R News 2007/1 PDF Rnews 2007 1
PDF Rnews_2007-1 CRAN: R News
User Manual: PDF CRAN - Contents of R News
Open the PDF directly: View PDF ![]() .
.
Page Count: 58
- Editorial
- Viewing Binary Files with the hexView Package
- FlexMix: An R Package for Finite Mixture Modelling
- Using R to Perform the AMMI Analysis on Agriculture Variety Trials
- Inferences for Ratios of Normal Means
- Working with Unknown Values
- A New Package for Fitting Random Effect Models
- Augmenting R with Unix Tools
- POT: Modelling Peaks Over a Threshold
- Backtests
- Review of John Verzani's Book Using R for Introductory Statistics
- DSC 2007
- New Journal: Annals of Applied Statistics
- Forthcoming Events: useR! 2007
- Changes in R 2.5.0
- Changes on CRAN
- R Foundation News
- R News Referees 2006
News
The Newsletter of the R Project Volume 7/1, April 2007
Editorial
by Torsten Hothorn
Welcome to the first issue of R News for 2007, which
follows the release of Rversion 2.5.0. This major revi-
sion, in addition to many other features, brings bet-
ter support of JAVA and Objective C to our desks.
Moreover, there is a new recommended package,
codetools, which includes functions that automagi-
cally check Rcode for possible problems.
Just before the release of R2.5.0 the fifth devel-
oper conference on “Directions in Statistical Com-
puting” was held in Auckland, NZ, the birthplace of
R. Hadley Wickham reports on the highlights of this
meeting. The Ruser community is not only active in
conferences. Volume 7, like the preceding volumes
of R News since 2001, wouldn’t be what it is without
the outstanding support of our referees. The editorial
board would like to say “Thank you!” to all who con-
tributed criticism and encouragement during the last
year—the complete list of referees in 2006 is given at
the end of this issue.
The scientific part of Volume 7 starts with an
article by Paul Murrell, our former editor-in-chief,
on handling binary files with tools provided by the
hexView package. Andrew Robinson teaches how
Rusers can make use of standard Unix tools, for
example mail for auto-generating large amounts of
email (not spam!). Many of us are regularly con-
fronted with data lacking a unique definition of miss-
ing values–the gdata package can help in this situa-
tion, as Gregor Gorjanc explains.
Bettina Grün and Fritz Leisch give an introduc-
tion to the flexmix package for finite mixture mod-
eling, analyzing a dataset on 21 different whiskey
brands. The analysis of field agricultural experi-
ments by means of additive main effect multiplica-
tive interactions is discussed by Andrea Onofri and
Egidio Ciriciofolo. Tests and confidence intervals for
ratios of means, such as ratios of regression coeffi-
cients, implemented in package mratio are described
by Gemechis Dilba and colleagues. The npmlreg
package for fitting random effect models is intro-
duced by Jochen Einbeck and his co-workers. Math-
ieu Ribatet models peaks over a threshold by POT,
and financial instruments like stocks or options are
(back-)tested by Kyle Campbell and colleagues.
Finally, I would like to remind everyone that the
next “useR!” conference is taking place in Ames,
Iowa, August 8–10. I hope to see you there!
Torsten Hothorn
Ludwig–Maximilians–Universität München
Germany
Torsten.Hothorn@R-project.org
Contents of this issue:
Editorial ...................... 1
Viewing Binary Files with the hexView Package 2
FlexMix: An R Package for Finite Mixture
Modelling .................... 8
Using R to Perform the AMMI Analysis on
Agriculture Variety Trials . . . . . . . . . . . 14
Inferences for Ratios of Normal Means . . . . . 20
Working with Unknown Values . . . . . . . . . 24
A New Package for Fitting Random Effect
Models...................... 26
Augmenting R with Unix Tools . . . . . . . . . 30
POT: Modelling Peaks Over a Threshold . . . . 34
Backtests ...................... 36
Review of John Verzani’s Book
Using R for Introductory Statistics . . . . . . 41
DSC2007...................... 42
New Journal: Annals of Applied Statistics . . . 43
Forthcoming Events: useR!2007......... 43
Changes in R 2.5.0 . . . . . . . . . . . . . . . . . 43
ChangesonCRAN ................ 51
R Foundation News . . . . . . . . . . . . . . . . 56
R News Referees 2006 . . . . . . . . . . . . . . . 56

Vol. 7/1, April 2007 2
Viewing Binary Files with the hexView
Package
by Paul Murrell
I really like plain text files.
I like them because I can see exactly what is in
them. I can even easily modify the file if I’m feeling
dangerous. This makes me feel like I understand the
file.
I am not so fond of binary files. I always have to
use specific software to access the contents and that
software only shows me an interpretation of the ba-
sic content of the file. The raw content is hidden from
me.
Sometimes I want to know more about a real bi-
nary file, for example when I need to read data in a
binary format that no existing R function will read.
When things go wrong, like when an R workspace
file becomes “corrupt”, I may have a strong need to
know more.
Hex editors are wonderful tools that provide a
view of the raw contents of a binary (or text) file,
whether just to aid in understanding the file or to in-
spect or recover a file. The hexView package is an
attempt to bring this sort of facility to R.
Viewing raw text files
The viewRaw() function reads and displays the raw
content of a file. The content is displayed in three
columns: the left column provides a byte offset
within the file, the middle column shows the raw
bytes, and the right column displays each byte as an
ASCII character. If the byte does not correspond to
a printable ASCII character then a full stop is dis-
played.
As a simple example, we will look at a plain text
file, "rawTest.txt", that contains a single line of
text. This file was created using the following code
(on a Linux system).
> writeLines("test pattern", "rawTest.txt")
A number of small example files are included as
part of the hexView package and the hexViewFile()
function is provided to make it convenient to refer
to these files. The readLines() function from the
base package reads in the lines of a plain text file as
a vector of strings, so the plain text content of the file
"rawTest.txt" can be retrieved as follows.
> readLines(hexViewFile("rawTest.txt"))
[1] "test pattern"
The following code uses the viewRaw() function
from hexView to display the raw contents of this file.
> viewRaw(hexViewFile("rawTest.txt"))
0 : 74 65 73 74 20 70 61 74 | test pat
8 : 74 65 72 6e 0a | tern.
As this example shows, by default, the raw bytes
are printed in hexadecimal format. The first byte
in this file is 74, which is 7 ∗16 +4=116 in dec-
imal notation—the ASCII code for the character t.
This byte pattern can be seen several times in the file,
wherever there is a tcharacter.
The machine argument to the viewRaw() function
controls how the raw bytes are displayed. It defaults
to "hex" for hexadecimal output, but also accepts the
value "binary", which means that the raw bytes are
printed in binary format, as shown below.
> viewRaw(hexViewFile("rawTest.txt"),
machine="binary")
0 : 01110100 01100101 01110011 | tes
3 : 01110100 00100000 01110000 | t p
6 : 01100001 01110100 01110100 | att
9 : 01100101 01110010 01101110 | ern
12 : 00001010 | .
One noteworthy feature of this simple file is the
last byte, which has the hexadecimal value 0a (or
00001010 in binary; the decimal value 10) and no
printable ASCII interpretation. This is the ASCII
code for the newline or line feed (LF) special char-
acter that indicates the end of a line in text files. This
is a simple demonstration that even plain text files
have details that are hidden from the user by stan-
dard viewing software; viewers will show text on
separate lines, but do not usually show the “charac-
ter” representing the start of a new line.
The next example provides a more dramatic
demonstration of hidden details in text files. The file
we will look at contains the same text as the previous
example, but was created on a Windows XP system
with Notepad using “Save As...” and selecting “Uni-
code” as the “Encoding”. The readLines() function
just needs the file to be opened with the appropriate
encoding, then it produces the same result as before.
> readLines(
file(hexViewFile("rawTest.unicode"),
encoding="UCS-2LE"))
[1] "test pattern"
However, the raw content of the file is now very dif-
ferent.
> viewRaw(hexViewFile("rawTest.unicode"))
R News ISSN 1609-3631

Vol. 7/1, April 2007 3
0 : ff fe 74 00 65 00 73 00 | ..t.e.s.
8 : 74 00 20 00 70 00 61 00 | t. .p.a.
16 : 74 00 74 00 65 00 72 00 | t.t.e.r.
24 : 6e 00 0d 00 0a 00 | n.....
It is fairly straightforward to identify some parts of
this file. The ASCII codes from the previous example
are there again, but there is an extra 00 byte after each
one. This reflects the fact that, on Windows, Unicode
text is stored using two bytes per character1.
Instead of the 13 bytes in the original file, we
might expect 26 bytes in this file, but there are actu-
ally 30 bytes. Where did the extra bytes come from?
The first two bytes at the start of the file are a
byte order mark (BOM). With two bytes to store for
each character, there are two possible orderings of
the bytes; for example, the two bytes for the charac-
ter tcould be stored as 74 00 (called little endian) or
as 00 74 (big endian). The BOM tells software which
order has been used. Another difference occurs at the
end of the file. The newline character is there again
(with an extra 00), but just before it there is a 0d char-
acter (with an extra 00). This is the carriage return
(CR) character. On Windows, a new line in a text file
is signalled by the combination CR+LF, but on UNIX
a new line is just indicated by a single LF.
As this example makes clear, software sometimes
does a lot of work behind the scenes in order to dis-
play even “plain text”.
Viewing raw binary files
An example of a binary file is the native binary for-
mat used by R to store information via the save()
function. The following code was used to create the
file "rawTest.bin".
> save(rnorm(50), file="rawTest.bin")
We can view this file with the following code; the
nbytes argument is used to show the raw data for
only the first 80 bytes.
> viewRaw(hexViewFile("rawTest.bin"),
nbytes=80)
0 : 1f 8b 08 00 00 00 00 00 | ........
8 : 00 03 01 c0 01 3f fe 52 | .....?.R
16 : 44 58 32 0a 58 0a 00 00 | DX2.X...
24 : 00 02 00 02 05 00 00 02 | ........
32 : 03 00 00 00 04 02 00 00 | ........
40 : 00 01 00 00 10 09 00 00 | ........
48 : 00 01 7a 00 00 00 0e 00 | ..z.....
56 : 00 00 32 3f e7 60 e6 49 | ..2?. .I
64 : c6 fe 0d 3f e1 3b c5 2f | ...?.;./
72 : bb 4e 18 bf c4 9e 0f 1a | .N......
This is a good example of a binary file that is in-
triguing to view, but there is little hope of retriev-
ing any useful information because the data has been
compressed (encoded). In other cases, things are a
not so hopeless, and it is not only possible to view the
raw bytes, but also to see useful patterns and struc-
tures.
The next example looks at a binary file with a
much simpler structure. The file "rawTest.int" only
contains (uncompressed) integer values and was cre-
ated by the following code.
> writeBin(1:50, "rawTest.int", size=4)
This file only contains the integers from 1 to 50,
with four bytes used for each integer. The raw con-
tents are shown below; this time the nbytes argu-
ment has been used to show only the raw data for
the first 10 integers (the first 40 bytes).
> viewRaw(hexViewFile("rawTest.int"),
nbytes=40)
0 : 01 00 00 00 02 00 00 00 | ........
8 : 03 00 00 00 04 00 00 00 | ........
16 : 05 00 00 00 06 00 00 00 | ........
24 : 07 00 00 00 08 00 00 00 | ........
32 : 09 00 00 00 0a 00 00 00 | ........
None of the bytes correspond to printable ASCII
characters in this case, so the right column of out-
put is not terribly interesting. The viewRaw() func-
tion has two arguments, human and size, which con-
trol the way that the raw bytes are interpreted and
displayed. In this case, rather than interpreting each
byte as an ASCII character, it makes sense to interpret
each block of four bytes as an integer. This is done in
the following code using human="int" and size=4.
> viewRaw(hexViewFile("rawTest.int"),
nbytes=40, human="int", size=4)
0 : 01 00 00 00 02 00 00 00 | 1 2
8 : 03 00 00 00 04 00 00 00 | 3 4
16 : 05 00 00 00 06 00 00 00 | 5 6
24 : 07 00 00 00 08 00 00 00 | 7 8
32 : 09 00 00 00 0a 00 00 00 | 9 10
With this simple binary format, we can see how
the individual integers are being stored. The integer
1is stored as the four bytes 01 00 00 00, the integer
2as 02 00 00 00, and so on. This clearly demon-
strates the idea of little endian byte order; the least-
significant byte, the value 1, is stored first. In big en-
dian byte order, the integer 1would be 00 00 00 01
(as we shall see later).
The other option for interpreting bytes is "real"
which means that each block of size bytes is inter-
preted as a floating-point value. A simple example
1Over-simplification alert! Windows used to use the UCS-2 encoding, which has two bytes per character, but now it uses UTF-16, which
has two or four bytes per character. There are only two bytes per character in this case because these are common english characters.
R News ISSN 1609-3631

Vol. 7/1, April 2007 4
is provided by the file "rawTest.real", which was
generated by the following code. I have deliberately
used big endian byte order because it will make it
easier to see the structure in the resulting bytes.
> writeBin(1:50/50, "rawTest.real", size=8,
endian="big")
Here is an example of reading this file and inter-
preting each block of 8 bytes as a floating-point num-
ber. This also demonstrates the use of the width ar-
gument to explicitly control how many bytes are dis-
played per line of output.
> viewRaw(hexViewFile("rawTest.real"),
nbytes=40, human="real", width=8,
endian="big")
0 : 3f 94 7a e1 47 ae 14 7b | 0.02
8 : 3f a4 7a e1 47 ae 14 7b | 0.04
16 : 3f ae b8 51 eb 85 1e b8 | 0.06
24 : 3f b4 7a e1 47 ae 14 7b | 0.08
32 : 3f b9 99 99 99 99 99 9a | 0.10
Again, we are able to see how individual floating-
point values are stored. The following code takes this
a little further and allows us to inspect the bit repre-
sentation of the floating point numbers. The output
is shown in Figure 1.
> viewRaw(hexViewFile("rawTest.real"),
nbytes=40, human="real",
machine="binary", width=8,
endian="big")
The bit representation adheres to the IEEE Stan-
dard for Binary Floating-Point Arithmetic (IEEE,
1985;Wikipedia, 2006). Each value is stored in the
form sign ×mantissa ×2exponent. The first (left-most)
bit indicates the sign of the number, the next 11 bits
describe the exponent and the remaining 52 bits de-
scribe the mantissa. The mantissa is a binary frac-
tion, with bit icorresponding to 2−i.
For the first value in "rawTest.real", the first bit
has value 0indicating a positive number, the expo-
nent bits are 0111111 1001 = 1017, from which we
subtract 1023 to get −6, and the mantissa is an im-
plicit 1 plus 0 ×2−1+1×2−2+0×2−3+0×2−4+
0×2−5+1×2−6... =1.28.2So we have the value
1.28 ×2−6=0.02.
Viewing a Binary File in Blocks
As the examples so far have hopefully demonstrated,
being able to see the raw contents of a file can be a
very good way to teach concepts such as endianness,
character encodings, and floating-point representa-
tions of real numbers. Plus, it is just good fun to poke
around in a file and see what is going on.
In this section, we will look at some more ad-
vanced functions from the hexView package, which
will allow us to take a more detailed look at more
complex binary formats and will allow us to perform
some more practical tasks.
We will start by looking again at R’s native bi-
nary format. The file "rawTest.XDRint" contains
the integers 1 to 50 saved as a binary R object
and was produced using the following code. The
compress=FALSE is important to allow us to see the
structure of the file.
> save(1:50, file="rawTest.XDRint",
compress=FALSE)
We can view (the first 80 bytes of) the raw file us-
ing viewRaw() as before and this does show us some
interesting features. For example, we can see the text
RDX2 at the start of the file (it is common for files to
have identifying markers at the start of the file). If
we look a little harder, we can also see the first few
integers (1 to 9); the data is stored in an XDR format
(Wikipedia,2006a), which uses big endian byte or-
der, so the integers are in consecutive blocks of four
bytes that look like this: 00 00 00 01, then 00 00 00
02, and so on.
> viewRaw(hexViewFile("rawTest.XDRint"),
width=8, nbytes=80)
0 : 52 44 58 32 0a 58 0a 00 | RDX2.X..
8 : 00 00 02 00 02 04 00 00 | ........
16 : 02 03 00 00 00 04 02 00 | ........
24 : 00 00 01 00 00 10 09 00 | ........
32 : 00 00 01 78 00 00 00 0d | ...x....
40 : 00 00 00 32 00 00 00 01 | ...2....
48 : 00 00 00 02 00 00 00 03 | ........
56 : 00 00 00 04 00 00 00 05 | ........
64 : 00 00 00 06 00 00 00 07 | ........
72 : 00 00 00 08 00 00 00 09 | ........
It is clear that there is some text in the file and that
there are some integers in the file, so neither viewing
the whole file as characters nor viewing the whole
file as integers is satisfactory. What we need to be
able to do is view the text sections as characters and
the integer sections as integers. This is what the func-
tions memFormat(),memBlock(), and friends are for.
The memBlock() function creates a description of
a block of memory, specifying how many bytes are
in the block; the block is interpreted as ASCII char-
acters. The atomicBlock() function creates a de-
scription of a memory block that contains a single
value of a specified type (e.g., a four-byte integer),
and the vectorBlock() function creates a descrip-
tion of a memory block consisting of 1 or more mem-
ory blocks.
A number of standard memory blocks are prede-
fined: integer4 (a four-byte integer) and integer1,
2At least, as close as it is possible to get to 1.28 with a finite number of bits. Another useful thing about viewing raw values is that it
makes explicit the fact that most decimal values do not have an exact floating-point representation.
R News ISSN 1609-3631

Vol. 7/1, April 2007 5
0 : 00111111 10010100 01111010 11100001 01000111 10101110 00010100 01111011 | 0.02
8 : 00111111 10100100 01111010 11100001 01000111 10101110 00010100 01111011 | 0.04
16 : 00111111 10101110 10111000 01010001 11101011 10000101 00011110 10111000 | 0.06
24 : 00111111 10110100 01111010 11100001 01000111 10101110 00010100 01111011 | 0.08
32 : 00111111 10111001 10011001 10011001 10011001 10011001 10011001 10011010 | 0.10
Figure 1: The floating point representation of the numbers 0.02 to 0.10 following IEEE 754 in big endian byte
order.
integer2, and integer8;real8 (an eight-byte
floating-point number, or double) and real4; and
ASCIIchar (a single-byte character). There is also
a special ASCIIline memory block for a series of
single-byte characters terminated by a newline.
The memFormat() function collects a num-
ber of memory block descriptions together and
viewFormat() reads the memory blocks and displays
them.
As an example, the following code reads in the
"RDX2" header line of the file "rawTest.XDRint",
treats the next 39 bytes as just raw binary, ignoring
any structure, then reads the first nine integers (as in-
tegers). A new memory block description is needed
for the integers because the XDR format is big endian
(the predefined integer4 is little endian). The names
of the memory blocks within the format are used to
separate the blocks of output.
> XDRint <- atomicBlock("int", endian="big")
> viewFormat(hexViewFile("rawTest.XDRint"),
memFormat(saveFormat=ASCIIline,
rawBlock=memBlock(39),
integers=vectorBlock(XDRint,
9)))
========saveFormat
0 : 52 44 58 32 0a | RDX2.
========rawBlock
5 : 58 0a 00 00 00 02 00 | X......
12 : 02 04 00 00 02 03 00 | .......
19 : 00 00 04 02 00 00 00 | .......
26 : 01 00 00 10 09 00 00 | .......
33 : 00 01 78 00 00 00 0d | ..x....
40 : 00 00 00 32 | ...2
========integers
44 : 00 00 00 01 00 00 00 02 | 1 2
52 : 00 00 00 03 00 00 00 04 | 3 4
60 : 00 00 00 05 00 00 00 06 | 5 6
68 : 00 00 00 07 00 00 00 08 | 7 8
76 : 00 00 00 09 | 9
The raw 39 bytes can be further broken down—
see the description of R’s native binary format on
pages 11 and 12 of the “R Internals” manual (R De-
velopment Core Team,2006) that is distributed with
R—but that is beyond the scope of this article.
Extracting Blocks from a Binary
File
As well as viewing different blocks of a binary
file, we may want to extract the values from each
block. For this purpose, the readFormat() function
is provided to read a binary format, as produced
by memFormat(), and generate a "rawFormat" object
(but not explicitly print it3). A "rawFormat" object is
a list with a component "blocks" that is itself a list
of "rawBlock" objects, one for each memory block
defined in the memory format. A "rawBlock" object
contains the raw bytes read from a file.
The blockValue() function extracts the inter-
preted value from a "rawBlock" object. The
blockString() function is provided specifically
for extracting a null-terminated string from a
"rawBlock" object.
The following code reads in the file
"rawTest.XDRint" and just extracts the 50 integer
values.
> XDRfile <-
readFormat(hexViewFile("rawTest.XDRint"),
memFormat(saveFormat=ASCIIline,
rawBlock=memBlock(39),
integers=vectorBlock(XDRint,
50)))
> blockValue(XDRfile$blocks$integers)
[1] 1 2 3 4 5 6 7 8 9 10 11 12 13
[14] 14 15 16 17 18 19 20 21 22 23 24 25 26
[27] 27 28 29 30 31 32 33 34 35 36 37 38 39
[40] 40 41 42 43 44 45 46 47 48 49 50
A Caution
On a typical 32-bit platform, R uses 4 bytes for rep-
resenting integer values in memory and 8 bytes for
floating-point values. This means that there may be
limits on what sort of values can be interpreted cor-
rectly by hexView.
For example, if a file contains 8-byte integers, it is
possible to view each set of 8 bytes as an integer, but
on my system R can only represent an integer using
4 bytes, so 4 of the bytes are (silently) dropped. The
following code demonstrates this effect by reading
3There is a readRaw() function too.
R News ISSN 1609-3631

