally predictive of false positive calls, rather than distinguishing if those MQ0 reads support
reference or variant.
• Obtain SOR (Somatic Odds Ratio) from BAM les instead of VarDict’s VCF le.
• Fixed a typo in the SomaticSeq.Wrapper.sh script that did not handle inclusion region correctly.
6.9 Version 2.2.2
• Got around an occasional unexplained issue in then ada package were the SOR is sometimes categorized
as type, by forcing it to be numeric.
• Defaults PASS score from 0.7 to 0.5, and make them tunable in the SomaticSeq.Wrapper.sh script
(--pass-threshold and --lowqual-threshold).
6.10 Version 2.2.3
• Incorporated Strelka2 since it’s now GPLv3.
• Added another R script (ada_model_builder_ntChange.R) that uses nucleotide substitution pattern
as a feature. Limited experiences have shown us that it improves the accuracy, but it’s not heavily
tested yet.
• If a COSMIC site is labeled SNP in the COSMIC VCF le, if_cosmic and CNT will be labeled as 0.
The COSMIC ID will still appear in the ID column. This will not change any results because both of
those features are turned o in the training R script.
• Fixed a bug: if JointSNVMix2 is not included, the values should be “NaN” instead of 0’s. This is to
keep consistency with how we handle all other callers.
17
6.11 Version 2.2.4
• Resolved a bug in v2.2.3 where the VCF les of Strelka INDEL and Scalpel clash on GATK Com-
bineVariants, by outputting a temporary VCF le for Strelka INDEL without the sample columns.
• Caller classication: consider if_Scalpel = 1 only if there is a SOMATIC ag in its INFO.
6.12 Version 2.2.5
• Added a dockerle. Docker repo at https://hub.docker.com/r/lethalfang/somaticseq/.
• Ability to use vcfsort.pl instead of GATK CombineVariants to merge VCF les.
6.13 Version 2.3.0
• Moved some scripts to the utilities directory to clean up the clutter.
• Added the split_Bed_into_equal_regions.py to utilities, which will split a input BED le into multiple
BED les of equal size. This is to be used to parallelize large WGS jobs.
• Made compatible with MuTect2 from GATK4.
• Removed long options for the SomaticSeq.Wrapper.sh script because it’s more readable this way.
• Added a script to add “GT” eld to Strelka’s VCF output before merging it with other VCF les.
That was what caused GATK CombineVariants errors mentioned in v2.2.4’s release notes.
• Added a bunch of scripts at utilities/dockered_pipelines that can be used to submit (requiring Sun
Grid Engine or equivalent) dockerized pipeline to a computing cluster.
6.14 Version 2.3.1
• Improve the automated run script generator at utilities/dockered_pipelines.
• No change to SomaticSeq algorithm
6.15 Version 2.3.2
• Added run script generators for dockerized BAMSurgeon pipelines at utilities/dock-
ered_pipelines/bamSurgeon
• Added an error message to r_scripts/ada_model_builder_ntChange.R when TrueVariants_or_False
don’t have both 0’s and 1’s. Other than this warning message change, no other change to SomaticSeq
algorithm.
6.16 Version 2.4.0
• Restructured the utilities scripts.
• Added the utilities/lter_SomaticSeq_VCF.py script that “demotes” PASS calls to LowQual based
on a set of tunable hard lters.
• BamSurgeon scripts invokes modied BamSurgeon script that splits a BAM le without the need to
sort by read name. This works if the BAM les have proper read names, i.e., 2 and only 2 identical
read names for each paired-end reads.
• No change to SomaticSeq algorithm
18
6.17 Version 2.4.1
• Updated some docker job scripts.
• Added a script that converts some items in the VCF’s INFO eld into the sample eld, to pre-
cipitate the need to merge multiple VCF les into a single multi-sample VCF, i.e., utilities/refor-
mat_VCF2SEQC2.py.
• No change to SomaticSeq algorithm
6.18 Version 2.5.0
• In modify_VJSD.py, get rid of VarDict’s END tag (in single sample mode) because it causes problem
with GATK CombineVariants.
• Added limited single-sample support, i.e., ssSomaticSeq.Wrapper.sh is the wrapper script. singleSam-
ple_callers_singleThread.sh is the wrapper script to submit single-sample mutation caller scripts.
• Added run scripts for read alignments and post-alignment processing, i.e,. FASTQ →BAM, at utili-
ties/dockered_pipelines/alignments.
• Fixed a bug where the last two CD4 numbers were both alternate concordant reads in the output VCF
le, when the last number should’ve been alternate discordant reads.
• Changed the output le names from Trained.s(SNV|INDEL).vcf and Untrained.s(SNV|INDEL).vcf to
SSeq.Classied.s(SNV|INDE).vcf and Consensus.s(SNV|INDEL).vcf. No change to the actual tumor-
normal SomaticSeq algorithm.
• Added utilities/modify_VarDict.py to VarDict’s “complex” variant calls (e.g., GCA>TAC) into SNVs
when possible.
• Modied r_scripts/ada_model_builder_ntChange.R to allow you to ignore certain features, e.g.,
r_scripts/ada_model_builder_ntChange.R Training_Data.tsv nBAM_REF_BQ tBAM_REF_BQ
SiteHomopolymer_Length ...
Everything after the input le are features to be ignored during training.
Also added r_scripts/ada_cross_validation.R.
6.19 Version 2.5.1
• Additional passable parameters options to pass extra parameters to somatic mutation callers. Fixed a
bug where the “two-pass” parameter is not passed onto Scalpel in multiThreads scripts.
• Ignore Strelka_QSS and Strelka_TQSS for indel training in the SomaticSeq.Wrapper.sh script.
6.20 Version 2.5.2
• Ported some pipeline scripts to singularities at utilities/singularities.
6.21 Version 2.6.0
• VarScan2_Score is no longer extracted from VarScan’s output. Rather, it’s now calculated directly
using Fisher’s Exact Test, which reproduces VarScan’s output, but will have a real value when VarScan2
does not output a particular variant.
• Incorporate TNscope’s output VCF into SomaticSeq, but did not incorporate TNscope caller into the
dockerized workow because we don’t have distribution license.
19
6.22 Version 2.6.1
• Optimized memory for singularity scripts.
• Updated utilities/bamQC.py and added utilities/trimSoftClippedReads.py (removed soft-clipped bases
on soft-clipped reads)
• Added some docker scripts at utilities/dockered_pipelines/QC
6.23 Version 2.7.0
• Added another feature: consistent/inconsistent calls for paired reads if the position is covered by both
forward and reverse reads. However, they’re excluded as training features in SomaticSeq.Wrapper.sh
script for the time being.
• Change non-GCTA characters to N in VarDict.vcf le to make it conform to VCF le specications.
6.24 Version 2.7.1
• Without –gatk $PATH/TO/GenomeAnalysisTK.jar in the SomaticSeq.Wrapper.sh script, it will use
utilities/getUniqueVcfPositions.py and utilities/vcfsorter.pl to (in lieu of GATK3 CombineVariants) to
combine all the VCF les.
• Fixed bugs in the docker/singularities scripts where extra arguments for the callers are not correctly
passed onto the callers.
6.25 Version 2.7.2
• Make compatible with .cram format
• Fixed a bug where Strelka-only calls are not considered by SomaticSeq.
6.26 Version 2.8.0
• The program is now designed to crash if the VCF le(s) are not sorted according to the .fasta reference
le.
7 Contact Us
For suggestions, bug reports, or technical support, please post in
https://github.com/bioinform/somaticseq/issues. The developers are alerted when issues are created
there. Alternatively, you may also email li_tai.fang@roche.com.
20