Vol. 7/1, April 2007 6
the file "testRaw.int" and interpreting its contents
as 8-byte integers.
> viewRaw(hexViewFile("rawTest.int"),
nbytes=40, human="int", size=8)
0 : 01 00 00 00 02 00 00 00 | 1
8 : 03 00 00 00 04 00 00 00 | 3
16 : 05 00 00 00 06 00 00 00 | 5
24 : 07 00 00 00 08 00 00 00 | 7
32 : 09 00 00 00 0a 00 00 00 | 9
An extended example:
Reading EViews Files
On November 18 2006, Dietrich Trenkler sent a mes-
sage to the R-help mailing list asking for a func-
tion to read files in the native binary format used
by Eviews, an econometrics software package (http:
//www.eviews.com/). No such function exists, but
John C Frain helpfully pointed out that an unoffi-
cial description of the basic structure of Eviews files
had been made available by Allin Cottrell (creator of
gretl, the Gnu Regression, Econometrics and Time-
series Library). The details of Allin Cottrell’s reverse-
engineering efforts are available on the web (http:
//www.ecn.wfu.edu/~cottrell/eviews_format/).
In this section, we will use the hexView package
to explore an Eviews file and produce a new function
for reading files in this format. The example data file
we will use is from Ramu Ramanathan’s Introduc-
tory Econometrics text (Ramanathan,2002). The data
consists of four variables measured on single family
homes in University City, San Diego, in 1990:
price: sale price in thousands of dollars.
sqft: square feet of living area.
bedrms: number of bedrooms.
baths: number of bath rooms.
The data are included in both plain text for-
mat, as "data4-1.txt", and Eviews format, as
"data4-1.wf1", as part of the hexViews package.4
For later comparison, the data from the plain text for-
mat are shown below, having been read in with the
read.table() function.
> read.table(hexViewFile("data4-1.txt"),
col.names=c("price", "sqft",
"bedrms", "baths"))
price sqft bedrms baths
1 199.9 1065 3 1.75
2 228.0 1254 3 2.00
3 235.0 1300 3 2.00
4 285.0 1577 4 2.50
5 239.0 1600 3 2.00
6 293.0 1750 4 2.00
7 285.0 1800 4 2.75
8 365.0 1870 4 2.00
9 295.0 1935 4 2.50
10 290.0 1948 4 2.00
11 385.0 2254 4 3.00
12 505.0 2600 3 2.50
13 425.0 2800 4 3.00
14 415.0 3000 4 3.00
An Eviews file begins with a header, starting with
the text “New MicroTSP Workfile” and including im-
portant information about the size of the header and
the number of variables and the number of observa-
tions in the file. The following code defines an ap-
propriate "memFormat" object for this header infor-
mation.
> EViewsHeader <-
memFormat(firstline=memBlock(80),
headersize=integer8,
unknown=memBlock(26),
numvblesplusone=integer4,
date=vectorBlock(ASCIIchar, 4),
unkown=memBlock(2),
datafreq=integer2,
startperiod=integer2,
startobs=integer4,
unkown=memBlock(8),
numobs=integer4)
We can use readFormat() to read this header from
the file as follows. The number of variables reported
is one greater than the actual number of variables
and also includes two “boiler plate” variables that
are always included in Eviews files (hence 7 instead
of the expected 4).
> data4.1.header <-
readFormat(hexViewFile("data4-1.wf1"),
EViewsHeader)
> data4.1.header
=========firstline
0 : 4e 65 77 20 4d 69 | New Mi
6 : 63 72 6f 54 53 50 | croTSP
12 : 20 57 6f 72 6b 66 | Workf
18 : 69 6c 65 00 00 00 | ile...
24 : d8 5e 0e 01 00 00 | .^....
30 : 00 00 00 00 08 00 | ......
36 : 15 00 00 00 00 00 | ......
42 : ff ff ff ff 21 00 | ....!.
48 : 00 00 00 00 00 00 | ......
54 : 00 00 06 00 00 00 | ......
60 : 0f 00 00 00 06 00 | ......
66 : 00 00 01 00 01 00 | ......
72 : 66 03 00 00 00 00 | f.....
78 : 00 00 | ..
=========headersize
80 : 90 00 00 00 00 00 00 00 | 144
=========unknown
88 : 01 00 00 00 01 00 | ......
4The original source of the files was: http://ricardo.ecn.wfu.edu/pub/gretl_cdrom/data/
R News ISSN 1609-3631

Vol. 7/1, April 2007 7
94 : 00 00 01 00 00 00 | ......
100 : 00 00 00 00 00 00 | ......
106 : 00 00 00 00 00 00 | ......
112 : 00 00 | ..
=========numvblesplusone
114 : 07 00 00 00 | 7
=========date
118 : d5 b7 0d 3a | ...:
=========unkown
122 : 06 00 | ..
=========datafreq
124 : 01 00 | 1
=========startperiod
126 : 00 00 | 0
=========startobs
128 : 01 00 00 00 | 1
=========unkown
132 : 00 5d 67 0e 01 59 | .]g..Y
138 : 8b 41 | .A
=========numobs
140 : 0e 00 00 00 | 14
We can extract some pieces of information from
this header and use them to look at later parts of the
file.
> headerSize <-
blockValue(
data4.1.header$blocks$headersize)
> numObs <-
blockValue(
data4.1.header$blocks$numobs)
At a location 26 bytes beyond the header size,
there are several blocks describing each variable in
the Eviews file. Each of these blocks is 70 bytes long
and contains information on the variable name and
the location within the file where the data values re-
side for that variable. The following code creates a
description of a block containing variable informa-
tion, then uses readFormat() to read the information
for the first variable (the number of bath rooms); the
offset argument is used to start reading the block
at the appropriate location within the file. We also
extract the location of the data for this variable.
> EViewsVbleInfo <-
memFormat(unknown=memBlock(6),
recsize=integer4,
memsize=integer4,
ptrtodata=integer8,
vblename=vectorBlock(ASCIIchar,
32),
ptrtohistory=integer8,
vbletype=integer2,
unknown=memBlock(6))
> data4.1.vinfo <-
readFormat(hexViewFile("data4-1.wf1"),
EViewsVbleInfo,
offset=headerSize + 26)
> data4.1.vinfo
=========unknown
170 : 00 00 00 00 0b 00 | ......
=========recsize
176 : 86 00 00 00 | 134
=========memsize
180 : 70 00 00 00 | 112
=========ptrtodata
184 : f6 03 00 00 00 00 00 00 | 1014
=========vblename
192 : 42 41 54 48 53 00 00 00 00 00 | BATHS.....
202 : 00 00 00 00 00 00 00 00 00 00 | ..........
212 : 00 00 00 00 00 00 00 00 00 00 | ..........
222 : 00 00 | ..
=========ptrtohistory
224 : 00 00 00 00 d5 b7 0d 3a | 0
=========vbletype
232 : 2c 00 | 44
=========unknown
234 : 60 02 10 00 01 00 | .....
> firstVbleLoc <-
blockValue(data4.1.vinfo$blocks$ptrtodata)
The data for each variable is stored in a block
containing some preliminary information followed
by the data values stored as eight-byte floating-point
numbers. The code below creates a description of a
block of variable data and then reads the data block
for the first variable.
> EViewsVbleData <- function(numObs) {
memFormat(numobs=integer4,
startobs=integer4,
unknown=memBlock(8),
endobs=integer4,
unknown=memBlock(2),
values=vectorBlock(real8,
numObs))
}
> viewFormat(hexViewFile("data4-1.wf1"),
EViewsVbleData(numObs),
offset=firstVbleLoc)
=========numobs
1014 : 0e 00 00 00 | 14
=========startobs
1018 : 01 00 00 00 | 1
=========unknown
1022 : 00 00 00 00 00 00 | ......
1028 : 00 00 | ..
=========endobs
1030 : 0e 00 00 00 | 14
=========unknown
1034 : 00 00 | ..
=========values
1036 : 00 00 00 00 00 00 fc 3f | 1.75
1044 : 00 00 00 00 00 00 00 40 | 2.00
1052 : 00 00 00 00 00 00 00 40 | 2.00
1060 : 00 00 00 00 00 00 04 40 | 2.50
1068 : 00 00 00 00 00 00 00 40 | 2.00
1076 : 00 00 00 00 00 00 00 40 | 2.00
1084 : 00 00 00 00 00 00 06 40 | 2.75
1092 : 00 00 00 00 00 00 00 40 | 2.00
1100 : 00 00 00 00 00 00 04 40 | 2.50
1108 : 00 00 00 00 00 00 00 40 | 2.00
1116 : 00 00 00 00 00 00 08 40 | 3.00
1124 : 00 00 00 00 00 00 04 40 | 2.50
1132 : 00 00 00 00 00 00 08 40 | 3.00
1140 : 00 00 00 00 00 00 08 40 | 3.00
This manual process of exploring the file struc-
ture can easily be automated within a function. The
hexView package includes such a function under the
R News ISSN 1609-3631

Vol. 7/1, April 2007 8
name readEViews(). With this function, we can read
in the data set from the Eviews file as follows.
> readEViews(hexViewFile("data4-1.wf1"))
Skipping boilerplate variable
Skipping boilerplate variable
BATHS BEDRMS PRICE SQFT
1 1.75 3 199.9 1065
2 2.00 3 228.0 1254
3 2.00 3 235.0 1300
4 2.50 4 285.0 1577
5 2.00 3 239.0 1600
6 2.00 4 293.0 1750
7 2.75 4 285.0 1800
8 2.00 4 365.0 1870
9 2.50 4 295.0 1935
10 2.00 4 290.0 1948
11 3.00 4 385.0 2254
12 2.50 3 505.0 2600
13 3.00 4 425.0 2800
14 3.00 4 415.0 3000
This solution is not the most efficient way to read
Eviews files, but the hexView package does make it
easy to gradually build up a solution, it makes it easy
to view the results, and it does provide a way to solve
the problem without having to resort to C code.
Summary
The hexView package provides functions for view-
ing the raw byte contents of files. This is useful for
exploring a file structure and for demonstrating how
information is stored on a computer. More advanced
functions make it possible to read quite complex bi-
nary formats using only R code.
Acknowledgements
At the heart of the hexView package is the
readBin() function and the core facilities for work-
ing with "raw" binary objects in R code (e.g.,
rawToChar()); thanks to the R-core member(s) who
were responsible for developing those features.
I would also like to thank the anonymous re-
viewer for useful comments on early drafts of this
article.
Bibliography
IEEE Standard 754 for Binary Floating-Point Arithmetic.
IEEE computer society, 1985. 4
R Development Core Team. R Internals. R Foun-
dation for Statistical Computing, Vienna, Austria,
2006. URL http://www.R-project.org. ISBN 3-
900051-14-3. 5
R. Ramanathan. INTRODUCTORY ECONOMET-
RICS WITH APPLICATIONS. Harcourt College, 5
edition, 2002. ISBN 0-03-034342-9. 6
Wikipedia. External data representation —
wikipedia, the free encyclopedia, 2006a. URL
http://en.wikipedia.org/w/index.php?
title=External_Data_Representation&oldid=
91734878. [Online; accessed 3-December-2006]. 4
Wikipedia. IEEE floating-point standard —
wikipedia, the free encyclopedia, 2006b. URL
http://en.wikipedia.org/w/index.php?
title=IEEE_floating-point_standard&oldid=
89734307. [Online; accessed 3-December-2006]. 4
Paul Murrell
Department of Statistics
The University of Auckland
New Zealand
paul@stat.auckland.ac.nz
FlexMix: An R Package for Finite Mixture
Modelling
by Bettina Grün and Friedrich Leisch
Introduction
Finite mixture models are a popular method for
modelling unobserved heterogeneity or for approx-
imating general distribution functions. They are ap-
plied in a lot of different areas such as astronomy, bi-
ology, medicine or marketing. An overview on these
models with many examples for applications is given
in the recent monographs McLachlan and Peel (2000)
and Frühwirth-Schnatter (2006).
Due to this popularity there exist many (stand-
alone) software packages for finite mixture mod-
elling (see McLachlan and Peel,2000;Wedel and Ka-
makura,2001). Furthermore, there are several dif-
ferent R packages for fitting finite mixture models
available on CRAN. Packages which use the EM algo-
R News ISSN 1609-3631

Vol. 7/1, April 2007 9
rithm for model estimation are flexmix,fpc,mclust,
mixreg,mixtools, and mmlcr. Packages with other
model estimation methods are bayesmix,depmix,
moc,vabayelMix and wle. A short description
of these packages can be found in the CRAN task
view on clustering (http://cran.at.r-project.
org/src/contrib/Views/Cluster.html).
Finite mixture models
A finite mixture model is given by a convex combina-
tion of Kdifferent components, i.e. the weights of the
components are non-negative and sum to one. For
each component it is assumed that it follows a para-
metric distribution or is given by a more complex
model, such as a generalized linear model (GLM).
In the following we consider finite mixture den-
sities h(·|·)with Kcomponents, dependent variables
yand (optional) independent variables x:
h(y|x,w,Θ) =
K
∑
k=1
πk(w,α)f(y|x,ϑk)
where ∀w,α:
πk(w,α)≥0∀k∧
K
∑
k=1
πk(w,α) = 1
and
ϑk6=ϑl∀k6=l.
We assume that the component distributions f(·|·)
are from the same distributional family with compo-
nent specific parameters ϑk. The component weights
or prior class probabilities πkoptionally depend on
the concomitant variables wand the parameters α
and are modelled through multinomial logit models
as suggested for example in Dayton and Macready
(1988). A similar model class is also described in
McLachlan and Peel (2000, p. 145). The model can
be estimated using the EM algorithm (see Dempster
et al.,1977;McLachlan and Peel,2000) for ML estima-
tion or using MCMC methods for Bayesian analysis
(see for example Frühwirth-Schnatter,2006).
A possible extension of this model class is to
either have mixtures with components where the
parameters of one component are fixed a-priori
(e.g. zero-inflated models; Grün and Leisch,2007b)
or to even allow different component specific mod-
els (e.g. for modelling noise in the data; Dasgupta
and Raftery,1998).
Design principles of FlexMix
The main reason for the implementation of the pack-
age was to allow easy extensibility and to have the
possibility for rapid prototyping in order to be able
to try out new mixture models. The package was im-
plemented using S4 classes and methods.
The EM algorithm provides a common basis for
estimation of a general class of finite mixture mod-
els and the package flexmix tries to enable the user
to exploit this commonness. flexmix provides the E-
step and takes care of all data handling while the user
is supposed to supply the M-step via model drivers
for the component-specific model and the concomi-
tant variable model. For the M-step available func-
tions for weighted maximum likelihood estimation
can be used as for example glm() for fitting GLMs or
multinom() in MASS for multinomial logit models.
Currently model drivers are available for
model-based clustering of multivariate Gaus-
sian distributions with diagonal or unrestricted
variance-covariance matrices (FLXMCmvnorm()) and
multivariate Bernoulli and Poisson distributions
(FLXMCmvbinary() and FLXMCmvpois()) where the
dimensions are mutually independent. flexmix does
not provide functionality for estimating mixtures
of Gaussian distributions with special variance-
covariance structures, as this functionality has al-
ready been implemented in the R package mclust
(Fraley and Raftery,2006).
For mixtures of regressions the Gaussian, bino-
mial, Poisson and gamma distribution can be speci-
fied (FLXMRglm()). If some parameters are restricted
to be equal over the components the model driver
FLXMRglmfix() can be used. Zero-inflated Poisson
and binomial regression models can be fitted us-
ing FLXMRziglm(). For an example of zero-inflated
models see example("FLXMRziglm"). For the con-
comitant variable models either constant component
weights (default) can be used or multinomial logit
models (FLXPmultinom()) can be fitted.
Estimation problems can occur if the components
become too small during the EM algorithm. In or-
der to avoid these problems a minimum size can be
specified for each component. This is especially im-
portant for finite mixtures of multivariate Gaussian
distributions where full variance-covariance matri-
ces are estimated for each component.
Further details on the implementation and the de-
sign principles as well as exemplary applications of
the package can be found in the accompanying vi-
gnettes "flexmix-intro" which is an updated ver-
sion of Leisch (2004) and "regression-examples"
and in Grün and Leisch (2007a). Note that this article
uses the new version 2.0 of the package, where the
names of some driver functions have changed com-
pared with older versions of flexmix.
Exemplary applications
In the following we present two examples for using
the package. The first example demonstrates model-
based clustering, i.e., mixtures without independent
variables, and the second example gives an applica-
tion for fitting mixtures of generalized linear regres-
R News ISSN 1609-3631
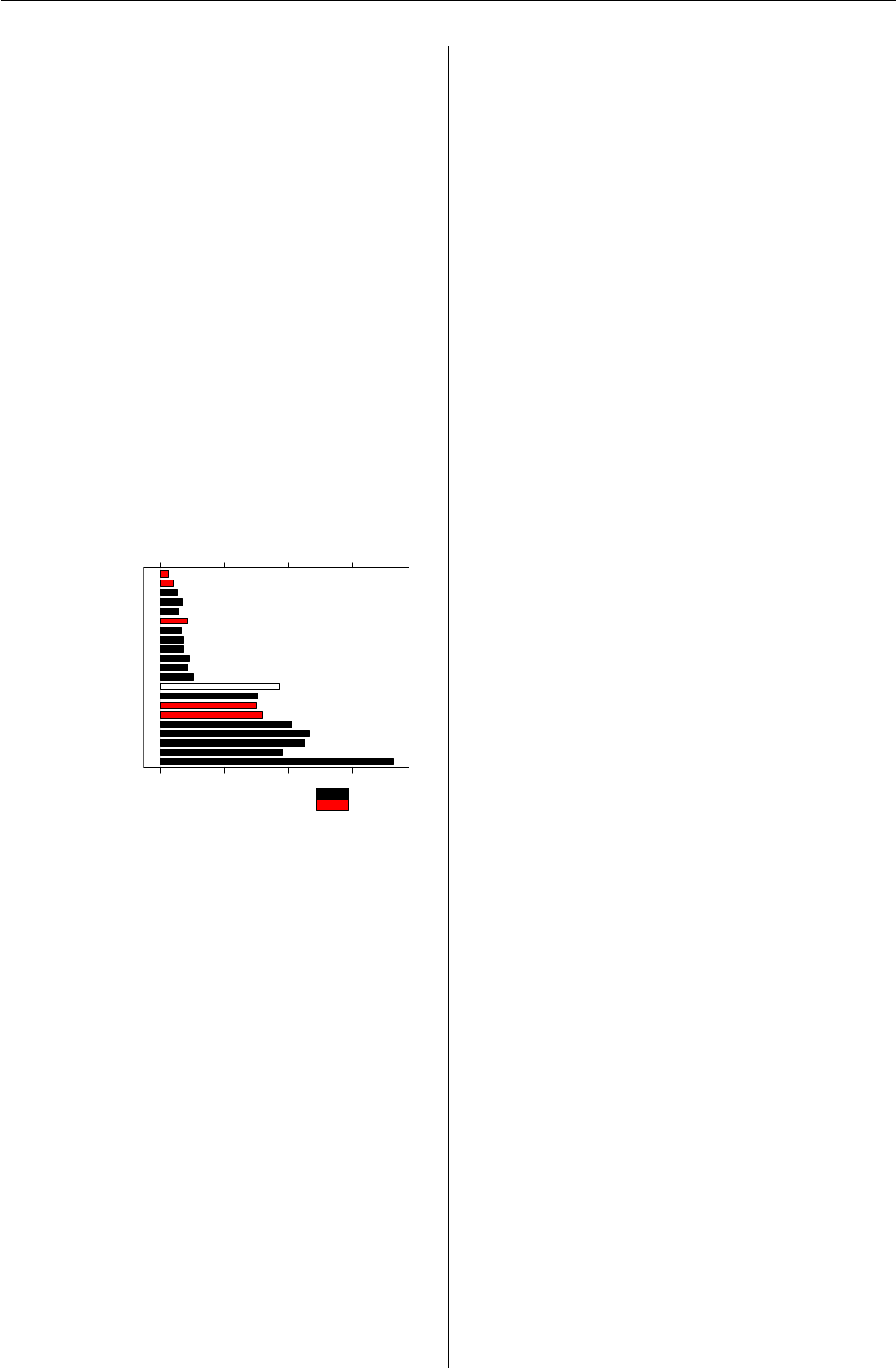
Vol. 7/1, April 2007 10
sion models.
Model-based clustering
The following dataset is taken from Edwards and Al-
lenby (2003) who refer to the Simmons Study of Me-
dia and Markets. It contains all households which
used any whiskey brand during the last year and
provides a binary incidence matrix on their brand
use for 21 whiskey brands during this year. This
means only the information on the different brands
used in a household is available.
We first load the package and the dataset. The
whiskey dataset contains observations from 2218
households. The relative frequency of usage for each
brand is given in Figure 1. Additional information
is available for the brands indicating the type of
whiskey: blend or single malt.
R> library("flexmix")
R> data("whiskey")
R> set.seed(1802)
Probability
Chivas Regal
Johnnie Walker Red Label
Johnnie Walker Black Label
Dewar's White Label
J&B
Glenlivet
Glenfiddich
Cutty Sark
Other brands
Pinch (Haig)
Ballantine
Clan MacGregor
Black & White
Passport
Grant's
Macallan
Ushers
Scoresby Rare
White Horse
Knockando
Singleton
0.0 0.1 0.2 0.3 Blend
Single Malt
Figure 1: Relative frequency of the whiskey brands.
We fit a mixture of binomial distributions to the
dataset where the variables in each component spe-
cific models are assumed to be independent. The
EM algorithm is repeated nrep = 3 times using ran-
dom initialization, i.e. each observation is assigned
to one component with an a-posteriori probability of
0.9 and 0.1 otherwise and the component is selected
with equal probability.
R> wh_mix <- stepFlexmix(Incidence ~ 1,
+ weights = ~ Freq, data = whiskey,
+ model = FLXMCmvbinary(truncated = TRUE),
+ control = list(minprior = 0.005),
+ k = 1:7, nrep = 3)
Model-based clustering uses no explanatory vari-
ables, hence the right hand side of the formula
Incidence ~ 1 is constant. The model driver
is FLXMCmvbinary() with argument truncated =
TRUE, as the number of non-users is not available and
a truncated likelihood is maximized in each M-step
again using the EM-algorithm. We vary the number
of components for k = 1:7. The best solution with
respect to the log-likelihood for each of the differ-
ent numbers of components is returned in an object
of class "stepFlexmix". The control argument can
be used to control the fitting with the EM algorithm.
With minprior the minimum relative size of the com-
ponents is specified, components falling below this
threshold are removed during the EM algorithm.
The dataset contains only the unique binary pat-
terns observed with the corresponding frequency.
We use these frequencies for the weights argument
instead of transforming the dataset to have one row
for each observation. The use of a weights argument
allows to use only the number of unique observa-
tions for fitting, which can substantially reduce the
size of the model matrix and hence speed up the es-
timation process. For this dataset this means that the
model matrix has 484 instead of 2218 rows.
Model selection can be made using information
criteria, as for example the BIC (see Fraley and
Raftery,1998). For this example the BIC suggests a
mixture with 5 components:
R> BIC(wh_mix)
1234
27705.1 26327.6 25987.7 25683.2
567
25647.0 25670.3 25718.6
R> wh_best <- getModel(wh_mix, "BIC")
R> wh_best
Call:
stepFlexmix(Incidence ~ 1,
weights = ~Freq, data = whiskey,
model = FLXMCmvbinary(truncated = TRUE),
control = list(minprior = 0.005),
k = 5, nrep = 3)
Cluster sizes:
12345
283 791 953 25 166
convergence after 180 iterations
The estimated parameters can be inspected using
accessor functions such as prior() or parameters().
R> prior(wh_best)
[1] 0.1421343 0.3303822
[3] 0.4311072 0.0112559
[5] 0.0851203
R> parameters(wh_best, component=4:5)[1:2,]
Comp.4
center.Singleton 0.643431
center.Knockando 0.601124
Comp.5
center.Singleton 2.75013e-02
center.Knockando 1.13519e-32
R News ISSN 1609-3631
Vol. 7/1, April 2007 11
Probability
Chivas Regal
Johnnie Walker Red Label
Johnnie Walker Black Label
Dewar's White Label
J&B
Glenlivet
Glenfiddich
Cutty Sark
Other brands
Pinch (Haig)
Ballantine
Clan MacGregor
Black & White
Passport
Grant's
Macallan
Ushers
Scoresby Rare
White Horse
Knockando
Singleton
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 1
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 2
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 3
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 4
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 5
Blend
Single Malt
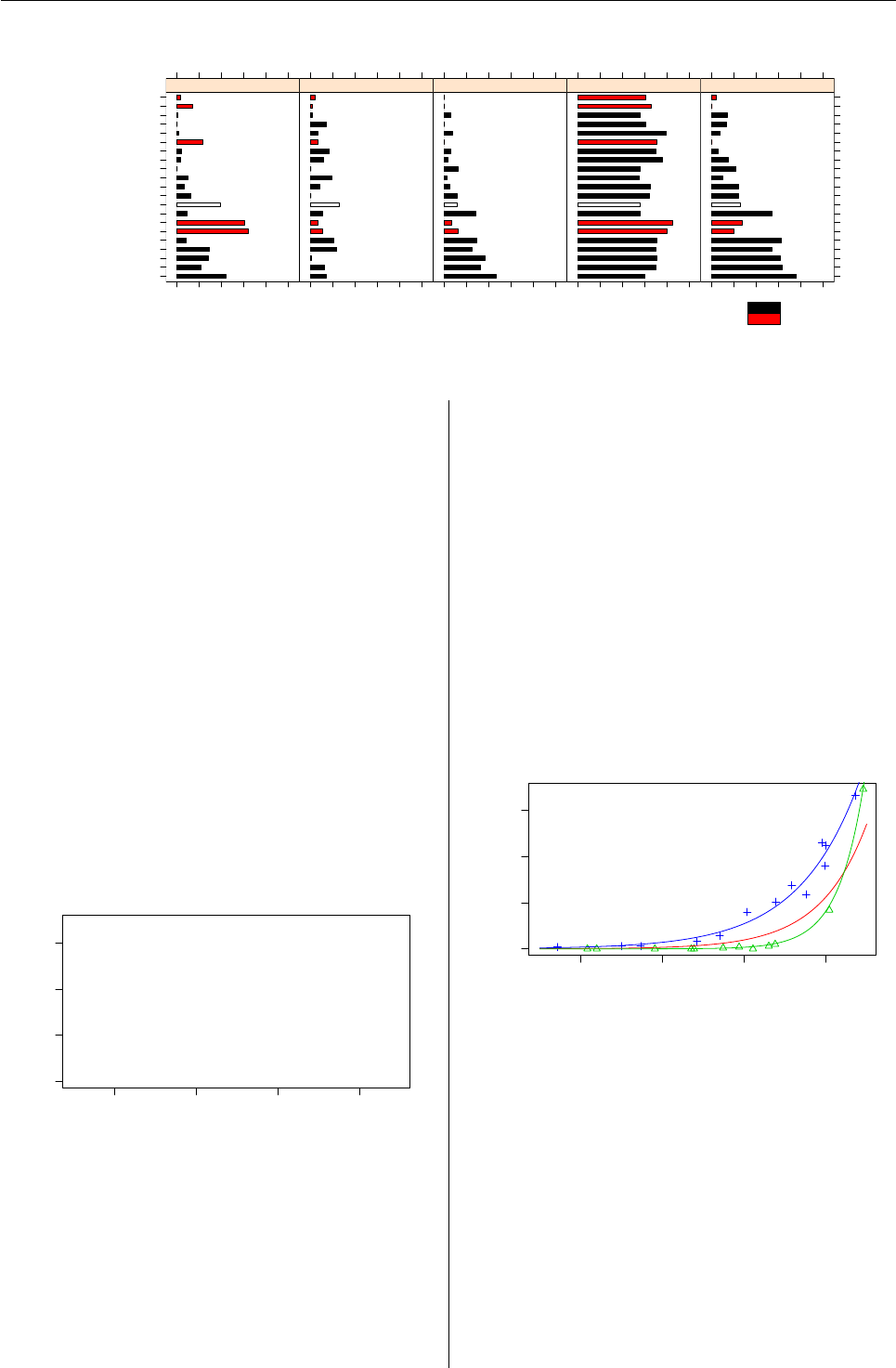
Figure 2: Estimated probability of usage for the whiskey brands for each component.
The fitted parameters of the mixture for each
component are given in Figure 2. It can be seen
that component 4 (1.1% of the households) contains
the households which bought the greatest number
of different brands and all brands to a similar ex-
tent. Households from component 5 (8.5%) also buy
a wide range of whiskey brands, but tend to avoid
single malts. Component 3 (43.1%) has a similar us-
age pattern as component 5 but buys less brands in
general. Component 1 (14.2%) seems to favour sin-
gle malt whiskeys and component 2 (33%) is espe-
cially fond of other brands and tends to avoid John-
nie Walker Black Label.
Mixtures of regressions
The patent data given in Wang et al. (1998) includes
70 observations on patent applications, R&D spend-
ing and sales in millions of dollar from pharmaceuti-
cal and biomedical companies in 1976 taken from the
National Bureau of Economic Research R&D Master-
file. The data is given in Figure 3.
●
●●●
●
●
●
●
●●
●
●
●
●●
●
●
●
●●
● ●
●●
●
●●
●●
●
● ●
●
●
●●● ●
●
●
●
● ●
●
●
●
●
●
●
●
● ● ●
●
●
●
●●
●
●
●
●
●
●
●
●
●
●
●
●
−2 0 2 4
0 50 100 150
lgRD
Patents
Figure 3: Patent dataset.
The model which is chosen as the best in Wang
et al. (1998) is a finite mixture of three Poisson regres-
sion models with Patents as dependent variable, the
logarithmized R&D spending lgRD as independent
variable and the R&D spending per sales RDS as con-
comitant variable. This model can be fitted in R with
the component-specific model driver FLXMRglm()
which allows fitting of finite mixtures of GLMs. As
concomitant variable model driver FLXPmultinom()
is used for a multinomial logit model where the pos-
terior probabilities are the dependent variables.
R> data("patent")
R> pat_mix <- flexmix(Patents ~ lgRD,
+ k = 3, data = patent,
+ model = FLXMRglm(family = "poisson"),
+ concomitant = FLXPmultinom(~RDS))
The observed values together with the fitted val-
ues for each component are given in Figure 4. The
coloring and characters used for plotting the ob-
servations are according to the component assign-
ment using the maximum a-posteriori probabili-
ties, which are obtained using cluster(pat_mix).
●● ●
●
●
●
●
●
●
●
●●
●
●●
●●
● ●
●
●
●●
●
●
● ●
●
●
●
●
●●
●
●
●
●
●
●
●
●
●
●
●
●
−2 0 2 4
0 50 100 150
lgRD
Patents
Figure 4: Patent data with fitted values for each com-
ponent.
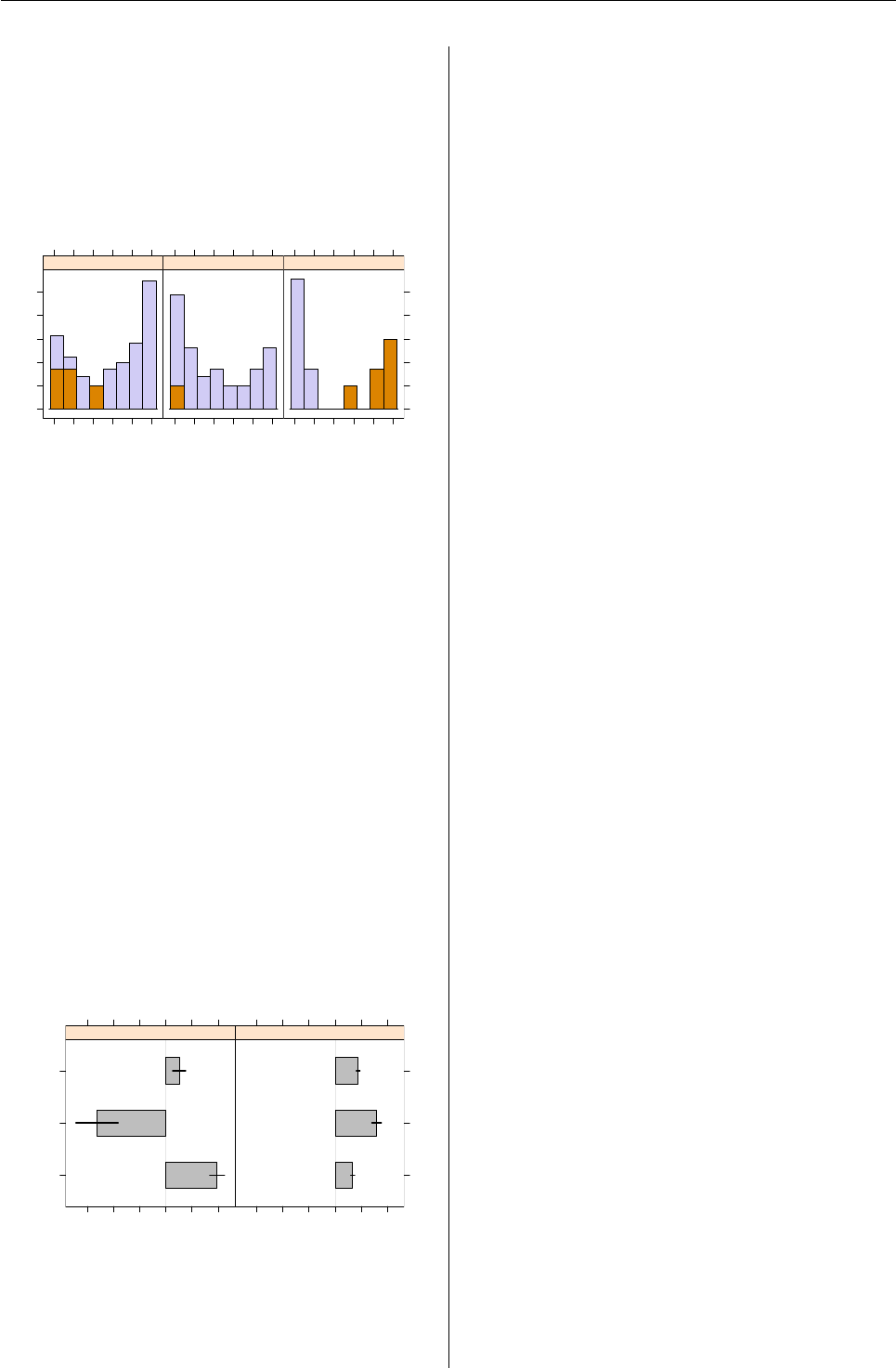
In Figure 5a rootogram of the posterior proba-
bilities of the observations is given. This is the de-
fault plot of the "flexmix" objects returned by the
fitting function. It can be used for arbitrary mix-
ture models and indicates how well the observations
are clustered by the mixture. For ease of interpre-
tation the observations with a-posteriori probability
less than eps=10−4are omitted as otherwise the peak
at zero would dominate the plot. The observations
where the a-posteriori probability is largest for the
third component are colored differently. The plot is
generated using the following command.
R News ISSN 1609-3631
Vol. 7/1, April 2007 12
R> plot(pat_mix, mark = 3)
The posteriors of all three components have modes
at 0 and 1, indicating well-separated clusters (Leisch,
2004). Note that the object returned by the plot func-
tion is of class "trellis", and that the plot itself
is produced by the corresponding show() method
(Sarkar,2002).
Rootogram of posterior probabilities > 1e−04
0
1
2
3
4
5
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 1
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 2
0.0 0.2 0.4 0.6 0.8 1.0
Comp. 3
Figure 5: Rootogram of the posterior probabilities.
Further details of the fitted mixture can be ob-
tained with refit() which returns the fitted values
together with approximate standard deviations and
significance tests, see Figure 6. The standard devi-
ations are only approximative because they are de-
termined separately for each component and it is not
taken into account that the components have been
estimated simultaneously. In the future functionality
to determine the standard deviations using either the
full Hesse matrix or the parametric bootstrap shall be
provided.
The estimated coefficients are given in Figure 7.
The black lines indicate the (approximative) 95%
confidence intervals. This is the default plot for the
objects returned by refit() and is obtained with the
following command.
R> plot(refit(pat_mix), bycluster = FALSE)
The argument bycluster indicates if the clus-
ters/components or the different variables are
used as conditioning variables for the panels.
Comp. 3
Comp. 2
Comp. 1
−3 −2 −1 0 1 2
(Intercept)
−3 −2 −1 0 1 2
lgRD
Figure 7: Estimated coefficients of the component
specific models with corresponding 95% confidence
intervals.
The plot indicates that the estimated coefficients
vary between all components even though the co-
efficients for lgRD are similar for the first and third
component. A smaller model where these coeffi-
cients are restricted to be equal can be fitted using
the model driver FLXMRglmfix(). The EM algorithm
can be initialized in the original solution using the
estimated posterior probabilities for the cluster ar-
gument. As in this case the first and third component
are restricted to have the same coefficient for lgRD,
the posteriors of the fitted mixture are used for ini-
tialization after reordering the components to have
these two components next to each other. The mod-
ified model is compared to the original model using
the BIC.
R> Model_2 <- FLXMRglmfix(family = "poisson",
+ nested = list(k = c(1,2),
+ formula = ~lgRD))
R> Post_1 <- posterior(pat_mix)[,c(2,1,3)]
R> pat_mix2 <- flexmix(Patents ~ 1,
+ concomitant = FLXPmultinom(~RDS),
+ data = patent, cluster = Post_1,
+ model = Model_2)
R> c(M_1 = BIC(pat_mix), M_2 = BIC(pat_mix2))
M_1 M_2
437.836 445.243
In this example, the original model is preferred
by the BIC.
Summary
flexmix provides infrastructure for fitting finite mix-
ture models with the EM algorithm and tools for
model selection and model diagnostics. We have
shown the application of the package for model-
based clustering as well as for fitting finite mixtures
of regressions.
In the future we want to implement new model
drivers, e.g., for generalized additive models with
smooth terms, as well as to extend the tools for
model selection, diagnostics and model validation.
Additional functionality will be added which allows
to fit mixture models with different component spe-
cific models. The implementation of zero-inflated
models has been a first step in this direction.
Acknowledgments
This research was supported by the Austrian Science
Foundation (FWF) under grant P17382.
Bibliography
A. Dasgupta and A. E. Raftery. Detecting features
in spatial point processes with clutter via model-
based clustering. Journal of the American Statistical
Association, 93(441):294–302, March 1998. 9
R News ISSN 1609-3631

Vol. 7/1, April 2007 13
R> refit(pat_mix)
Call:
refit(pat_mix)
Number of components: 3
$Comp.1
Estimate Std. Error z value Pr(>|z|)
(Intercept) 0.50819 0.12366 4.11 3.96e-05 ***
lgRD 0.87976 0.03328 26.43 < 2e-16 ***
---
Signif. codes: 0 *** 0.001 ** 0.01 * 0.05 . 0.1 1
$Comp.2
Estimate Std. Error z value Pr(>|z|)
(Intercept) -2.6368 0.4043 -6.522 6.95e-11 ***
lgRD 1.5866 0.0899 17.648 < 2e-16 ***
---
Signif. codes: 0 *** 0.001 ** 0.01 * 0.05 . 0.1 1
$Comp.3
Estimate Std. Error z value Pr(>|z|)
(Intercept) 1.96217 0.13968 14.05 <2e-16 ***
lgRD 0.67190 0.03572 18.81 <2e-16 ***
---
Signif. codes: 0 *** 0.001 ** 0.01 * 0.05 . 0.1 1
Figure 6: Details of the fitted mixture model for the patent data.
C. M. Dayton and G. B. Macready. Concomitant-
variable latent-class models. Journal of the American
Statistical Association, 83(401):173–178, March 1988.
9
A. P. Dempster, N. M. Laird, and D. B. Rubin. Maxi-
mum likelihood from incomplete data via the EM-
algorithm. Journal of the Royal Statistical Society B,
39:1–38, 1977. 9
Y. D. Edwards and G. M. Allenby. Multivariate anal-
ysis of multiple response data. Journal of Marketing
Research, 40:321–334, August 2003. 10
C. Fraley and A. E. Raftery. How many clusters?
which clustering method? answers via model-
based cluster analysis. The Computer Journal, 41(8):
578–588, 1998. 10
C. Fraley and A. E. Raftery. MCLUST version 3 for R:
Normal mixture modeling and model-based clus-
tering. Technical Report 504, University of Wash-
ington, Department of Statistics, September 2006.
9
S. Frühwirth-Schnatter. Finite Mixture and Markov
Switching Models. Springer Series in Statistics.
Springer, New York, 2006. ISBN 0-387-32909-9. 8,
9
B. Grün and F. Leisch. Fitting finite mixtures of
generalized linear regressions in R.Computational
Statistics & Data Analysis, 2007a. Accepted for pub-
lication. 9
B. Grün and F. Leisch. Flexmix 2.0: Finite mixtures
with concomitant variables and varying and fixed
effects. Submitted for publication, 2007b. 9
F. Leisch. FlexMix: A general framework for fi-
nite mixture models and latent class regression in
R. Journal of Statistical Software, 11(8), 2004. URL
http://www.jstatsoft.org/v11/i08/.9,12
G. J. McLachlan and D. Peel. Finite Mixture Models.
Wiley, 2000. 8,9
D. Sarkar. Lattice. R News, 2(2):19–23, June 2002. URL
http://CRAN.R-project.org/doc/Rnews/.12
P. Wang, I. M. Cockburn, and M. L. Puterman. Anal-
ysis of patent data — A mixed-Poisson-regression-
model approach. Journal of Business & Economic
Statistics, 16(1):27–41, 1998. 11
M. Wedel and W. A. Kamakura. Market Segmen-
tation — Conceptual and Methodological Founda-
tions. Kluwer Academic Publishers, second edi-
tion, 2001. ISBN 0-7923-8635-3. 8
Bettina Grün
Technische Universität Wien, Austria
Bettina.Gruen@ci.tuwien.ac.at
Friedrich Leisch
Ludwig-Maximilians-Universität München, Germany
Friedrich.Leisch@R-project.org
R News ISSN 1609-3631

Vol. 7/1, April 2007 14
Using R to Perform the AMMI Analysis
on Agriculture Variety Trials
by Andrea Onofri & Egidio Ciriciofolo
Introduction
Field agricultural experiments are generally planned
to evaluate the actual effect produced by man-
made chemical substances or human activities on
crop yield and quality, environmental health, farm-
ers’ income and so on. Field experiments include
the testing of new and traditional varieties (geno-
types), fertilizers (types and doses), pesticides (types
and doses) and cultural practices. With respect to
greenhouse or laboratory experiments, field trials
are much more strongly subjected to environmental
variability that forces researchers into repeating ex-
periments across seasons and/or locations. A sig-
nificant ’treatment x environment’ interaction may
introduce some difficulties in exploring the dataset,
summarizing results and determining which treat-
ment (genotype, herbicide, pesticide...) was the best.
In such conditions, the AMMI (Additive Main ef-
fect Multiplicative Interaction) analysis has been pro-
posed as an aid to visualize the dataset and explore
graphically its pattern and structure (Gollob,1968;
Zobel et al.,1988); this technique has received a par-
ticular attention from plant breeders (see for example
Abamu and Alluri,1998;Annichiarico et al.,1995;
Annichiarico,1997;Ariyo,1998) and recently it has
been stated as superior to other similar techniques,
such as the GGE (Gauch,2006). Unfortunately, such
technique is not yet very well exploited by agri-
cultural scientists, who often prefer a more tradi-
tional approach to data analysis, based on ANOVA
and multiple comparison testing. Without disre-
garding the importance of such an approach, one
cannot deny that sometimes this does not help un-
veil the underlying structure of experimental data,
which may be more important than hypothesis test-
ing, especially at the beginning of data analyses (Ex-
ploratory Data Analysis; NIST/SEMATECH,2004)
or at the very end, when graphs have to be drawn
for publication purposes.
To make more widespread the acceptance and the
use of such powerful tool within agronomists, it is
necessary to increase the availability of both practi-
cal information on how to perform and interpret an
AMMI analysis and simple software tools that give
an easily understandable output, aimed at people
with no specific and deep statistical training, such as
students and field technicians.
The aim of this paper was to show how R can be
easily used to perform an AMMI analysis and pro-
duce ’biplots’, as well as to show how these tools can
be very useful within variety trials in agriculture.
Some basic statistical aspects
The AMMI analysis combines the ANalysis OF VAri-
ance (ANOVA) and the Singular Value Decomposi-
tion (SVD) and it has been explained in detail by Gol-
lob (1968). If we specifically refer to a variety trial,
aimed at comparing the yield of several genotypes
in several environments (years and/or locations), the
ANOVA partitions the total sum of squares into two
main effects (genotypes and environments) plus the
interaction effect (genotypes x environments). This
latter effect may be obtained by taking the observed
averages for each ’genotype x environment’ combi-
nation and doubly-centering them (i.e., subtracting
to each data the appropriate genotype and environ-
ment means and adding back the grand mean). The
interaction effect is arranged on a two-way matrix γ
(one row for each genotype and one column for each
environment) and submitted to SVD, as follows:
γ=
r
∑
i=1
λi·gik ·ei j (1)
where ris the rank of γ,λiis the singular value
for principal component i,gik is the eigenvector score
for genotype kand Principal Component (PC) i(left
singular vector), while ei j is the eigenvector score for
environment jand PC i(right singular vector). If PC
scores are multiplied by the square root of the singu-
lar value, equation 1is transformed into:
γ=
r
∑
i=1λ0.5
i·gik λ0.5
i·ei j=
r
∑
i=1
Gik ·Ei j (2)
In this way the additive interaction in the
ANOVA model is obtained by multiplication of
genotype PC scores by environment PC scores, ap-
propriately scaled. If a reduced number of PCs is
used (r= 1 or 2, typically) a dimensionality reduction
is achieved with just a small loss in the descriptive
ability of the model. This makes it possible to plot
the interaction effect, via the PC scores for genotypes
and environments. Such graphs are called biplots, as
they contain two kinds of data; typically, a AMMI1
and a AMMI2 biplots are used: the AMMI1 biplot
has main effects (average yields for genotypes and
environments) on the x-axis and PC1 scores on the y-
axis, while the AMMI2 biplot has PC1 scores on the
x-axis and PC2 scores on the y-axis.
R News ISSN 1609-3631

Vol. 7/1, April 2007 15
Table 1: Field averages (three replicates) for six genotypes compared in seven years.
Genotype 1996 1997 1998 1999 2000 2001 2002 Average
COLOSSEO 6.35 6.46 6.70 6.98 6.44 7.07 4.90 6.41
CRESO 5.60 6.09 6.13 7.13 6.08 6.45 4.33 5.97
DUILIO 5.64 8.06 7.15 7.99 5.18 7.88 4.24 6.59
GRAZIA 6.26 6.74 6.35 6.84 4.75 7.30 4.34 6.08
IRIDE 6.04 7.72 6.39 7.99 6.05 7.71 4.96 6.70
SANCARLO 5.70 6.77 6.81 7.41 5.66 6.67 4.50 6.22
SIMETO 5.08 7.19 6.44 7.07 4.82 7.55 3.34 5.93
SOLEX 6.14 6.39 6.44 6.87 5.45 7.52 4.79 6.23
Average 5.85 6.93 6.55 7.29 5.55 7.27 4.42 6.27
The dataset
To show how the AMMI analysis can be easily per-
formed with R, we will use a dataset obtained from
a seven-years field trial on durum wheat, carried out
from 1996 to 2003 in central Italy, on a randomised
block design with three replicates. For the present
analysis, eight genotypes were chosen, as they were
constantly present throughout the years (Colosseo,
Creso, Duilio, Grazia, Iride, Sancarlo, Simeto, Solex).
Yield data referred to the standard humidity content
of 13% (Tab. 1) have been previously published in
Belocchi et al. (2003), Ciriciofolo et al. (2002); Ciri-
ciofolo et al. (2001); Desiderio et al. (2000), Desiderio
et al. (1999), Desiderio et al. (1998), Desiderio et al.
(1997). The interaction matrix (which is submitted to
SVD) is given in table 2.
The AMMI with R
To perform the AMMI analysis, an R function was
defined, as shown on page 17.
The AMMI() function requires as inputs a vector
of genotype codes (factor), a vector of environment
codes (factor), a vector of block codes (factor) and a
vector of yields (numerical). PC is the number of PCs
to be considered (set to 2 by default) and biplot is the
type of biplot to be drown (1 for AMM1 and 2 for
AMMI2). It should be noted that the script is very el-
ementary and that it does not use any difficult func-
tion or programming construct. It was simply coded
by translating the algebraic procedure proposed by
Gollob (1968) into R statements, which is a very easy
task, even without a specific programming training.
Wherever possible, built-in R functions were used, to
simplify the coding process and to facilitate the adap-
tation of the script to other kinds of AMMI models.
The first part uses the function tapply() to cal-
culate some descriptive statistics, such as genotype
means, environment means and ’genotype x envi-
ronment’ means, which are all included in the final
output.
The second part uses the function aov() to per-
form the ANOVA by using a randomised block de-
sign repeated in different environments with a dif-
ferent randomisation in each environment (LeClerg
et al.,1962). The interaction matrix γis calculated
by using the function model.tables() applied to the
output of the function aov(); the way the R script
is coded, the interaction matrix is actually the trans-
pose of the matrix shown in table 2, but this does not
change much in terms of the results. The interaction
matrix is then submitted to SVD, by using the built-
in R function svd().
The significant PCs are assessed by a series of F
tests as shown by Zobel et al. (1988) and PC scores,
genotype means and environment means are used to
produce biplots, by way of the functions plot() and
points().
Table 2: Interaction effects for the dataset in table 1.
Genotype 1996 1997 1998 1999 2000 2001 2002
COLOSSEO 0.35 -0.62 0.00 -0.45 0.74 -0.34 0.33
CRESO 0.04 -0.54 -0.13 0.14 0.82 -0.52 0.20
DUILIO -0.54 0.80 0.28 0.38 -0.70 0.29 -0.51
GRAZIA 0.60 -0.01 -0.02 -0.27 -0.62 0.21 0.10
IRIDE -0.24 0.37 -0.59 0.28 0.07 0.01 0.11
SANCARLO -0.10 -0.11 0.31 0.17 0.16 -0.55 0.12
SIMETO -0.43 0.60 0.23 0.13 -0.40 0.62 -0.75
SOLEX 0.33 -0.50 -0.07 -0.38 -0.07 0.29 0.40
R News ISSN 1609-3631

Vol. 7/1, April 2007 16
$Additive_ANOVA
Df Sum Sq Mean Sq F value Pr(>F)
Environments 6 159.279 26.547 178.3996 < 2.2e-16 ***
Genotypes 7 11.544 1.649 11.0824 2.978e-10 ***
Blocks(Environments) 14 3.922 0.280 1.8826 0.03738 *
Environments x Genotypes 42 27.713 0.660 4.4342 6.779e-10 ***
Residuals 98 14.583 0.149
---
Signif. codes: 0 *** 0.001 ** 0.01 * 0.05 . 0.1 1
$Mult_Interaction
Effect SS DF MS F Prob.
1 PC1 18.398624 12 1.5332187 10.303612 7.958058e-13
2 PC2 5.475627 10 0.5475627 3.679758 3.339881e-04
3 PC3 1.961049 8 0.2451311 1.647342 1.212529e-01
4 Residuals 1.877427 12 0.1564522 1.051398 4.094193e-01
$Environment_scores
PC1 PC2 PC3
1996 0.4685599 -0.62599974 0.01665148
1997 -0.8859669 0.21085535 -0.19553672
1998 -0.1572887 -0.00567589 0.80162642
1999 -0.3139136 0.51881710 -0.13286326
2000 0.8229290 0.59868592 -0.03330554
2001 -0.5456613 -0.49726356 -0.18138908
2002 0.6113417 -0.19941917 -0.27518331
$Genotype_scores
PC1 PC2 PC3
COLOSSEO 0.74335025 -0.02451524 0.1651197989
CRESO 0.63115567 0.47768803 -0.0001969871
DUILIO -0.87632103 0.17923645 0.1445152042
GRAZIA -0.07625519 -0.74659598 -0.0108977060
IRIDE -0.12683903 0.28634343 -0.7627600696
SANCARLO 0.18186612 0.35076556 0.3753706117
SIMETO -0.78109997 0.04751457 0.1740113396
SOLEX 0.30414317 -0.57043681 -0.0851621918
Figure 1: Results from ANOVA and AMMI analyses.
Results
Results (Fig. 1) show a highly significant ’genotypes
x environments’ interaction (GE) on the ANOVA,
that does not permit to define an overall ranking of
varieties across environments.
The SVD decomposition of the interaction matrix
was performed by extracting three PCs, though only
the first two are significant. It is possible to observe
that the first PC accounts for 66% of the interaction
sum of squares, while the second one accounts for an
additional 20%.
The AMMI1 biplot shows contemporarily main
effects (genotypes and environments average yields)
and interaction, as PC1 scores (Fig. 2). This graph is
relevant as it accounts for 87% of total data variabil-
ity. Figure 2: AMMI1 biplot.
R News ISSN 1609-3631

Vol. 7/1, April 2007 17
AMMI <- function(variety, envir, block, yield, PC = 2, biplot = 1) {
## 1 - Descriptive statistics
overall.mean <- mean(yield)
envir.mean <- tapply(yield, envir, mean)
var.mean <- tapply(yield, variety, mean)
int.mean <- tapply(yield, list(variety,envir), mean)
envir.num <- length(envir.mean)
var.num <- length(var.mean)
## 2 - ANOVA (additive model)
variety <- factor(variety)
envir <- factor(envir)
block <- factor(block)
add.anova <- aov(yield ~ envir + block %in% envir + variety + envir:variety)
modelTables <- model.tables(add.anova, type = "effects", cterms = "envir:variety")
int.eff <- modelTables$tables$"envir:variety"
add.anova.residual.SS <- deviance(add.anova)
add.anova.residual.DF <- add.anova$df.residual
add.anova.residual.MS <- add.anova.residual.SS/add.anova.residual.DF
anova.table <- summary(add.anova)
row.names(anova.table[[1]]) <- c("Environments", "Genotypes", "Blocks(Environments)",
"Environments x Genotypes", "Residuals")
## 3 - SVD
dec <- svd(int.eff, nu = PC, nv = PC)
if (PC > 1) {
D <- diag(dec$d[1:PC])
} else {
D <- dec$d[1:PC]
}
E <- dec$u %*% sqrt(D)
G <- dec$v %*% sqrt(D)
Ecolnumb <- c(1:PC)
Ecolnames <- paste("PC", Ecolnumb, sep = "")
dimnames(E) <- list(levels(envir), Ecolnames)
dimnames(G) <- list(levels(variety), Ecolnames)
## 4 - Significance of PCs
numblock <- length(levels(block))
int.SS <- (t(as.vector(int.eff)) %*% as.vector(int.eff))*numblock
PC.SS <- (dec$d[1:PC]^2)*numblock
PC.DF <- var.num + envir.num - 1 - 2*Ecolnumb
residual.SS <- int.SS - sum(PC.SS)
residual.DF <- ((var.num - 1)*(envir.num - 1)) - sum(PC.DF)
PC.SS[PC + 1] <- residual.SS
PC.DF[PC + 1] <- residual.DF
MS <- PC.SS/PC.DF
F <- MS/add.anova.residual.MS
probab <- pf(F, PC.DF, add.anova.residual.DF, lower.tail = FALSE)
percSS <- PC.SS/int.SS
rowlab <- c(Ecolnames, "Residuals")
mult.anova <- data.frame(Effect = rowlab, SS = PC.SS, DF = PC.DF, MS = MS, F = F, Prob. = probab)
## 5 - Biplots
if (biplot == 1) {
plot(1, type = n , xlim = range(c(envir.mean, var.mean)), ylim = range(c(E[,1], G[,1])), xlab = "Yield",
ylab = "PC 1")
points(envir.mean, E[,1], col = "red", lwd = 5)
plot(1, type = n , xlim = range(c(envir.mean, var.mean)), ylim = range(c(E[,1], G[,1])), xlab = "Yield",
ylab = "PC 1")
points(envir.mean, E[,1], "n", col = "red", lwd = 5)
text(envir.mean, E[,1], labels = row.names(envir.mean), adj = c(0.5, 0.5), col = "red")
points(var.mean, G[,1], "n", col = "blue", lwd = 5)
text(var.mean, G[,1], labels = row.names(var.mean), adj = c(0.5, 0.5), col = "blue")
abline(h = 0, v = overall.mean, lty = 5)
} else {
plot(1, type = n , xlim = range(c(E[,1], G[,1])), ylim = range(c(E[,2], G[,2])), xlab = "PC 1",
ylab = "PC 2")
points(E[,1], E[,2], "n",col = "red", lwd = 5)
text(E[,1], E[,2], labels = row.names(E),adj = c(0.5,0.5),col = "red")
points(G[,1],G[,2], "n", col = "blue", lwd = 5)
text(G[,1], G[,2], labels = row.names(G),adj = c(0.5, 0.5), col = "blue")
}
## 6 - Other results
list(Genotype_means = var.mean, Environment_means = envir.mean, Interaction_means = int.mean,
Additive_ANOVA = anova.table, Mult_Interaction = mult.anova, Environment_scores = E,
Genotype_scores = G)
}
R News ISSN 1609-3631
Vol. 7/1, April 2007 18
To read this biplot, it is necessary to remember
that genotypes and environments on the right side
of the graph shows yield levels above the average.
Besides, genotypes and environments laying close
to the x-axis (PC 1 score close to 0) did not interact
with each other, while data with positive/negative
score on y-axis interacted positively with environ-
ments characterised by a score of same sign.
Indeed, environmental variability was much
higher than genotype variability (Fig. 2, see also the
ANOVA in Fig. 1). Iride showed the highest aver-
age yield and did not interact much with the envi-
ronment (PC1 score close to 0). Duilo ranked over-
all second, but showed a high interaction with the
environment, i.e., its yield was above the average
in 1997 (first ranking), 2001 (first ranking) and 1999
(second ranking), while it was below the average in
1996, 2000 and 2002. Colosseo gave also a good av-
erage yield, but its performances were very positive
in 1996, 2000 and 2002, while they were below the
average in 1997, 2000 and 2002.
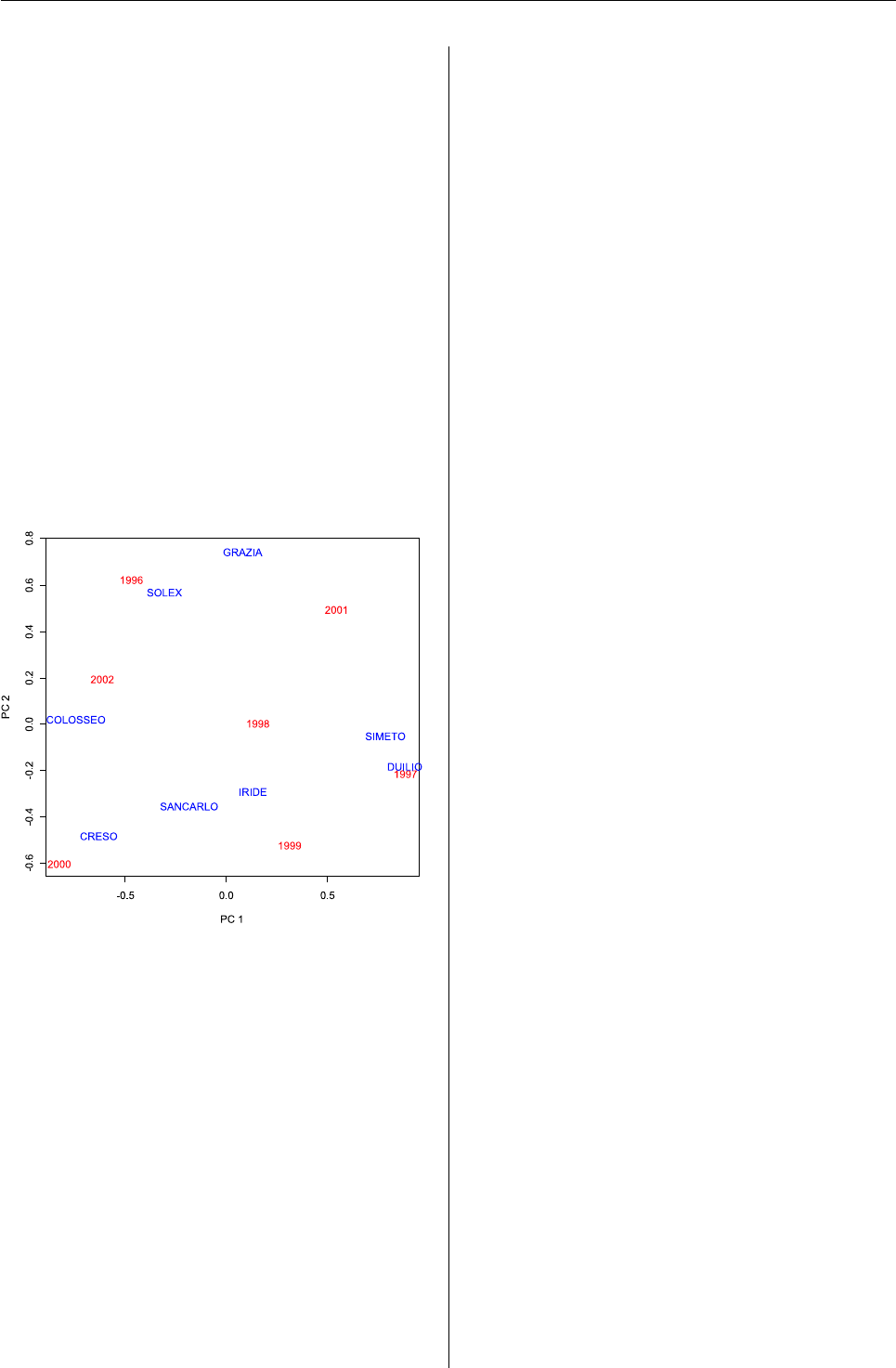
Figure 3: AMMI2 biplot.
The AMMI2 biplot (Fig. 3) is more informative on
the GE interaction as it accounts for 86% of the sum
of squares of this latter effect. Remember that geno-
types and environments in the center of the graph
did not show a relevant interaction, while genotypes
and environment lying close on the external parts of
the graph interacted positively. Duilio and Simeto
were particularly brilliant in 1997 (compared to their
average performances; notice in tab. 1that Simeto
was the third in this year, which is very good com-
pared to its seventh position on the ranking based
on average yield). Solex and Grazia were brilliant
in 1997 (they were third and second respectively, in
spite of the eighth and fifth ranking based on aver-
age yield). Likewise, Creso and Colosseo were the
best in 2000 and 2002, while Iride and Sancarlo inter-
acted positively with 1999.
Discussion and conclusions
The above example should be discussed with refer-
ence to two main aspects: the AMMI analysis and
the use of R. Concerning the first aspect, the example
confirms the graphical power of the biplots; indeed,
all the above comments are just an excerpt of what
can be easily grasped at first sight from the AMMI1
and AMMI2 biplots. It is worth to notice that obtain-
ing such information from table 1is not as immediate
and quick. Of course, the AMMI analysis should be
followed by other procedures to explore the relation-
ship between the behaviour of each variety and the
environmental conditions of each year.
It is also necessary to mention that in the present
example the analyses were aimed only at graphically
exploring the underlying structure of the dataset. In
other cases, whenever hypothesis testing is more im-
portant, the F procedure employed on the script may
be too liberal and other techniques may be better
suited to evaluate the significance of PCs (Cornelius,
1993).
Concerning R, the above example confirms that
this environment can be easily used to perform sta-
tistical analyses in the agricultural field. Thanks to
its ability to deal with linear models and to the fa-
cilities for matrix manipulation, it is very easy to ac-
complish also rather complex statistical tasks, such
as the AMMI analysis. Indeed, calculations can be
performed having in mind the usual algebraic nota-
tion, as one can find in statistical literature, without
a deep knowledge of programming constructs. In-
deed, this script has been coded in an elementary
fashion, following the calculation pattern proposed
by Gollob (1968) and including some built-in R func-
tions when possible.
Of course, it is necessary to mention that this
elementary coding style may be useful for simple
scripts, but should not be regarded as optimal, es-
pecially for more advanced applications. Indeed, in
such cases an object-oriented approach is much more
advisable to exploit the statistical power of R. In any
case, elementary scripts such this one may be always
used as the starting point to perform other types of
statistical analyses. In particular, with slight mod-
ifications, this script (available on: www.unipg.it/
~onofri/software.htm) could be used to draw the
GGE biplot, that has received a certain attention in
the last years (Yan and Tinker,2005).
However, when re-using this script, one should
bear in mind some limitations. Indeed, it is impor-
tant to notice that this script has been aimed at a spe-
cific experimental design (completely randomised
block experiment repeated across years or environ-
ments), as commonly found in field variety trials.
Other designs will require some adaptations into
the code and unbalanced designs (especially those
with missing combinations of genotypes and envi-
ronments) should not be analysed with this script.
R News ISSN 1609-3631

Vol. 7/1, April 2007 19
Furthermore, the ’environment’ effect has been con-
sidered as ’fixed’ and changes to the code should be
made in case it should be considered as ’random’.
In spite of the the above limitations, it is clear that
also users with a limited background in computer
programming (which is often the case in agriculture)
can benefit from the use of R: an elementary knowl-
edge of R statements and functions is already enough
to perform also the ’less traditional’ statistical analy-
sis, with a very slight effort.
Bibliography
F. J. Abamu and K. Alluri. Ammi analysis of rainfed
lowland rice (oryza sativa) trials in nigeria. Plant
Breeding, 117(4):395–397, 1998. 14
P. Annichiarico. Addittive main effects and multi-
plicative interaction (ammi) analysis of genotype-
location interaction in variety trials repeated over
years. Theorethycal applied genetics, 94:1072–1077,
1997. 14
P. Annichiarico, F. Bozzo, G. Parente, F. Gusmeroli,
V. Mair, O. Marguerettaz, and D. Orlandi. Analysis
of adaptation of grass/legume mixtures to italian
alpine and subalpine zones through an addittive
main effects and multiplicative interaction model.
Grass and Forage Science, 50:405–413, 1995. 14
O. J. Ariyo. Use of additive main effects and mul-
tiplicative interaction model to analyse multiloca-
tion soybean varietal trials. J.Genet. Breed, (53):129–
134, 1998. 14
A. Belocchi, M. Fornara, E. Ciriciofolo, E. Bianco-
latte, S. D. Puglia, G. Olimpieri, V. Vecchiarelli,
E. Gosparini, C. Piccioni, and E. Desiderio. Scelta
delle varietà di frumento duro: risultati delle
prove varietali 2002-2003 in Lazio ed Umbria.
L’Informatore Agrario, supplemento 1, 2003. 15
E. Ciriciofolo, A. Belocchi, E. Biancolatte, S. D.
Puglia, M. Fornara, G. Olimpieri, C. Piccioni, and
E. Desiderio. Scelta delle varietà di frumento duro:
risultati delle prove varietali 2000-2001 in Lazio ed
Umbria. L’Informatore Agrario, supplemento 1, 2001.
15
E. Ciriciofolo, A. Belocchi, E. Biancolatte, S. D.
Puglia, M. Fornara, G. Olimpieri, C. Piccioni, and
E. Desiderio. Scelta delle varietà di frumento duro:
risultati delle prove varietali 2001-2002 in Lazio ed
Umbria. L’Informatore Agrario, supplemento 1, 2002.
15
P. L. Cornelius. Statistical tests and retention of terms
in the Addittive Main Effects and Multiplicative
interaction model for cultivar trials. Crop Science,
33:1186–1193, 1993. 18
E. Desiderio, A. Belocchi, E. Biancolatte, E. Ciricio-
folo, S. D. Puglia, G. Olimpieri, C. Piccioni, and
M. Fornara. Scelta delle varietà di frumento duro:
risultati delle prove varietali 1996-1997 in Lazio ed
Umbria. L’Informatore Agrario, supplemento 1, 1997.
15
E. Desiderio, E. Ciriciofolo, A. Belocchi, E. Bian-
colatte, S. D. Puglia, G. Olimpieri, C. Piccioni,
M. Fornara, and M. Magini. Scelta delle varietà di
frumento duro: risultati delle prove varietali 1997-
1998 in Lazio ed Umbria. L’Informatore Agrario,
supplemento 1, 1998. 15
E. Desiderio, E. Ciriciofolo, A. Belocchi, E. Bian-
colatte, S. D. Puglia, G. Olimpieri, C. Piccioni,
M. Fornara, and M. Magini. Scelta delle varietà di
frumento duro: risultati delle prove varietali 1998-
1999 in Lazio ed Umbria. L’Informatore Agrario,
supplemento 1, 1999. 15
E. Desiderio, E. Biancolatte, E. Ciriciofolo, S. D.
Puglia, M. Fornara, G. Olimpieri, C. Piccioni, and
L. Frongia. Scelta delle varietà di frumento duro:
risultati delle prove varietali 1999-2000 in Lazio ed
Umbria. L’Informatore Agrario, supplemento 1, 35:
52–55, 2000. 15
H. F. Gollob. A statistical model which combines
features of factor analytic and analysis of variance
techniques. Psychometrika, 33(1):73–114, 1968. 14,
15,18
H. G. Gauch. Statistical analysis of yield trials by
AMMI and GGE. Crop Science, 46(4):1488–1500,
2006. 14
E. L. LeClerg, W. H. Leonard, and A. G. Clark. Field
Plot Technique. Burgess Publishing Company, Min-
neapolis, Minnesota, 1962. 15
NIST/SEMATECH. E-Handbook of Statistical Meth-
ods. 1: Exploratory Data Analysis.http://www.itl.
nist.gov/div898/handbook/, 2004. 14
W. Yan and N. A. Tinker. An integrated biplot analy-
sis system for displaying, interpreting, and explor-
ing genotype ×environment interaction. Crop Sci-
ence, 45(3):1004–1016, 2005. 18
R. W. Zobel, M. J. Wright, and H. G. Gauch. Statis-
tical analysis of a yield trial. Agronomy Journal, 80:
388–393, 1988. 14,15
Andrea Onofri & Egidio Ciriciofolo
Department of Agricultural and Environmental Science
University of Perugia
onofri@unipg.it
egicir@unipg.it
R News ISSN 1609-3631

Vol. 7/1, April 2007 20
Inferences for Ratios of Normal Means
by Gemechis Dilba, Frank Schaarschmidt, Ludwig A.
Hothorn
Introduction
Inferences concerning ratios of means of normally
distributed random variables or ratios of regression
coefficients arise in a variety of problems in biomedi-
cal research. For example, in tests for non-inferiority
of one or more experimental treatments against a
positive control, it is often easier to define and also
to interpret the non-inferiority margin as percent-
age changes (or fraction retained compared to the
mean of the control group). In bioassay problems,
one is also interested in ratios of regression coef-
ficients, for instance in parallel line or slope ratio
assays. Our aim here is to introduce an R exten-
sion package called mratios which can perform infer-
ences about one or more such ratio parameters in the
general linear model. For two-sample problems, the
package is capable of constructing Fieller confidence
intervals and performing the related tests when the
group variances are assumed homogeneous or het-
erogeneous. In simultaneous inferences for multiple
ratios, the package can (i) perform multiple tests, (ii)
construct simultaneous confidence intervals using a
variety of techniques, and (iii) calculate the sample
sizes required for many-to-one comparisons in si-
multaneous tests for non-inferiority (or superiority)
based on relative margins. We demonstrate the func-
tionality of the package by using several data exam-
ples.
Two-sample Problem
The two-sample problem is one of the standard
methods routinely used in practice. Here the interest
is in comparing the means of two independent nor-
mally distributed random variables in terms of the
ratio of their means. This can be accomplished by
using the t.test.ratio function. If the variances are
homogeneous, this function performs a ratio format-
ted t-test (also known as Sasabuchi test) and com-
putes Fieller’s confidence interval. If variance homo-
geneity is not tenable (the default), the test proposed
by Tamhane and Logan (2004) is performed using
Satterthwaite adjusted degrees of freedom. For con-
fidence interval estimation under variance hetero-
geneity, Satterthwaite degrees of freedom depends
on the unknown ratio. To circumvent this problem,
we plug in the maximum likelihood estimate of the
ratio (i.e., ratio of sample means) in the approximate
expression for the number of degrees of freedom.
Example 1. Consider the mutagenicity assay data
described in the mratios package. A first step in
the analysis of the data could be to test whether the
active control (cyclophosphamide at dose 25mg/kg)
results in a significantly higher number of mutations
than the vehicle control. The data appear to be het-
eroscedastic, and therefore we use the unequal vari-
ances option (the default) to compare the two treat-
ments.
> library("mratios")
> data("Mutagenicity")
> muta2 <- subset(Mutagenicity, Treatment ==
+ "Vehicle" | Treatment == "Cyclo25")
> t.test.ratio(MN ~ Treatment, data = muta2,
+ alternative = "greater")
Ratio t-test for unequal variances
data: Cyclo25 and Vehicle
t = 5.0071, df = 3.07, p-value = 0.0073
alternative hypothesis: true ratio of means
is greater than 1
95 percent confidence interval:
5.110079 Inf
sample estimates:
mean Cyclo25 mean Vehicle
25.000000 2.571429
Cyclo25/Vehicle
9.722222
Note that when testing a ratio of means against
1, the p-value computed by the t.test.ratio func-
tion is exactly the same as that computed by t.test
when testing the difference of means against 0.
Simultaneous Inferences
In this section we consider inferential problems in-
volving one or more ratio parameters. The basic dis-
tribution underlying the analyses is the multivariate
t-distribution. Under the assumption of normality
and homogeneous variance for the error terms, the
joint distribution of the test statistics associated with
the various contrasts of interest follows a multivari-
ate t-distribution. For the computation of the related
multivariate tprobabilities and equi-coordinate crit-
ical points, we refer to Hothorn et al. (2001).
Multiple Tests
Assume a normal one-way ANOVA model with ho-
mogeneous variances. The interest is in simultane-
ous tests for several ratios of linear combinations of
the treatment means. Such tests for ratio hypothe-
ses (ratios of normal means) appear, for example,
in tests for non-inferiority (or superiority) of sev-
eral experimental treatments compared to a control
R News ISSN 1609-3631

Vol. 7/1, April 2007 21
(placebo). These are so called many-to-one compar-
isons. In the R-function simtest.ratio, most of the
routinely used multiple comparison procedures [e.g.,
many-to-one (Dunnett type), all pairs (Tukey type),
sequence (successive comparisons of ordered treat-
ment effects)] are implemented in the context of ratio
hypotheses. In general, the function also allows for
any user-defined contrast matrices.
Let γj=c0
jµ/d0
jµ,j=1, . . . , rdenote the ra-
tios of interest, where µ= (µ1,...,µk)0is a vector
of the treatment means, cjand djare known vectors
of real constants each of dimension k×1, and ris the
number of ratios. To specify the ratios, we define two
contrast matrices, namely, numerator and denomina-
tor contrast matrices. The numerator contrast matrix
is a matrix whose row vectors are c0
1,...,c0
r, and the
denominator contrast matrix is a matrix whose row
vectors are d0
1,...,d0
r. Therefore, the dimensions of
both the numerator and denominator contrast ma-
trices are each r×k. Further, let (ψ1,...,ψr)0de-
note the set of margins against which we test the r
ratios. Then, for example, for one-sided upper-tailed
alternative hypotheses, the hypotheses of interest are
H0j:γj≤ψjversus H1j:γj>ψj,j=1, . . . , r.
Given a data frame containing the observa-
tions, the contrast matrices, the vector of margins,
and the family-wise type I error rate, the func-
tion simtest.ratio calculates the point estimates of
the ratios, the test statistics, the raw p-values and
the multiplicity adjusted p-values. The adjusted p-
values are computed by adapting the results of West-
fall et al. (1999) for ratio hypotheses and general con-
trasts.
In general, note that the function simtest.ratio
allows for varying margins for the set of compar-
isons. This can be quite appealing, for example, in
test problems involving a mixture of non-inferiority
and superiority hypotheses.
Example 2. Bauer et al. (1998) analyzed data
from a multi-dose experiment including a positive
control and placebo. In the experiment, patients
with chronic stable angina pectoris were random-
ized to five treatment arms (placebo, three doses of
a new compound, and an active control). The re-
sponse variable is the difference in the duration of
an exercise test before and after treatment. Now,
due to the unavailability of the original data val-
ues, we randomly generated independent samples
(from a normal distribution) that satisfy the sum-
mary statistics given in Table II of Bauer et al.
(1998). This data set is available in the mratios pack-
age. The interest is in simultaneous tests for non-
inferiority of the three doses versus the active con-
trol by including the placebo. Following Pigeot et
al. (2003), the hypotheses can succinctly be formu-
lated as H0i:(µj−µ2)/(µ1−µ2)≤0.9 versus
H1i:(µj−µ2)/(µ1−µ2)>0.9, j=3, 4, 5, where
µi,i=1, ..., 5 denote the means for the active con-
trol, placebo, dose 50, dose 100 and dose 150, consec-
utively. In this example, the non-inferiority margins
are all set to 0.9.
> data("AP")
> NC <- rbind(N1 = c(0, -1, 1, 0, 0),
+ N2 = c(0, -1, 0, 1, 0),
+ N3 = c(0, -1, 0, 0, 1))
> DC <- rbind(D1 = c(1, -1, 0, 0, 0),
+ D2 = c(1, -1, 0, 0, 0),
+ D3 = c(1, -1, 0, 0, 0))
> ap.test <- simtest.ratio(pre_post ~
+ treatment, data = AP, Num.Contrast = NC,
+ Den.Contrast = DC, Margin.vec = 0.9,
+ alternative = "greater")
> ap.test
Alternative hypotheses: Ratios greater
than margins
margin estimate statistic
N1/D1 0.9 5.306 2.9812
N2/D2 0.9 4.878 2.7152
N3/D3 0.9 1.969 0.7236
p.value.raw p.value.adj
N1/D1 0.001554 0.004429
N2/D2 0.003505 0.009799
N3/D3 0.234952 0.451045
By using the command summary(ap.test), one
can get further information — for example, the corre-
lation matrix under the null hypotheses and the crit-
ical point (equi-coordinate percentage point of the
multivariate t-distribution).
Simultaneous Confidence Intervals
Unlike in multiple testing, in simultaneous estima-
tion of the ratios γj=c0
jµ/d0
jµ,j=1, . . . , r, the joint
distribution of the associated t-statistics follows a
multivariate t-distribution with a correlation matrix
that depends on the unknown ratios. This means that
the critical points that are required for confidence in-
terval construction depend on these unknown pa-
rameters. There are various methods of dealing with
this problem. They are (i) using the unadjusted in-
tervals (Fieller confidence intervals without multi-
plicity adjustments); (ii) Bonferroni (Fieller intervals
with simple Bonferroni adjustments); (iii) a method
called MtI which consists of replacing the unknown
correlation matrix of the multivariate t-distribution
by an identity matrix of the same dimension accord-
ing to Sidak and Slepian inequalities (Hochberg and
Tamhane,1987) for two- and one-sided confidence
intervals, respectively; and (iv) plug-in (plugging the
maximum likelihood estimates of the ratios into the
unknown correlation matrix). The latter method is
known to have good simultaneous coverage proba-
bilities and hence it is set as a default method in the
R functions to be introduced. For details regarding
these methodologies, we refer to Dilba et al. (2006a).
R News ISSN 1609-3631

Vol. 7/1, April 2007 22
The sci.ratio function is used to construct si-
multaneous CIs for ratios of linear combinations of
treatment means in a one-way ANOVA model. Sev-
eral standard contrast types (e.g., Dunnett, Tukey, se-
quence, and many others) are implemented in this
function. The default contrast is many-to-one com-
parisons (Dunnett type) with the mean of the first
level of the factor (in alpha-numeric order) taken as
the denominator of the ratios. In addition, this func-
tion has an option for user-defined contrast matrices.
Example 3. Recall the data from the multi-dose
experiment in Example 2 above. Now, suppose that
the interest is to calculate simultaneous lower 95%
confidence limits for the ratios of the three doses and
the active control to the placebo. Noting that placebo
is the second level in the alpha-numeric order of the
treatments, we use the following R code to calculate
the limits.
> ap.sci <- sci.ratio(pre_post ~
+ treatment, data = AP, type = "Dunnett",
+ base = 2, alternative = "greater",
+ method = "MtI")
The graph of the confidence intervals can be obtained
by applying the plot function to the object in which
the confidence interval estimates are stored, see Fig-
ure 1.
> plot(ap.sci)
●
●
●
●
Lower 95 % simultaneous CI for ratios (method: Slepian)
AC/D0
D100/D0
D150/D0
D50/D0
0.5 1.0 1.5 2.0
|
|
|
|
Figure 1: Graphical visualization of the ap.sci ob-
ject.
The sci.ratio.gen function is a more general
function that can construct simultaneous confidence
intervals for ratios of linear combinations of coeffi-
cients in the general linear model. For this function,
it is necessary to specify the vector of responses, the
design matrix, and the numerator and denominator
contrast matrices.
Example 4. Consider the problem of simulta-
neously estimating relative potencies in a multiple
slope ratio assay. Jensen (1989) describes an exper-
iment in which three preparations are compared to
a control. The response variable (Y) is pantothenic
acid content of plant tissues. The model is Yi j =
α+βiXi j +i j,i=0, 1, 2, 3; j=1, . . . , ni, where
the Xi js are the dose levels and i=0 refers to the
control group. The vector of regression coefficients
is (α,β0,β1,β2,β3)0. Now using the data in Table 5
of Jensen (1989), the interest is to construct simul-
taneous CIs for βi/β0,i=1, 2, 3. The function
sci.ratio.gen needs the response vector Yand the
design matrix X as an input.
> data(SRAssay)
> Y <- SRAssay[, "Response"]
> X <- model.matrix(Response ~ Treatment:Dose,
+ data = SRAssay)
> NC <- matrix(c(0, 0, 1, 0, 0,
+ 0, 0, 0, 1, 0,
+ 0, 0, 0, 0, 1),
+ nrow = 3, byrow = TRUE)
> DC <- matrix(c(0, 1, 0, 0, 0,
+ 0, 1, 0, 0, 0,
+ 0, 1, 0, 0, 0),
+ nrow = 3, byrow = TRUE)
> s.ratio <- sci.ratio.gen(Y, X,
+ Num.Contrast = NC, Den.Contrast = DC)
> s.ratio
Two-sided 95 % simultaneous confidence
intervals for ratios:
estimate lower upper
C1 1.1217 1.0526 1.1964
C2 0.7193 0.6603 0.7805
C3 0.7537 0.6942 0.8157
Using the command summary(s.ratio), one can
also get further details regarding the fitted regres-
sion model, the contrast matrices and an estimate of
the correlation matrix (when the plug-in method is
used). The estimate of the correlation matrix used
for critical point calculation can also be obtained as
> s.ratio[["CorrMat.est"]]
[,1] [,2] [,3]
[1,] 1.0000000 0.4083451 0.4260802
[2,] 0.4083451 1.0000000 0.3767098
[3,] 0.4260802 0.3767098 1.0000000
Note that by choosing the option for method
as ’Sidak’, one gets the results reported by Jensen
(1989).
Before closing this section, we give two important
remarks.
i) According to the Slepian inequality (Hochberg
and Tamhane,1987), it is appropriate to use the MtI
method for estimating one-sided simultaneous confi-
dence limits only when all the elements of the corre-
lation matrix are non-negative. Therefore, if some of
the (estimated) correlations are negative, sci.ratio
and sci.ratio.gen functions report a warning mes-
sage about the inappropriateness of the MtI method.
ii) In simultaneous CI estimation (using either
sci.ratio or simtest.ratio.gen), one may en-
counter the case where some of the contrasts in the
R News ISSN 1609-3631

Vol. 7/1, April 2007 23
denominators of the ratios are not significantly dif-
ferent from zero. In this situation, NSD (standing for
“non-significant denominator”) will be printed. For
instance, in Example 2 above, since there is no signif-
icant difference between the placebo and the active
control, one gets NSD in constructing the related si-
multaneous CIs for the three ratios.
Sample Size Calculation
Consider the design of a special problem in simulta-
neous comparison of m≥2 treatments with a control
for non-inferiority (or superiority), where the mar-
gins are expressed as a percentage of the mean of
the control group. For sample size calculation, we
implement a method based on normal approxima-
tion to the exact method which involves inversion of
a univariate (multivariate) non-central t-distribution
(see Dilba et al. (2006b) for details on the exact
method). Given the number of comparisons (m), the
non-inferiority (superiority) margin (rho), the power
(Power), the coefficient of variation of the control
group (CV0), the percentage (of the mean of the con-
trol group) to be detected (rho.star), the family-
wise type-I error rate (alpha), and the kind of power
to be controlled (by default minimal power), the
function n.ratio calculates the sample size required
in a balanced design.
Example 5. Suppose that we have a response
variable where large response values indicate better
treatment benefits. The following R code calculates
the sample size required per treatment in designing
a non-inferiority trial with four treatment arms (in-
cluding the control).
> n.ratio(m = 3, rho = 0.7, Power = 0.8,
+ CV0 = 0.5, rho.star = 0.95,
+ alpha = 0.05, Min.power = TRUE)
Number of observations per treatment = 52
Total number of observations = 208
If the aim is to control the complete power, we set
Min.power to FALSE.
For the two-sample design (m=1), the sample
sizes required in the non-inferiority trials discussed
by Laster and Johnson (2003) can be calculated as a
special case.
Remarks
We conclude by giving some general remarks regard-
ing the four basic functions in the mratios package.
• In two-sample ratio problems with homoge-
neous variances, t.test.ratio is a special case
of simtest.ratio and sci.ratio.
• The simtest.ratio function with all the el-
ements of the vector of margins equal to 1
gives the same result as the analysis based on
the difference of treatment means. Thus, the
difference-based test is a special case of the
ratio-based test with the thresholds set to 1.
• The sci.ratio function is a special case of
sci.ratio.gen for the one-way layout.
Bibliography
P. Bauer, J. Röhmel, W. Maurer and L.A. Hothorn.
Testing strategies in multi-dose experiments in-
cluding active control. Statistics in Medicine, 17:
2133-2146, 1998. 21
G. Dilba, F. Bretz and V. Guiard. Simultaneous con-
fidence sets and confidence intervals for multiple
ratios. Journal of Statistical Planning and Inference,
136:2640–2658, 2006a. 21
G. Dilba, F. Bretz, L.A. Hothorn and V. Guiard.
Power and sample size computations in simul-
taneous tests for non-inferiority based on rela-
tive margins. Statistics in Medicine, 25:1131–1147,
2006b. 23
J. Hochberg and A. Tamhane. Multiple comparison
procedures. Wiley, New York, 1987, p366. 21,22
T. Hothorn, F. Bretz and A. Genz. On Multivariate t
and Gauss Probabilities. R News, 1(2):27–29, 2001.
20
G. R. Jensen. Joint confidence sets in multiple dilu-
tion assays. Biometrical Journal, 31:841–853, 1989.
22
L. L. Laster and M. F. Johnson. Non-inferiority tri-
als: the ‘at least as good as’ criterion. Statistics in
Medicine, 22:187–200, 2003.
I. Pigeot, J. Schäfer, J. Röhmel and D. Hauschke.
Assessing non-inferiority of a new treatment in a
three-arm clinical trial including a placebo. Statis-
tics in Medicine, 22:883–899, 2003. 21
A. C. Tamhane and B. R. Logan. Finding the maxi-
mum safe dose level for heteroscedastic data. Jour-
nal of Biopharmaceutical Statistics, 14:843–856, 2004.
20
P. H. Westfall, R. D. Tobias, D. Rom, R. D. Wolfinger
and Y. Hochberg. Multiple comparisons and mul-
tiple tests using the SAS system. Cary, NC, SAS In-
stitute Inc,1999, 65–81.
21
G. Dilba, F. Schaarschmidt, L. A. Hothorn
Institute of Biostatistics
Faculty of Natural Sciences
Leibniz University of Hannover, Germany
dilba@biostat.uni-hannover.de
schaarschmidt@biostat.uni-hannover.de
hothorn@biostat.uni-hannover.de
R News ISSN 1609-3631

Vol. 7/1, April 2007 24
Working with Unknown Values
The gdata package
by Gregor Gorjanc
Introduction
Unknown or missing values can be represented in
various ways. For example SAS uses .(dot), while
R uses NA, which we can read as Not Available.
When we import data into R, say via read.table
or its derivatives, conversion of blank fields to NA
(according to read.table help) is done for logical,
integer,numeric and complex classes. Addition-
ally, the na.strings argument can be used to spec-
ify values that should also be converted to NA. In-
versely, there is an argument na in write.table and
its derivatives to define value that will replace NA
in exported data. There are also other ways to im-
port/export data into R as described in the RData
Import/Export manual (R Development Core Team,
2006). However, all approaches lack the possibility
to define unknown value(s) for some particular col-
umn. It is possible that an unknown value in one
column is a valid value in another column. For ex-
ample, I have seen many datasets where values such
as 0, -9, 999 and specific dates are used as column
specific unknown values.
This note describes a set of functions in package
gdata1(Warnes ,2006): isUnknown,unknownToNA and
NAToUnknown, which can help with testing for un-
known values and conversions between unknown
values and NA. All three functions are generic (S3)
and were tested (at the time of writing) to work
with: integer,numeric,character,factor,Date,
POSIXct,POSIXlt,list,data.frame and matrix
classes.
Description with examples
The following examples show simple usage of these
functions on numeric and factor classes, where
value 0(beside NA) should be treated as an unknown
value:
> library("gdata")
> xNum <- c(0, 6, 0, 7, 8, 9, NA)
> isUnknown(x=xNum)
[1] FALSE FALSE FALSE FALSE FALSE FALSE
TRUE
The default unknown value in isUnknown is NA,
which means that output is the same as is.na — at
least for atomic classes. However, we can pass the
argument unknown to define which values should be
treated as unknown:
> isUnknown(x=xNum, unknown=0)
[1] TRUE FALSE TRUE FALSE FALSE FALSE
FALSE
This skipped NA, but we can get the expected an-
swer after appropriately adding NA into the argu-
ment unknown:
> isUnknown(x=xNum, unknown=c(0, NA))
[1] TRUE FALSE TRUE FALSE FALSE FALSE
TRUE
Now, we can change all unknown values to NA
with unknownToNA. There is clearly no need to add
NA here. This step is very handy after importing
data from an external source, where many differ-
ent unknown values might be used. Argument
warning=TRUE can be used, if there is a need to be
warned about “original” NAs:
> xNum2 <- unknownToNA(x=xNum, unknown=0)
[1] NA 6 NA 7 8 9 NA
Prior to export from R, we might want to change
unknown values (NA in R) to some other value. Func-
tion NAToUnknown can be used for this:
> NAToUnknown(x=xNum2, unknown=999)
[1] 999 6 999 7 8 9 999
Converting NA to a value that already exists in x
issues an error, but force=TRUE can be used to over-
come this if needed. But be warned that there is no
way back from this step:
> NAToUnknown(x=xNum2, unknown=7,
force=TRUE)
[1]7677897
Examples below show all peculiarities with class
factor.unknownToNA removes unknown value from
levels and inversely NAToUnknown adds it with a
warning. Additionally, "NA" is properly distin-
guished from NA. It can also be seen that the
argument unknown in functions isUnknown and
unknownToNA need not match the class of x(other-
wise factor should be used) as the test is internally
done with %in%, which nicely resolves coercing is-
sues.
> xFac <- factor(c(0, "BA", "RA", "BA",
NA, "NA"))
[1] 0 BA RA BA <NA> NA
Levels: 0 BA NA RA
> isUnknown(x=xFac)
[1] FALSE FALSE FALSE FALSE TRUE FALSE
> isUnknown(x=xFac, unknown=0)
[1] TRUE FALSE FALSE FALSE FALSE FALSE
> isUnknown(x=xFac, unknown=c(0, NA))
[1] TRUE FALSE FALSE FALSE TRUE FALSE
1package version 2.3.1
R News ISSN 1609-3631

Vol. 7/1, April 2007 25
> isUnknown(x=xFac, unknown=c(0, "NA"))
[1] TRUE FALSE FALSE FALSE FALSE TRUE
> isUnknown(x=xFac, unknown=c(0, "NA", NA))
[1] TRUE FALSE FALSE FALSE TRUE TRUE
> xFac <- unknownToNA(x=xFac, unknown=0)
[1] <NA> BA RA BA <NA> NA
Levels: BA NA RA
> xFac <- NAToUnknown(x=xFac, unknown=0)
[1] 0 BA RA BA 0 NA
Levels: 0 BA NA RA
Warning message:
new level is introduced: 0
These two examples with classes numeric and
factor are fairly simple and we could get the same
results with one or two lines of R code. The real ben-
efit of the set of functions presented here is in list
and data.frame methods, where data.frame meth-
ods are merely wrappers for list methods.
We need additional flexibility for list/data.frame
methods, due to possibly having multiple unknown
values that can be different among list components
or data.frame columns. For these two methods, the
argument unknown can be either a vector or list,
both possibly named. Of course, greater flexibil-
ity (defining multiple unknown values per compo-
nent/column) can be achieved with a list.
When a vector/list object passed to the
argument unknown is not named, the first
value/component of a vector/list matches the first
component/column of a list/data.frame. This
can be quite error prone, especially with vectors.
Therefore, I encourage the use of a list. In case
vector/list passed to argument unknown is named,
names are matched to names of list or data.frame.
If lengths of unknown and list or data.frame do not
match, recycling occurs.
The example below illustrates the application of
the described functions to a list which is composed
of previously defined and modified numeric (xNum)
and factor (xFac) classes. First, function isUnknown
is used with 0as an unknown value. Note that we
get FALSE for NAs as has been the case in the first ex-
ample.
> xList <- list(a=xNum, b=xFac)
$a
[1]060789NA
$b
[1] 0 BA RA BA 0 NA
Levels: 0 BA NA RA
> isUnknown(x=xList, unknown=0)
$a
[1] TRUE FALSE TRUE FALSE FALSE FALSE
FALSE
$b
[1] TRUE FALSE FALSE FALSE TRUE FALSE
We need to add NA as an unknown value. How-
ever, we do not get the expected result this way!
> isUnknown(x=xList, unknown=c(0, NA))
$a
[1] TRUE FALSE TRUE FALSE FALSE FALSE
FALSE
$b
[1] FALSE FALSE FALSE FALSE FALSE FALSE
This is due to matching of values in the argument
unknown and components in a list; i.e., 0is used for
component aand NA for component b. Therefore, it
is less error prone and more flexible to pass a list
(preferably a named list) to the argument unknown,
as shown below.
> xList1 <- unknownToNA(x=xList,
+ unknown=list(b=c(0, "NA"), a=0))
$a
[1] NA 6 NA 7 8 9 NA
$b
[1] <NA> BA RA BA <NA> <NA>
Levels: BA RA
Changing NAs to some other value (only one per
component/column) can be accomplished as fol-
lows:
> NAToUnknown(x=xList1,
+ unknown=list(b="no", a=0))
$a
[1]0607890
$b
[1] no BA RA BA no no
Levels: BA no RA
Warning message:
new level is introduced: no
A named component .default of a list passed
to argument unknown has a special meaning as it will
match a component/column with that name and any
other not defined in unknown. As such it is very use-
ful if the number of components/columns with the
same unknown value(s) is large. Consider a wide
data.frame named df. Now .default can be used
to define unknown value for several columns:
> df <- unknownToNA(x=df,
+ unknown=(.default=0,
+ col1=999,
+ col2="unknown"))
If there is a need to work only on some com-
ponents/columns you can of course “skip” columns
with standard R mechanisms, i.e., by subsetting list
or data.frame objects:
> cols <- c("col1", "col2")
> df[, cols] <- unknownToNA(x=df[, cols],
+ unknown=(col1=999,
+ col2="unknown"))
R News ISSN 1609-3631

Vol. 7/1, April 2007 26
Summary
Functions isUnknown,unknownToNA and NAToUnknown
provide a useful interface to work with various rep-
resentations of unknown/missing values. Their use
is meant primarily for shaping the data after import-
ing to or before exporting from R. I welcome any
comments or suggestions.
Bibliography
R Development Core Team. R Data Import/Export,
2006. URL http://cran.r-project.org/
manuals.html. ISBN 3-900051-10-0. 24
G. R. Warnes. gdata: Various R program-
ming tools for data manipulation, 2006. URL
http://cran.r-project.org/src/contrib/
Descriptions/gdata.html. R package version
2.3.1. Includes R source code and/or documenta-
tion contributed by Ben Bolker, Gregor Gorjanc
and Thomas Lumley. 24
Gregor Gorjanc
University of Ljubljana, Slovenia
gregor.gorjanc@bfro.uni-lj.si
A New Package for Fitting Random Effect
Models
The npmlreg package
by Jochen Einbeck, John Hinde, and Ross Darnell
Introduction
Random effects have become a standard concept in
statistical modelling over the last decades. They
enter a wide range of applications by providing a
simple tool to account for such problems as model
misspecification, unobserved (latent) variables, un-
observed heterogeneity, and the like. One of the most
important model classes for the use of random effects
is the generalized linear model. Aitkin (1999) noted
that “the literature on random effects in generalized
linear models is now extensive,” and this is certainly
even more true today.
However, most of the literature and the imple-
mented software on generalized linear mixed mod-
els concentrates on a normal random effect distri-
bution. An approach that avoids specifying this
distribution parametrically was provided by Aitkin
(1996a), using the idea of ’Nonparametric Maximum
Likelihood’ (NPML) estimation (Laird,1978). The
random effect distribution can be considered as an
unknown mixing distribution and the NPML esti-
mate of this is a finite discrete distribution. This can
be determined by fitting finite mixture distributions
with varying numbers of support points, where each
model is conveniently fitted using a straightforward
EM algorithm.
This approach is implemented in GLIM4 (Aitkin
and Francis,1995). Despite being a quite pow-
erful tool, the current GLIM-based software is
computationally limited and the GLIM system is
no longer widely used. Though the alternatives
C.A.MAN (Böhning et al.,1992) and the Stata pro-
gram gllamm (Skrondal and Rabe-Hesketh,2004)
cover parts of GLIMs capacities (in the latter case
based on Newton-Raphson instead of EM), no R im-
plementation of NPML estimation existed. The pack-
age npmlreg (Einbeck et al.,2006), which we wish to
introduce to the R community in this article, is de-
signed to fill this gap.
NPML estimation
Assume there is given a set of explanatory vec-
tors x1,...,xnand a set of observations y1,...,yn
sampled from an exponential family distribution1
f(yi|β,φi)with dispersion2parameter φi. In a gen-
eralized linear model, predictors and response are
assumed to be related through a link function h,
µi≡E(yi|β,φi) = h(ηi)≡h(x0
iβ),
and the variance Var(yi|β,φi) = φiv(µi)depends on
a function v(µi)which is entirely determined by the
choice of the particular exponential family. However,
often the actual variance in the data is larger than the
variance according to this strict mean-variance rela-
tionship. This effect is commonly called overdisper-
sion, reasons for which might be, e.g., correlation in
the data or important explanatory variables not in-
cluded in the model. In order to account for addi-
tional unexplained variability of the individual ob-
servations, a random effect ziwith density g(z)is in-
1In the present implementation, Gaussian, Poisson, Binomial, and Gamma distributed responses are supported
2For binomial and Poisson models, φi≡1. For Gaussian and Gamma models, the dispersion may be specified as constant, i.e., φi≡φ,
or as depending on the observation i. The theory in this section is provided for the most general case, i.e., variable dispersion.
R News ISSN 1609-3631

Vol. 7/1, April 2007 27
cluded into the linear predictor
ηi=x0
iβ+zi.
The likelihood can be approximated by a finite mix-
ture
L=
n
∏
i=1Zf(yi|zi,β,φi)g(zi)dzi≈
n
∏
i=1(K
∑
k=1
fik πk),
where fik =f(yi|zk,β,φk),zkare the mass points
and πktheir masses. The score equations, obtained
by setting the partial derivatives of the log-likelihood
`=log Lequal to zero,
∂`
∂zk
=0, ∂`
∂β=0, ∂`
∂φk
=0,
turn out to be weighted versions of the single-
distribution score equations, with weights wik =
πkfik /∑lπlfil .
The weights wik can be interpreted as posterior
probabilities that the observation yicomes from com-
ponent k. The score equation for the mixture propor-
tions,
∂`−λ(∑πk−1)
∂πk
=0,
gives the ML estimate ˆπk=1
n∑iwik, which can be
nicely interpreted as the average posterior probabil-
ity for component k. The parameters β,zkand πkcan
now be simultaneously estimated by the EM algo-
rithm, whereby the missing information is the com-
ponent membership of the observations:
E-Step Adjust weights wik =P(obs. icomes from
comp. k)
M-Step Update parameter estimates fitting a
weighted GLM with weights wik.
As starting values for the EM algorithm one uses
Gauss-Hermite integration points and masses. The
location of these starting points can be scaled in-
wards or outwards by means of a tuning parameter
tol, which is by default set to 0.5.
This procedure, and its straightforward extension
to random coefficient models, is implemented in the
function alldist, while variance component models
can be fitted with allvc; see Aitkin et al. (2005), pp
474ff and 485ff for details.
The function alldist
The main functions of this package are alldist and
allvc, the names of which were adapted from the
homonymous macros in GLIM4. The functions can
be used in a similar manner to the R function glm.
As an example for alldist, we consider data
from a study on lung cancer mortality presented in
Tsutakawa (1985). The data were recorded in the 84
largest Missouri cities from 1972-1981 and give the
number of lung cancer deaths of males aged 45-54 as
well as the city sizes3. The data were analyzed by
Tsutakawa (1985) and Aitkin (1996b), both authors
considering logit models of type
log pi
1−pi
=zi,
where ziis a random effect associated with the i−th
city and piis its associated mortality rate. While Tsu-
takawa fitted a Poisson model with a normal ran-
dom effect, Aitkin opted for a binomial model with
an unspecified random effect distribution. We follow
Aitkin and will leave the random effect unspecified,
but as lung cancer death is a rather rare event (the
crude rate does not exceed 0.03 in any of the cities), it
seems natural to work with Poisson models. Hence,
one can write log(pi) = zi, or equivalently, in terms
of the means µi=nipi,
log(µi) = log(ni) + zi,
where the logarithm of the city sizes niappears as
an offset. The two-point solution is then obtained
via the function alldist, using the same notation
as for a usual glm fit, except that the random term
and the number of mass points k=2 have also to be
specified. The resulting object (which we name, say,
missouri.np2) is of class ‘glmmNPML’ and its printed
output is given by
> print(missouri.np2)
Call: alldist(formula = Deaths ~1, random =
~1, family = poisson(link = "log"), data =
missouri, k = 2, offset = log(Size))
Coefficients:
MASS1 MASS2
-4.844 -4.232
Mixture proportions:
MASS1 MASS2
0.8461624 0.1538376
-2 log L: 355.3
One minor difference to a glm output is that the
disparity (−2 log L) is displayed instead of the de-
viance, but the latter one is immediately obtained via
> missouri.np2$dev
[1] 92.49207,
which is slightly better than the deviance 93.10 re-
ported in Aitkin (1996b) for the corresponding two-
mass point binomial logit model fitted with GLIM4.
As also observed by Aitkin, the disparity does not
3The data set missouri is part of this R package
R News ISSN 1609-3631
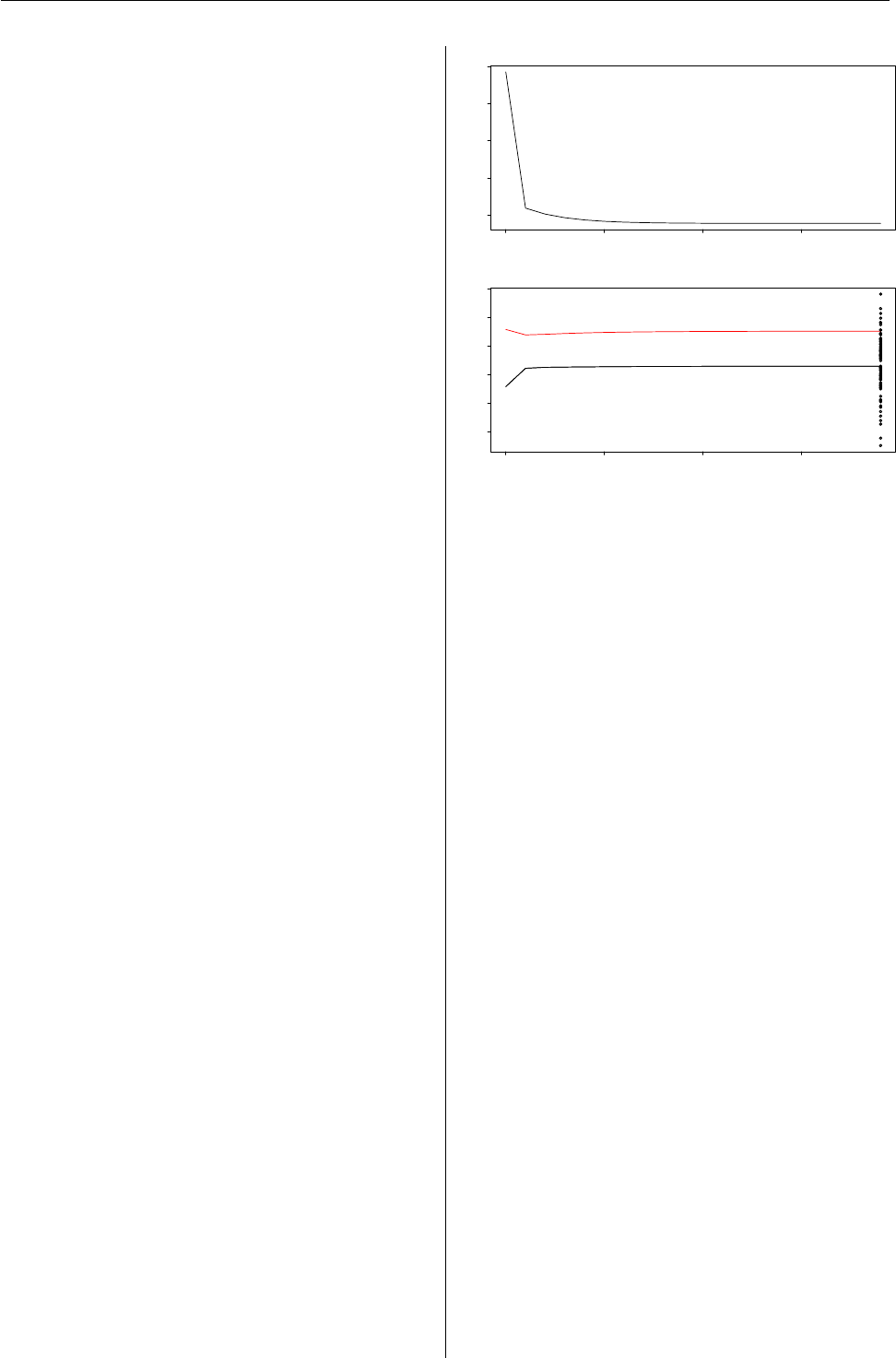
Vol. 7/1, April 2007 28
fall significantly when increasing the number of
mass points further.
Empirical Bayes predictions (generally, h(x0
iˆ
β+
ˆ
zi)for the number of deaths per city can be obtained
by the use of fitted(missouri.np2), or equiv-
alently, missouri.np2$fitted.values, or equiva-
lently, predict(missouri.np2, type="response").
Dividing this quantity by missouri$Size, one ob-
tains estimated or ‘shrunken’ rates which can be
compared to the 6th column in Table 4, Aitkin
(1996b). The shrunken rates are less variable than
the crude rates and hence are useful for small area
estimation problems. The posterior probabilities wik
can be obtained from the fitted model (in analogy to
the 8th and 9th column of Table 4, Aitkin,1996b), via
the component $post.prob. Further, ’posterior in-
tercepts’ for the construction of ’league tables’ are
stored in component $post.int — see Sofroniou
et al. (2006) for an example of their application.
Methods for most generic functions that
are applied to fitted model objects, such
as update(), coefficients(), residuals(),
fitted(), summary(), predict() and plot(),
have been defined for the glmmNPML class. In some
cases (the first four generic functions listed above)
the default method is used; in other cases (the last
three generics) explicit methods are provided. The
plot() function offers four different plots: (i) dis-
parity vs. EM iterations; (ii) EM trajectories; (iii)
Empirical Bayes predictions vs. true response; and
(iv) posterior probabilities against the residuals of
the fixed part of the GLM fit in the last EM itera-
tion. Plots (i) and (ii) are generated by default when
running alldist and are depicted in Fig. 1for the
model fitted above.
One observes that the EM trajectories converge
essentially to the fixed part residuals of cities no. 4
and 84 (in Tsutakawa’s list), which have populations
of 54155 and 22514, respectively, being much larger
than the majority of the other cities with only several
hundreds of inhabitants (For the very interested, the
numerical values associated with the disparity plot
and the EM trajectories, as well as the residuals plot-
ted vertically in the latter plot, are available in com-
ponent $Misc).
This was a simple example without any covari-
ates. In general an arbitrary number of fixed effects
can be specified, and the random component can be
an intercept (∼1) or a single variable giving a model
with a random slope and random intercept.
0 5 10 15
360 380 400 420 440
EM iterations
−2logL
0 5 10 15
−6.0 −5.5 −5.0 −4.5 −4.0 −3.5
EM iterations
mass points
Figure 1: Top: Disparity (−2 log L) trend; bottom:
EM trajectories. The vertical dots in the latter plot are
the residuals of the fixed part of the fitted random
effect model (Note that these residuals are not cen-
tered around 0, in contrast to the residuals of a simple
fixed effect model. The plotted residuals, generally
h−1(yi)−x0
iˆ
β, represent the random effect distribu-
tion and are on the same scale as the mass points).
The function allvc
The function alldist is designed to fit simple
overdispersion models (i.e., one has a random effect
on the individual observations). However, often one
wishes to introduces shared random effects, e.g. for
students from the same class or school, for the same
individual observed repeatedly over time (longitudi-
nal data), or in small area estimation problems. This
leads to variance component models, which can be
fitted in npmlreg using the function allvc. As an
example, let us consider the Oxford boys data from
Brush and Harrison (1990), which were analyzed
with NPML using GLIM4 by Aitkin et al. (2005).
The data set is part of the R package nlme (Pin-
heiro et al.,2005) and contains the heights of 26
boys, measured in Oxford on nine occasions over
two years. The boys, indexed by the factor Subject,
represent the upper level (primary sampling units,
PSU), and the particular measurements at different
time points correspond to the lower-level units (sec-
ondary sampling units, SSU).
As suggested by (Aitkin et al.,2005, p. 495), we
fit a Gaussian model with unspecified random effect
distribution and K=7 mass points,
(Oxboys.np7 <- allvc(height ~ age, random =
~1|Subject, data = Oxboys, k=7))$disparity
[1] 1017.269
R News ISSN 1609-3631

Vol. 7/1, April 2007 29
which confirms the GLIM4 result. Aitkin et al. state
that for all models using K>7 the fitted mass
points ‘are duplicated with the same total mass as
in lower-dimension models’, and hence consider this
7- mass point model as ‘sufficiently complex’. How-
ever, fitting for comparison a simple linear mixed
model with function lmer in package lme4 (Bates
and Sarkar,2006)
(fm1 <- lmer(height ~ age + (1|Subject),
data = Oxboys, method = "ML"))
gives the MLdeviance, i.e., disparity, of 940.569. As
this model is based on a normal random effect distri-
bution, NPML should be superior to this, or at least
competitive with it. Therefore, we went on fitting
models with K=8 and K=9 mass points, yielding
(Oxboys.np8 <- allvc(height ~ age, random =
~1|Subject, data = Oxboys, k=8))$disparity
[1] 931.3752
(Oxboys.np9 <- allvc(height ~ age, random =
~1|Subject, data = Oxboys, k=9,
tol=0.3))$disparity
[1] 916.0921
For both models, all mass points are indeed distinct.
For instance, for the 9-mass point model one has
Estimate Std. Error
age 6.523719 0.05094195
MASS1 130.200174 0.16795344
MASS2 138.416628 0.09697325
MASS3 143.382397 0.09697225
MASS4 147.350113 0.07511877
MASS5 150.954327 0.06857400
MASS6 153.869256 0.11915344
MASS7 156.178153 0.09676418
MASS8 159.521550 0.16795354
MASS9 164.883640 0.11876372
This increased performance compared to the
GLIM code is due to the installation of a ‘damping’
mechanism in the first cycles of the EM algorithm;
see Einbeck and Hinde (2006) for details.
Further increase of Kdoes not yield major drops
in disparity, so we continue to work with nine mass
points, and extend the model by allowing the lin-
ear trend to vary across boys. The function call then
takes the form
(Oxboys.np9s <- allvc(height ~ age, random =
~age|Subject, data = Oxboys, k = 9,
tol=0.3))
The difference in disparities is
> Oxboys.np9$disparity - Oxboys.np9s$disparity
[1] 102.1071
on
> Oxboys.np9$df.res - Oxboys.np9s$df.res
[1] 8
degrees of freedom, showing clear heterogeneity in
the slope.
Summary
We have introduced the R package npmlreg, which
is in some parts a simple translation from the corre-
sponding GLIM4 code, but, in other parts, contains
substantial extensions and methodological improve-
ments. In particular, we mention the possibility to
fit Gamma models and to work with dispersion pa-
rameters varying smoothly over components, and, as
already noted, the installation of a damping proce-
dure. We note finally that for the sake of compara-
bility all implemented features are also available for
Gaussian Quadrature instead of NPML (leaving the
zkand πkfixed and equal to Gauss-Hermite integra-
tion points and masses). The R package is available
on CRAN. Future developments will include 3-level
models and multicategory responses.
Acknowledgements
The work on this R package was supported by
Science Foundation Ireland Basic Research Grant
04/BR/M0051. We are grateful to N. Sofroniou for
sharing his GLIM code for the Missouri data and two
anonymous referees for pointing at the comparison
with package lme4.
Bibliography
M. Aitkin. A general maximum likelihood analy-
sis of overdispersion in generalized linear models.
Statistics and Computing, 6:251–262, 1996a. 26
M. Aitkin. Empirical Bayes shrinkage using posterior
random effect means from nonparametric maxi-
mum likelihood estimation in general random ef-
fect models. In IWSM 1996: Proceedings of the 11th
International Workshop on Statistical Modelling, Orvi-
eto, pages 87–94, 1996b. 27,28
M. Aitkin. A general maximum likelihood analysis
of variance components in generalized linear mod-
els. Biometrics, 55:218–234, 1999. 26
M. Aitkin and B. Francis. Fitting overdispersed
generalized linear models by nonparametric max-
imum likelihood. The GLIM Newsletter, 25:37–45,
1995. 26
M. Aitkin, B. Francis, and J. Hinde. Statistical Mod-
elling in GLIM 4. Oxford Statistical Science Series,
Oxford, UK., 2005. 27,28
R News ISSN 1609-3631

Vol. 7/1, April 2007 30
D. Bates and D. Sarkar. lme4: Linear mixed-effects mod-
els using S4 classes, 2006. R package version 0.995-2.
29
D. Böhning, P. Schlattmann, and B. Lind-
sey. Computer-assisted analysis of mixtures
(C.A.MAN): statistical algorithms. Biometrics, 48:
283–303, 1992. 26
G. Brush and G. A. Harrison. Components of growth
variation in human stature. Mathematical Medicine
and Biology, 7:77–92, 1990. 28
J. Einbeck and J. Hinde. A note on NPML estimation
for exponential family regression models with un-
specified dispersion parameter. Austrian Journal of
Statistics, 35:233–243, 2006. 29
J. Einbeck, R. Darnell, and J. Hinde. npmlreg: Non-
parametric maximum likelihood estimation for random
effect models, 2006. R package version 0.40. 26
N.M. Laird. Nonparametric maximum likelihood es-
timation of a mixing distribution. Journal of the
American Statistical Association, 73:805–811, 1978.
26
J. Pinheiro, D. Bates, S. DebRoy, and D. Sarkar. nlme:
Linear and nonlinear mixed effects models, 2005. R
package version 3.1-66. 28
A. Skrondal and S. Rabe-Hesketh. Generalized Latent
Variable Modelling. Chapman & Hall/CRC, Boca
Raton, 2004. 26
N. Sofroniou, J. Einbeck, and J. Hinde. Analyzing
Irish suicide rates with mixture models. In IWSM
2006: Proceedings of the 21th International Work-
shop on Statistical Modelling, Galway, pages 474–481,
2006. 28
R. Tsutakawa. Estimation of cancer mortality rates: a
Bayesian analysis of small frequencies. Biometrics,
41:69–79, 1985. 27
Jochen Einbeck
Durham University, UK
jochen.einbeck@durham.ac.uk
John Hinde
National University of Ireland, Galway, Rep. of Ireland
john.hinde@nuigalway.ie
Ross Darnell
The University of Queensland, Australia
r.darnell@uq.edu.au
Augmenting R with Unix Tools
Andrew Robinson
This article describes a small collection of Unix
command-line tools that can be used to augment R. I
introduce:
make which after judicious preparation will allow
one-word execution of large-scale simulations,
all the way from compilation of source code to
construction of a PDF report;
screen which will permit remote monitoring of pro-
gram execution progress with automatic pro-
tection against disconnection; and
mail which will allow automatic alerting under er-
ror conditions or script completion.
These programs will be part of the default instal-
lations of many flavours of Unix-like operating sys-
tems. Although these tools have many different ap-
plications, each is very useful for running R remotely
on a Unix-like operating system, which is the focus of
their usage in this article.
Regardless of what kind of computer is on your
own desk, you may find these tools useful; they do
not require you to be running a BSD or GNU/Linux
on your own machine.
R and screen
It is sometimes necessary to run R code that executes
for long periods of time upon remote machines. This
requirement may be because the local computing re-
sources are too slow, too unstable, or have insuffi-
cient memory.
For example, I have recently completed some
simulations that took more than a month on a rea-
sonably fast cluster of Linux computers. I devel-
oped, trialed, and profiled my code on my local, not-
particularly-fast, computer. I then copied the scripts
and data to a more powerful machine provided by
my employer and ran the simulations on R remotely
on that machine. To get easy access to the simula-
tions whilst they ran, I used screen.
screen is a so-called terminal multiplexor, which
allows us to create, shuffle, share, and suspend com-
mand line sessions within one window. It provides
protection against disconnections and the flexibility
to retrieve command line sessions remotely. screen
is particularly useful for R sessions that are running
on a remote machine.
We might use the following steps to invoke R
within screen:
1. log in remotely via secure shell,
R News ISSN 1609-3631
Vol. 7/1, April 2007 31
2. start screen,
3. start R,
4. source our script with echo=TRUE,
5. detach the screen session, using Ctrl-a d, and
6. log out.
The R session continues working in the background,
contained within the screen session. If we want to
revisit the session to check its progress, then we
1. log in remotely via secure shell,
2. start screen -r, which recalls the unattached
session,
3. examine the saved buffer; scrolling around,
copying and pasting as necessary,
4. detach the screen session, using Ctrl-a d, and
5. log out.
This approach works best if examining the buffer
is informative, which requires that the R script
be written to provide readable output or flag its
progress every so often. I find that the modest de-
crease in speed of looping is more than compensated
by a cheerful periodic reminder, say every 1000th
iteration, that everything is still working and that
there are only niterations left to run. I have also
been experimenting with various algorithms for R to
use the elapsed time, the elapsed number of simula-
tions, and the number of simulations remaining, to
estimate the amount of time left until the run is com-
plete.
screen offers other advantages. You can manage
several screens in one window, so editing a script re-
motely using, say, emacs, and then sourcing the script
in the remote R session in another screen, is quick
and easy. If you lose your connection, the session is
kept alive in the background, and can be re-attached,
using screen -r as above. If you have forgotten
to detach your session you can do so forcibly from
another login session, using screen -dr. You can
change the default history/scrollback buffer length,
and navigate the buffer using intuitive keystrokes.
Finally, you can share a screen session with other
people who are logged in to the same machine. That
is, each user can type and see what the other is typ-
ing, so a primitive form of online collaboration is
possible.
So, running an R session in screen provides a
simple and robust way to allow repeated access to
a simulation or a process.
More information about screen can be found
from http://www.gnu.org/software/screen/, or
man screen.
R and mail
It is very useful to be able to monitor an R session
that is running remotely. However, it would also be
useful if the session could alert us when the script has
completed, or when it has stopped for some reason,
including some pre-determined error conditions. We
can use mail for this purpose, if the computer is run-
ning an appropriate mail server, such as sendmail.
mail is an old email program, small enough to
be included by default on most Unix-like systems,
and featureless enough to be almost universally ig-
nored by users in favour of other programs, such as
mutt, or those with shiny graphical user interfaces.
However, mail does allow us to send email from the
command line. And, R can do pretty much anything
that can be done from the command line1. Of course,
a small amount of fiddling is necessary to make it
work. A simple function will help2. This function
will fail if any of the arguments contain single or
double quotes. So, craft your message carefully.
mail <- function(address, subject, message) {
system(paste("echo ", message,
" | mail -s ", subject,
" ", address, sep=""))
}
We can now send mail from R via, for example,
mail("andrewr", "Test", "Hello world!")
You can place calls to this function in strategic loca-
tions within your script. I call it at the end of the
script to let me know that the simulations are fin-
ished, and I can collect the results at my leisure. In
order to have R let us know when it has stopped, we
can use a function like this:
alert <- function() {
mail("andrewr", "Stopped", "Problem")
browser()
}
then in our script we call
options(error = alert)
Now in case of an error, R will send an email and
drop into the browser, to allow detailed follow-up. If
you have more than one machine running, then call-
ing hostname via system, and pasting that into the
subject line, can be helpful.
Alternatively, you can use mutt with exactly the
same command line structure as mail, if mutt is in-
stalled on your system. An advantage of mutt is
that it uses the MIME protocol for binary attachments.
That would enable you to, say, attach to your email
the PDF that your script has just created with Sweave
and pdflatex, or the cvs file that your script creates,
1In theory, it can do everything, I suppose. I haven’t tried.
2Caveat: this function is written to work on bash version 3.1.17. Your mileage may vary.
R News ISSN 1609-3631

Vol. 7/1, April 2007 32
or even the relevant objects saved from the session,
neatly packaged and compressed in a *.RData object.
Different flavours of Unix mail are available. You
can find out more about yours by man mail.
R and make
I have recently been working on bootstrap tests of
a model-fitting program that uses maximum likeli-
hood to fit models with multivariate normal and t
distributions. The program is distributed in FOR-
TRAN, and it has to be compiled every time that it is
run with a new model. I wanted to use a studentized
bootstrap for interval estimates, gather up all the out-
put, and construct some useful tables and graphics.
So, I need to juggle FORTRAN files, FORTRAN
executables, R source files, Sweave files, PDFs, and
so on. R does a great job preparing the input files,
calling the executable (using system()), and scoop-
ing up the output files. However, to keep everything
in sync, I use make.
Although it is commonly associated with build-
ing executable programs, make can control the con-
version of pretty much any file type into pretty much
any other kind of file type. For example, make can be
told how to convert a FORTRAN source file to an ex-
ecutable, and it will do so if and when the source file
changes. It can be told how and when to run an R
source file, and then how and when to call Sweave
upon an existing file, and then how and when to cre-
ate a PDF from the resulting L
A
T
EX file.
The other advantage that make offers is splitting
large projects into chunks. I use Sweave to bind my
documentation and analysis tightly together. How-
ever, maintaining the link between documentation
and analysis can be time-consuming. For example,
when documenting a large-scale simulation, I would
rather not run the simulation every time I correct a
spelling mistake.
One option is to tweak the number of runs. make
provides a flexible infrastructure for testing code, as
we can pass parameters, such as the number of runs
to perform, to R. For example, with an appropriate
Makefile, typing make test at the operating system
prompt will run the entire project with only 20 simu-
lations, whereas typing make will run the project with
2000 simulations.
Another option is to split the project into two
parts: the simulation, and the analysis, and run only
the second, unless important elements change in the
first. Again, this sort of arrangement can be con-
structed quite easily using make.
For example, I have an R source file called sim.r
that controls the simulations and produces a .RData
object called output.RData. The content is:
randoms <- runif(reps)
save.image("output.RData")
I also have a Sweave file called report.rnw which
provides summary statistics (and graphics, not in-
cluded here) for the runs. The content is:
\documentclass{article}
\begin{document}
<<>>=
load("output.RData")
@
We performed \Sexpr{reps} simulations.
\end{document}
The latter file often requires tweaking, depending
on the output. So, I want to separate the simulations
and the report writing, and only run the simulations
if I absolutely have to. Figure 1is an example Make-
file that takes advantage of both options noted above.
All the files referenced here are assumed to be held in
the same directory as the Makefile, but of course they
could be contained in subdirectories, which is gener-
ally a neater strategy.
I do not actually need make to do these tasks, but
it does simplify the operation a great deal. One ad-
vantage is that make provides conditional execution
without me needing to fiddle around to see what has
and has not changed. If I now edit report.rnw, and
type make, it won’t rerun the simulation, it will just
recompile report.rnw. On the other hand, if I make
a change to sim.r, or even run sim.r again (thus
updating the .RData object), make will rerun every-
thing. That doesn’t seem very significant here, but it
can be when you’re working on a report with numer-
ous different branches of analysis.
The other advantage (which my Makefile doesn’t
currently use) is that it provides wildcard match-
ing. So, if I have a bunch of Sweave files (one for
each chapter, say) and I change only one of them,
then make will identify which one has changed and
which ones depend on that change, and recompile
only those that are needed. But I don’t have to pro-
duce a separate line in the Makefile for each file.
Also, the Makefile provides implicit documenta-
tion for the flow of operations, so if I need to pass the
project on to someone else, all they need to do is call
make to get going.
More information on make can be found from
http://www.gnu.org/software/make/.
Andrew Robinson
Department of Mathematics and Statistics
University of Melbourne
Australia
A.Robinson@ms.unimelb.edu.au
R News ISSN 1609-3631

Vol. 7/1, April 2007 33
## Hashes indicate comments. Use these for documentation.
## Makefiles are most easily read from the bottom up, the first time!
# make is automatically interpreted as make report.pdf .
# Update report.pdf whenever report.tex changes, by running this code.
report.pdf : report.tex
( \
\pdflatex report.tex; \
while \grep -q "Rerun to get cross-references right." report.log;\
do \
\pdflatex report.tex; \
done \
)
# Update report.tex whenever report.rnw *or* output.RData changes,
# by running this code.
report.tex : report.rnw output.RData
echo "Sweave(\"report.rnw\")" | R --no-save --no-restore
# Update output.RData whenever sim.r changes, by running this code.
output.RData : sim.r
echo "reps <- 2000; source(\"sim.r\")" | R --no-save --no-restore
########################################################################
## The following section tells make how to respond to specific keywords.
.PHONY: test full neat clean
# make test cleans up, runs a small number of simulations,
# and then constructs a report
test:
make clean;
echo "reps <- 10; source(\"sim.r\")" | R --no-save --no-restore;
make
# make full cleans up and runs the whole project from scratch
full :
make clean;
make
# make neat cleans up temporary files - useful for archiving
neat :
rm -fr *.txt *.core fort.3 *~ *.aux *.log *.ps *.out
# make clean cleans up all created files.
clean :
rm -fr *.core fort.3 *~ *.exe *.tex *.txt *.lof *.lot *.tex \
*.RData *.ps *.pdf *.aux *.log *.out *.toc *.eps
Figure 1: The contents of a Makefile. Note that all the indenting is done by means of tab characters, not spaces.
This Makefile has been tested for both GNU make and BSD make.
R News ISSN 1609-3631
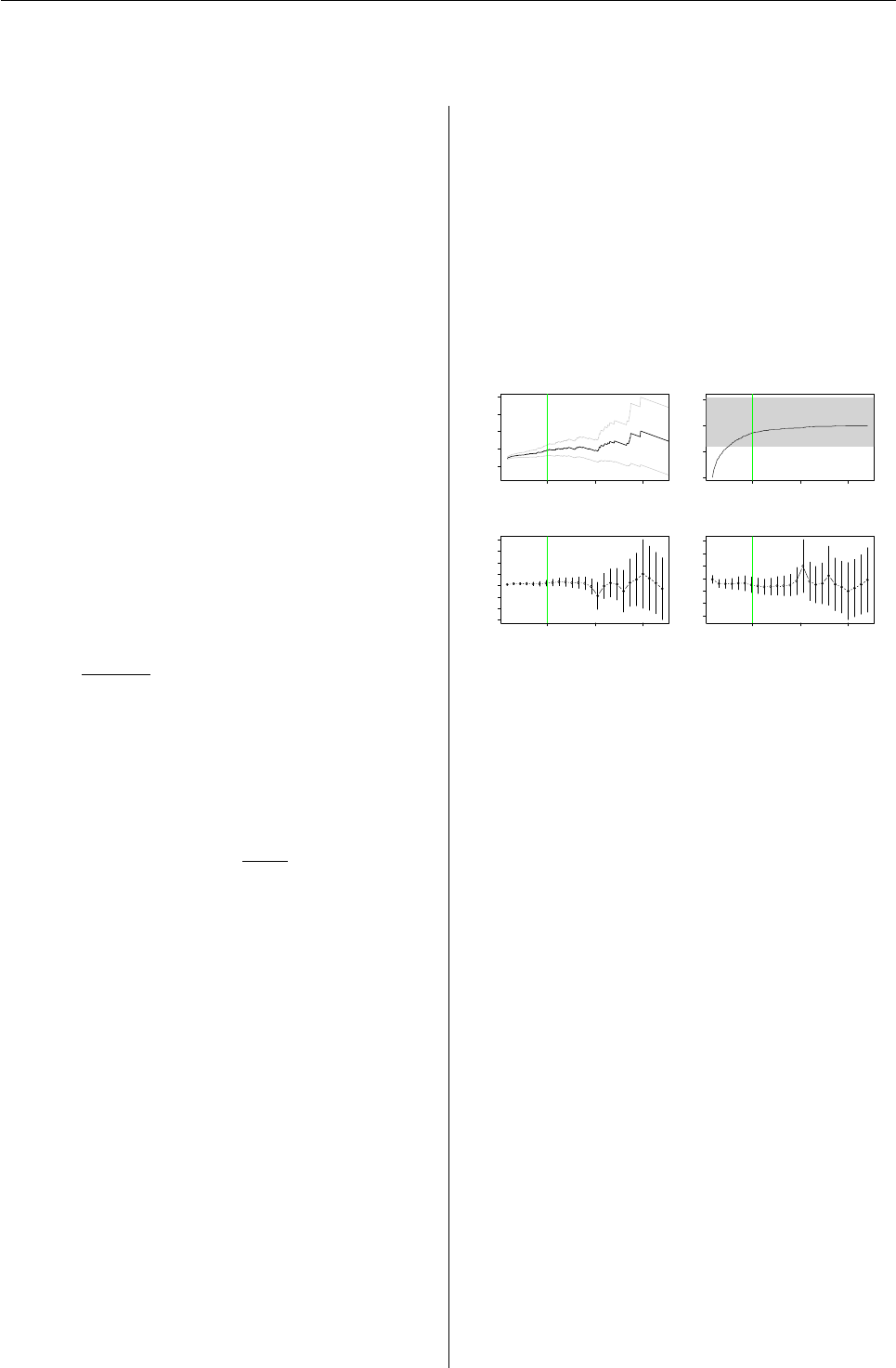
Vol. 7/1, April 2007 34
POT: Modelling Peaks Over a Threshold
by Mathieu Ribatet
The Generalised Pareto Distribution (GPD) is the
limiting distribution of normalised excesses over a
threshold, as the threshold approaches the endpoint
of the variable (Pickands,1975). The POT package
contains useful tools to perform statistical analysis
for peaks over a threshold using the GPD approxi-
mation.
There is many packages devoted to the extreme
value theory (evd,ismev,evir, . . . ); however, the
POT package is specialised in peaks over thresh-
old analysis. Moreover, this is currently the only
one which proposes many estimators for the GPD. A
user’s guide (as a package vignette) and two demos
are also included in the package.
Asymptotic Approximation
Let X1,...,Xnbe a series of i.id. random variables
with common distribution function F. Let Mn=
max {X1,...,Xn}. Suppose there exists constants
an>0 and bnsuch that:
IP Mn−bn
an
≤z−→ G(z),n→+∞
for z∈Rand where Gis a non degenerate distribu-
tion function. Then, for i=1, . . . , n, we have:
IP [Xi≤z|Xi>u]−→ H(z),u→uend (1)
with
H(y) = 1−1+ξy−µ
σ−1/ξ
+
,
where (µ,σ,ξ)are the location, scale and shape pa-
rameters respectively, σ>0 and z+=max(z, 0)and
uend is the right end-point of the variable Xi.
It is usual to fit the GPD to excesses over a
(high enough) threshold. Thus we suppose that the
asymptotic result given by equation (1) is (approxi-
mately) true for the threshold of interest.
Application:
Ardières River at Beaujeu
The ardieres data frame containing flood dis-
charges (in m3·s−1) over a period of 33 years of
the Ardières river at Beaujeau (FRANCE) is included
in the package. There are NA values in year 1994 as
a flood event damaged record instrumentation. We
use this dataset as an example for a typical univari-
ate analysis. First, we have to “extract” indepen-
dent events from the time series and select a suit-
able threshold such that asymptotic approximation
in equation (1) is good enough.
library("POT")
data("ardieres")
tmp <- clust(ardieres, 0.85, tim.cond = 7/365,
clust.max = TRUE)
par(mfrow=c(2,2))
mrlplot(tmp[,"obs"], xlim = c(0.85, 17))
diplot(tmp, u.range = c(0.85, 17))
tcplot(tmp[,"obs"], u.range = c(0.85, 17))
5 10 15
0 5 10 15 20
Mean Residual Life Plot
Threshold
Mean Excess
5 10 15
0.0 0.5 1.0 1.5
Dispersion Index Plot
Threshold
Dispersion Index
5 10 15
−30 −10 10 30
Threshold
Modified Scale
5 10 15
−1.0 0.0 1.0 2.0
Threshold
Shape
Figure 1: Tools for the threshold selection
The threshold selection stage is a compromise be-
tween bias and variance. On one hand, if a too
high threshold is selected, the bias decreases as the
asymptotic approximation in equation (1) is good
enough while the variance increases as there is not
enough data above this threshold. On the other
hand, by taking a lower threshold, the variance de-
creases as the number of observations is larger and
the bias increases as the asymptotic approximation
becomes poorer.
According to Fig. 1, a threshold around five
m3·s−1should be a “good compromise”. Indeed,
the mean residual life plot is “linear” on the range
(5, 7); for thresholds greater than 5, the dispersion
index estimates are “near” the theoretical value 1;
and both modified scale and shape estimates seem to
be constant on the range (5, 9). Thus, we select only
independent values above this threshold by invok-
ing:
events <- clust(ardieres, u = 5,
tim.cond = 7/365, clust.max = TRUE)
We can fit the GPD to those excesses accord-
ing several estimators by setting the method option.
There is currently 7 estimators: Moments "moments",
Unbiased and Biased Probability Weighted Moments
"pwmu", "pwmb", Minimum Density Power Diver-
gence "mdpd", medians "med", Pickands "pickands"
R News ISSN 1609-3631
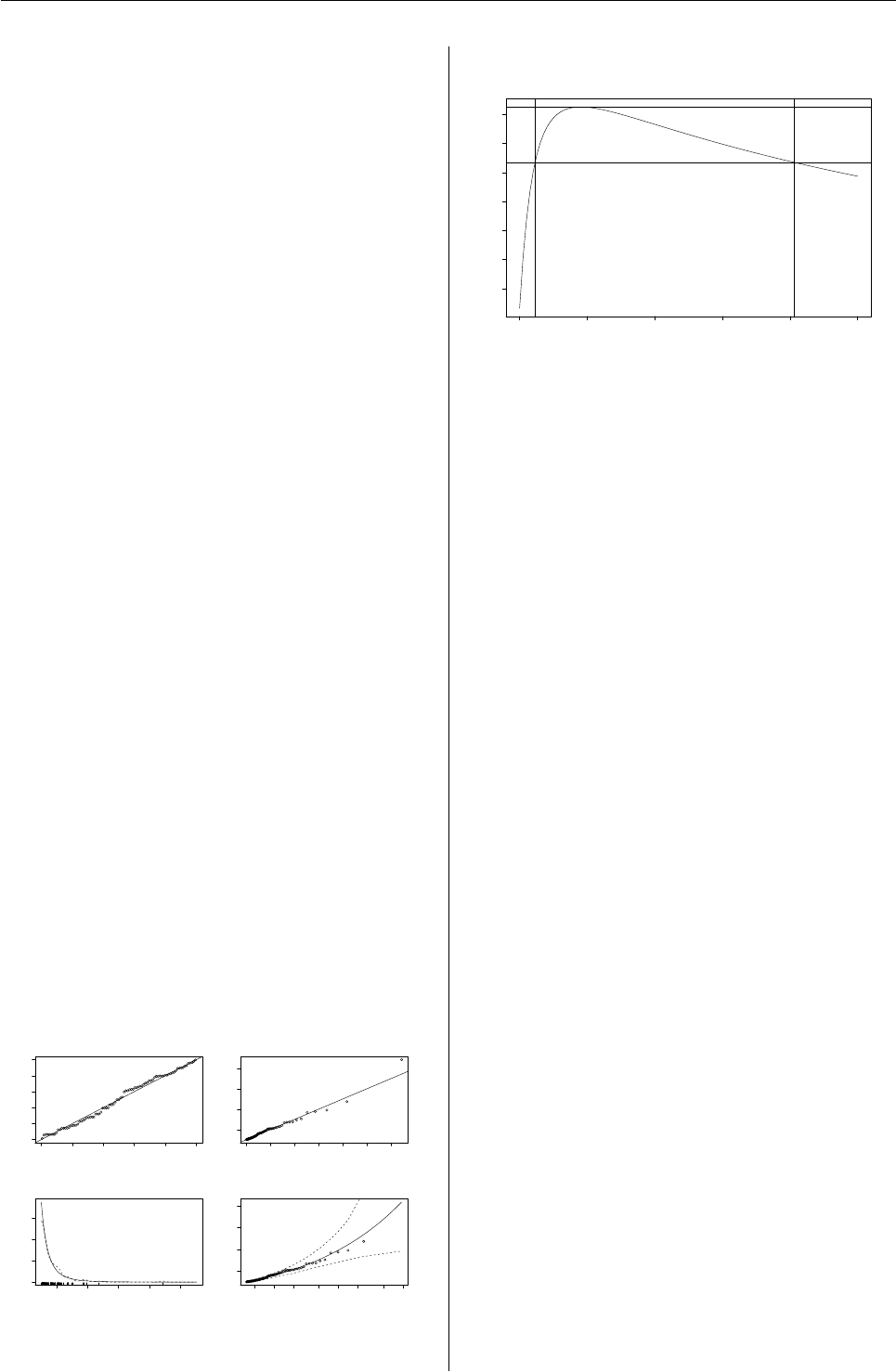
Vol. 7/1, April 2007 35
and Maximum Likelihood (the default) "mle" esti-
mators. References for these estimators can be found
in (Pickands,1975;Hosking and Wallis,1987;Coles,
2001;Peng and Welsh,2001) and (Juárez and Schu-
cany,2004). For example, if we want to fit the GPD
using the unbiased probability weighted moment es-
timator:
obs <- events[,"obs"]
pwmu <- fitgpd(obs, thresh = 5, "pwmu")
Here is the scale and shape parameter estimates
of the GPD for the 7 estimators implemented.
scale shape
mle 2.735991 0.2779359
mom 2.840792 0.2465661
pwmu 2.668368 0.2922964
pwmb 2.704665 0.2826697
mdpd 2.709254 0.2915759
med 2.135882 0.8939585
pick 2.328240 0.6648158
By invoking:
par(mfrow=c(2,2))
plot(pwmu, npy=2.63)
we obtain Fig. 2which depicts graphic tools for
model diagnostic. Profile likelihood confidence in-
tervals can also be computed, even for return levels
see Fig. 3, with:
gpd.pfrl(mle, 0.995, range = c(20, 120),
conf = 0.95)
0.0 0.2 0.4 0.6 0.8 1.0
0.0 0.4 0.8
Probability plot
Empirical
Model
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
5 10 15 20 25 30 35
10 20 30 40
QQ−plot
Model
Empirical
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−−−−−−−−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−
−−−−−−
−
10 20 30 40 50
0.0 0.1 0.2 0.3
Density Plot
Quantile
Density
0.5 1.0 2.0 5.0 20.0 50.0
10 20 30 40
Return Level Plot
Return Period (Years)
Return Level
Figure 2: Graphic tools for model diagnostic.
20 40 60 80 100 120
−205 −204 −203 −202 −201 −200 −199
Return Level
Profile Log−likelihood
Figure 3: 95% profile confidence interval for the re-
turn level associated to non exceedance probability
0.995.
Miscellaneous Features
The POT package can also:
• simulate and compute density, quantile and
distribution functions for the GPD;
• fit the GPD with a varying threshold using
MLE;
• fit the GPD with held fixed parameters using
MLE;
• perform analysis of variance for two nested
models;
• estimate the extremal index using two estima-
tors;
• display a L-Moment plot (Hosking and Wallis,
1997);
• compute sample L-moments;
• convert non exceedance probabilities to return
periods and vice-versa;
• compute “averaged” time series using an aver-
age mobile window.
Currently, most of the package developments are
devoted to bivariate peaks over threshold. For this
purpose, the POT package can also:
• fit a bivariate GPD using 6 parametric depen-
dence functions;
• fit a first order Markov chain with a fixed ex-
treme value dependence structure to all thresh-
old exceedances;
• simulate first order Markov chains with a fixed
extreme value dependence structure;
• plot the Pickands’ dependence and the spectral
density functions.
R News ISSN 1609-3631

Vol. 7/1, April 2007 36
Bibliography
S. Coles. An Introduction to Statistical Modelling of Ex-
treme Values. Springer Series in Statistics. London,
2001. 35
J. Hosking and J. Wallis. Parameter and quantile
estimation for the generalised Pareto distribution.
Technometrics, 29(3):339–349, 1987. 35
J. Hosking and J. Wallis. Regional Frequency Analysis.
Cambridge University Press, 1997. 35
S. Juárez and W. Schucany. Robust and efficient esti-
mation for the generalised Pareto distribution. Ex-
tremes, 7(3):237–251, 2004. ISSN 13861999. 35
L. Peng and A. Welsh. Robust estimation of the gen-
eralised Pareto distribution. Extremes, 4(1):53–65,
2001. 35
J. Pickands. Statistical inference using extreme order
statistics. Annals of Statistics, 3:119–131, 1975.
34,35
Mathieu Ribatet
Cemagref Unité de Recherche HH Lyon, France
ribatet@lyon.cemagref.fr
Backtests
by Kyle Campbell, Jeff Enos, Daniel Gerlanc and David
Kane
Introduction
The backtest package provides facilities for explor-
ing portfolio-based conjectures about financial in-
struments (stocks, bonds, swaps, options, et cetera).
For example, consider a claim that stocks for which
analysts are raising their earnings estimates perform
better than stocks for which analysts are lowering es-
timates. We want to examine if, on average, stocks
with raised estimates have higher future returns than
stocks with lowered estimates and whether this is
true over various time horizons and across different
categories of stocks. Colloquially, “backtest” is the
term used in finance for such tests.
Background
To demonstrate the capabilities of the backtest pack-
age we will consider a series of examples based on a
single real-world data set. StarMine1is a San Fran-
sisco research company which creates quantitative
equity models for stock selection. According to the
company:
StarMine Indicator is a 1-100 percentile rank-
ing of stocks that is predictive of future an-
alyst revisions. StarMine Indicator improves
upon basic earnings revisions models by:
• Explicitly considering management
guidance.
• Incorporating SmartEstimates,
StarMine’s superior estimates con-
structed by putting more weight on the
most accurate analysts.
• Using a longer-term (forward 12-month)
forecast horizon (in addition to the cur-
rent quarter).
StarMine Indicator is positively correlated to
future stock price movements. Top-decile
stocks have annually outperformed bottom-
decile stocks by 27 percentage points over the
past ten years across all global regions.
These ranks and other attributes of stocks are in
the starmine data frame, available as part of the
backtest package.
> data("starmine")
> names(starmine)
[1] "date" "id" "name"
[4] "country" "sector" "cap.usd"
[7] "size" "smi" "fwd.ret.1m"
[10] "fwd.ret.6m"
starmine contains selected attributes such as sec-
tor, market capitalisation, country, and various mea-
sures of return for a universe of approximately 6,000
securities. The data is on a monthly frequency from
January, 1995 through November, 1995. The number
of observations varies over time from a low of 4,528
in February to a high of 5,194 in November.
date count
1995-01-31 4593
1995-02-28 4528
1995-03-31 4569
1995-04-30 4708
1995-05-31 4724
1995-06-30 4748
1995-07-31 4878
1995-08-31 5092
1995-09-30 5185
1995-10-31 5109
1995-11-30 5194
1See www.starmine.com for details.
R News ISSN 1609-3631

Vol. 7/1, April 2007 37
The smi column contains the StarMine Indicator
score for each security and date if available. Here is
a sample of rows and columns from the data frame:
date name fwd.ret.1m fwd.ret.6m smi
1995-01-31 Lojack Corp 0.09 0.8 96
1995-02-28 Raymond Corp 0.05 0.1 85
1995-02-28 Lojack Corp 0.08 0.7 90
1995-03-31 Lojack Corp 0.15 1.0 49
1995-08-31 Supercuts Inc -0.11 -0.5 57
1995-10-31 Lojack Corp -0.40 -0.2 22
1995-11-30 Lojack Corp 0.20 0.4 51
Most securities (like LoJack above) have multiple
entries in the data frame, each for a different date.
The row for Supercuts indicates that, as of the close of
business on August 31, 1995, its smi was 57. During
the month of September, its return (i.e., fwd.ret.1m)
was -11%.
A simple backtest
Backtests are run by calling the function backtest to
produce an object of class backtest.
> bt <- backtest(starmine, in.var = "smi",
+ ret.var = "fwd.ret.1m")
starmine is a data frame containing all the infor-
mation necessary to conduct the backtest. in.var
and ret.var identify the columns containing the
input and return variables, respectively. backtest
splits observations into 5 (the default) quantiles, or
“buckets,” based on the value of in.var. Lower
(higher) buckets contain smaller (larger) values of
in.var. Each quantile contains an approximately
equal number of observations. This backtest creates
quantiles according to values in the smi column of
starmine.
[1,21] (21,40] (40,59] (59,82] (82,100]
6765 6885 6642 6600 6496
backtest calculates the average return within
each bucket. From these averages we calculate the
spread, or the difference between the average return
of the highest and lowest buckets.
Calling summary on the resulting object of class
backtest reports the in.var,ret.var, and by.var
used. We will use a by.var in later backtests.
> summary(bt)
Backtest conducted with:
1 in.var: smi;
1 ret.var: fwd.ret.1m;
and no by.var.
low 2 3 4 high spread
pooled 0.011 0.013 0.016 0.02 0.032 0.021
This backtest is an example of a pooled backtest. In
such a backtest, we assume that all observations are
exchangeable. This means that a quantile may con-
tain observations for any stock and from any date.
Quantiles may contain multiple observations for the
same stock.
The backtest summary shows that the average re-
turn for the highest bucket was 3.2%. This value is
the mean one month forward return of stocks with
smi values in the highest quantile. As the obser-
vations are exchangeable, we use every observation
in the starmine data frame with a non-missing smi
value. This means that the returns for LoJack from
both 1995-01-31 and 1995-02-28 would contribute to
the 3.2% mean of the high bucket.
The backtest suggests that StarMine’s model pre-
dicted performance reasonably well. On average,
stocks in the highest quantile returned 3.2% while
stocks in the lowest quantile returned 1.1%. The
spread of 2.1% suggests that stocks with high ratings
perform better than stocks with low ratings.
Natural backtests
Anatural backtest requires that the frequency of re-
turns and observations be the same.
A natural backtest approximates the following
implementation methodology: in the first period
form an equal weighted portfolio with long posi-
tions in the stocks in the highest quantile and short
positions in the stocks in the lowest quantile. Each
stock has an equal weight in the portfolio; if there
are 5 stocks on the long side, each stock has a weight
of 20%. Subsequently rebalance the portfolio every
time the in.var values change. If the observations
have a monthly frequency, the in.var values change
monthly and the portfolio must be rebalanced ac-
cordingly. When the in.var values change, rebalanc-
ing has the effect of exiting positions that have left
the top and bottom quantiles and entering positions
that have entered the top and bottom quantiles. If the
data contains monthly observations, we will form 12
portfolios per year.
To create a simple natural backtest, we again call
backtest using fwd.ret.1m. This is the only return
value in starmine for which we can construct a nat-
ural backtest of smi.
> bt <- backtest(starmine, id.var = "id",
+ date.var = "date", in.var = "smi",
+ ret.var = "fwd.ret.1m", natural = TRUE)
Natural backtests require a date.var and id.var,
the names of the columns in the data frame contain-
ing the dates of the observations and unique security
identifiers, respectively. Calling summary displays
the results of the backtest:
> summary(bt)
R News ISSN 1609-3631

Vol. 7/1, April 2007 38
Backtest conducted with:
1 in.var: smi;
1 ret.var: fwd.ret.1m;
and no by.var.
low 2 3 4 high spread
1995-01-31 0.003 0.011 0.003 -0.0001 0.019 0.016
1995-02-28 -0.008 -0.003 0.003 0.0072 0.013 0.021
1995-03-31 0.029 0.017 0.013 0.0225 0.037 0.008
1995-04-30 -0.002 -0.003 0.002 -0.0054 0.005 0.007
1995-05-31 0.010 0.013 0.019 0.0228 0.044 0.034
1995-06-30 0.072 0.059 0.057 0.0708 0.101 0.030
1995-07-31 0.033 0.030 0.034 0.0323 0.052 0.018
1995-08-31 -0.004 0.006 0.017 0.0119 0.024 0.028
1995-09-30 -0.055 -0.030 -0.031 -0.0219 -0.014 0.041
1995-10-31 0.030 0.032 0.040 0.0430 0.038 0.008
1995-11-30 0.013 0.016 0.021 0.0294 0.037 0.024
MEAN 0.011 0.014 0.016 0.0193 0.032 0.021
average turnover: 0.5
mean spread: 0.02
sd spread: 0.01
raw sharpe ratio: 2
Focus on the mean return of the highest quan-
tile for 1995-02-28 of 1.3%. backtest calculated this
value by first computing the 5 quantiles of the in-
put variable smi over all observations in starmine.
Among the observations that fall into the highest
quantile, those with date 1995-02-28 contribute to the
mean return of 1.3%. It is important to note that
the input variable quantiles are computed over the
whole dataset, as opposed to within each category
that may be defined by a date.var or by.var.
The bottom row of the table contains the mean
quantile return over all dates. On account of the
way we calculate quantile means, a single stock will
have more effect on the quantile mean if during that
month there are fewer stocks in the quantile. Sup-
pose that during January there are only 2 stocks in
the low quantile. The return of a single stock in Jan-
uary will account for 1
22 of the quantile mean. This
is different than a pooled backtest where every ob-
servation within a quantile has the same weight. In
a natural backtest, the weight of a single observation
depends on the number of observations for that pe-
riod.
Calling summary yields information beyond that
offered by the summary method of a pooled back-
test. The first piece of extra information is average
turnover. Turnover is the percentage of the portfolio
we would have to change each month if we imple-
mented the backtest as a trading strategy. For exam-
ple, covering all the shorts and shorting new stocks
would yield a turnover of 50% because we changed
half the portfolio. We trade stocks when they enter
or exit the extreme quantiles due to in.var changes.
On average, we would turn over 50% of this portfolio
each month.
The second piece of extra information is mean
spread. The spread was positive each month, so on
average the stocks with the highest smi values out-
performed the stocks with the lowest smi values.
On average, stocks in the highest quantile outper-
formed stocks in the lowest quantile by 2%. The
third piece of extra information, the standard devia-
tion of spread, is 1%. The spread varied from month
to month, ranging from a low of close to 0% to a high
of over 4%.
We define the fourth piece of extra information,
raw (non-annualized) Sharpe ratio, as return
risk . We set
return equal to mean spread return and use the stan-
dard deviation of spread return as a measure of risk.
More than one in.var
backtest allows for more than one in.var to be
tested simultaneously. Besides using smi, we will
test market capitalisation in dollars, cap.usd. This
is largely a nonsense variable since we do not expect
large cap stocks to outperform small cap stocks — if
anything, the reverse is true historically.
> bt <- backtest(starmine, id.var = "id",
+ date.var = "date", in.var = c("smi",
+ "cap.usd"), ret.var = "fwd.ret.1m",
+ natural = TRUE)
Because more than one in.var was specified,
only the spread returns for each in.var are dis-
played, along with the summary statistics for each
variable.
> summary(bt)
Backtest conducted with:
2 in.vars: smi, cap.usd;
1 ret.var: fwd.ret.1m;
and no by.var.
smi cap.usd
1995-01-31 0.016 -0.0138
1995-02-28 0.021 0.0017
1995-03-31 0.008 -0.0023
1995-04-30 0.007 -0.0052
1995-05-31 0.034 -0.0568
1995-06-30 0.030 -0.0143
1995-07-31 0.018 -0.0008
1995-08-31 0.028 0.0051
1995-09-30 0.041 0.0321
1995-10-31 0.008 0.0127
1995-11-30 0.024 0.0029
summary stats for in.var = smi:
average turnover: 0.5
mean spread: 0.02
sd spread: 0.01
raw sharpe ratio: 2
summary stats for in.var = cap.usd:
average turnover: 0.1
mean spread: -0.004
sd spread: 0.02
raw sharpe ratio: -0.2
R News ISSN 1609-3631
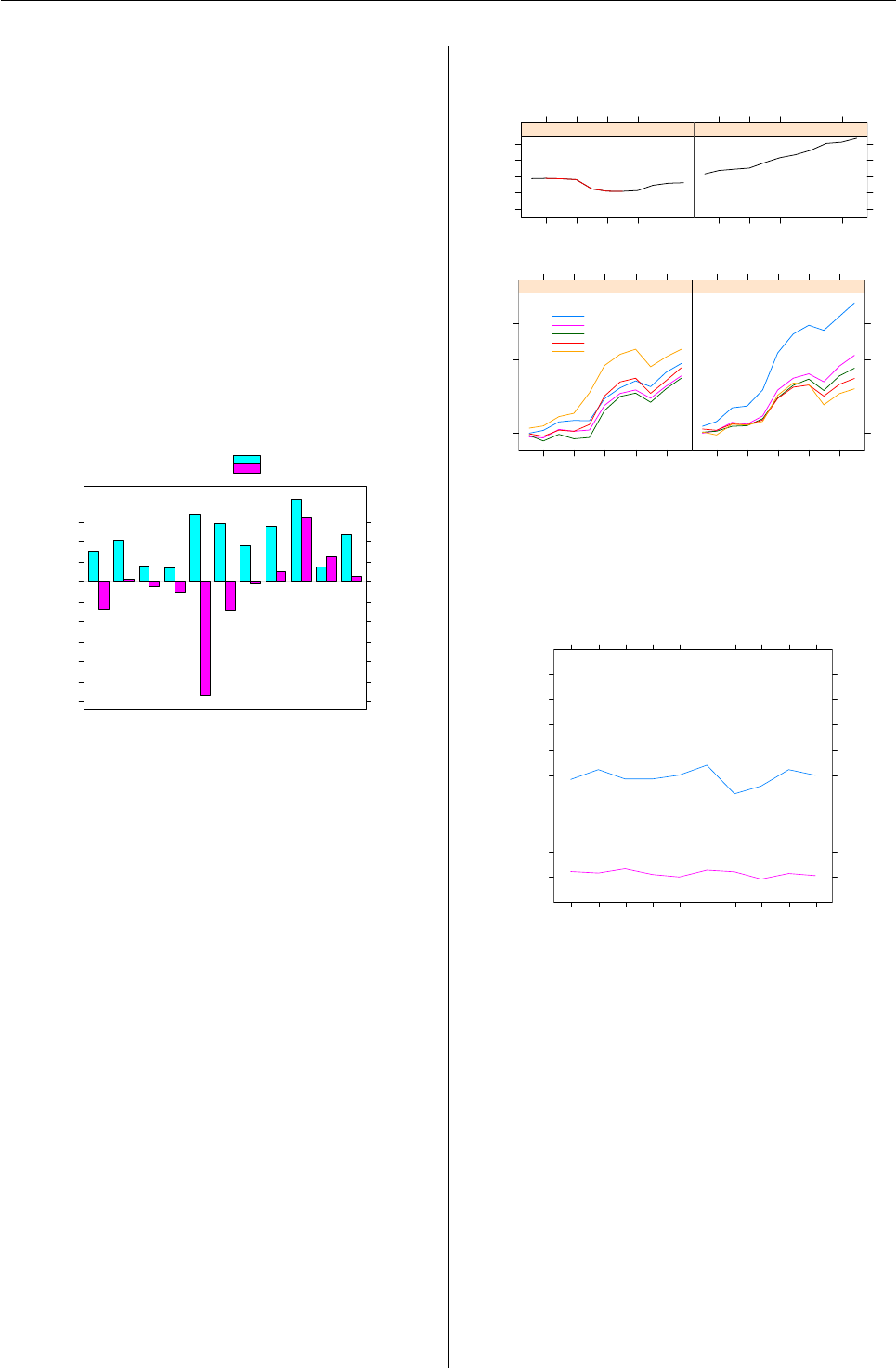
Vol. 7/1, April 2007 39
Viewing the results for the two input variables
side-by-side allows us to compare their performance
easily. As we expected, cap.usd as an input variable
did not perform as well as smi over our backtest pe-
riod. While smi had a positive return during each
month, cap.usd had a negative return in 6 months
and a negative mean spread. In addition, the spread
returns for cap.usd were twice as volatile as those of
smi.
There are several plotting facilities available in
backtest that can help illustrate the difference in
performance between these two signals. These plots
can be made from a natural backtest with any num-
ber of input variables. Below is a bar chart of the
monthly returns of the two signals together:
> plot(bt, type = "return")
Spread Return
Date
Return (%)
−6
−5
−4
−3
−2
−1
0
1
2
3
4
1995−01−31
1995−02−28
1995−03−31
1995−04−30
1995−05−31
1995−06−30
1995−07−31
1995−08−31
1995−09−30
1995−10−31
1995−11−30
smi
cap.usd
Figure 1: Monthly return spreads.
Returns for smi were consistently positive. Re-
turns for cap.usd were of low quality, but improved
later in the period. cap.usd had a particularly poor
return in June. We can also plot cumulative returns
for each input variable as shown in Figure 2.
The top region in this plot shows the cumulative
return of each signal on the same return scale, and
displays the total return and worst drawdown of the
entire backtest period. The bottom region shows the
cumulative return of the individual quantiles over
time. We can see that smi’s top quantile performed
best and lowest quantile performed worst. In con-
trast, cap.usd’s lowest quantile was its best perform-
ing.
Though it is clear from the summary above that
smi generated about 5 times as much turnover as
cap.usd, a plot is available to show the month-by-
month turnover of each signal, see Figure 3. This
chart shows that the turnover of smi was consistently
around 50% with lower turnover in September and
October, while the turnover of cap.usd was consis-
tently around 10%.
> plot(bt.save, type = "cumreturn.split")
Cumulative Spread Return
Return (%)
−20
−10
0
10
20
Mar May Jul Sep Nov
Total return: −3.9
Worst drawdown: −8
cap.usd
Mar May Jul Sep Nov
Total return: 23
Worst drawdown: −−
smi
Cumulative Quantile Return
Date
Return (%)
0
10
20
30
Mar May Jul Sep Nov
cap.usd
Mar May Jul Sep Nov
smi
quantile
high
Q4
Q3
Q2
low
Figure 2: Cumulative spread and quantile returns.
> plot(bt, type = "turnover")
Turnover
Date
Turnover (%)
10
20
30
40
50
60
70
80
90
Mar Apr May Jun Jul Aug Sep Oct Nov Dec
●
●
●●●
●
●
●
●
●
●●●
●●
●●
●
●●
smi
cap.usd
●
●
Figure 3: Monthly turnover.
Using by.var
In another type of backtest we can look at quan-
tile spread returns by another variable. Specifying
by.var breaks up quantile returns into categories de-
fined by the levels of the by.var column in the input
data frame. Consider a backtest of smi by sector:
> bt <- backtest(starmine, in.var = "smi",
+ ret.var = "fwd.ret.1m",
+ by.var = "sector")
> summary(bt)
Backtest conducted with:
R News ISSN 1609-3631

Vol. 7/1, April 2007 40
1 in.var: smi;
1 ret.var: fwd.ret.1m;
and by.var: sector.
low 2 3 4 high spread
Durbl 0.0063 0.007 0.009 0.007 0.01 0.004
Enrgy 0.0152 0.014 0.017 0.019 0.04 0.024
HiTec 0.0237 0.016 0.026 0.029 0.05 0.024
Hlth 0.0395 0.036 0.021 0.038 0.05 0.006
Manuf 0.0005 0.005 0.014 0.009 0.02 0.022
Money 0.0190 0.024 0.021 0.026 0.04 0.017
NoDur 0.0036 0.010 0.010 0.019 0.03 0.025
Other 0.0045 0.006 0.015 0.017 0.02 0.017
Shops 0.0020 0.004 0.005 0.017 0.03 0.026
Telcm 0.0277 0.014 0.022 0.023 0.03 0.005
Utils 0.0128 0.021 0.013 0.016 0.02 0.007
This backtest categorises observations by the
quantiles of smi and the levels of sector. The high-
est spread return of 2.6% occurs in Shops. Since
smi quantiles were computed before the observa-
tions were split into groups by sector, however, we
can not be sure how much confidence to place in
this result. There could be very few observations in
this sector or one of the top and bottom quantiles
could have a disproportionate number of observa-
tions, thereby making the return calculation suspect.
counts provides a simple check.
> counts(bt)
$smi
low 2 3 4 high
Durbl 348 349 261 231 223
Enrgy 246 250 158 130 64
HiTec 647 660 824 1004 1432
Hlth 380 377 410 464 424
Manuf 1246 1265 1279 1395 1576
Money 959 1265 1244 1095 875
NoDur 615 563 528 441 371
Other 1034 940 784 760 710
Shops 870 714 710 697 548
Telcm 186 177 140 129 95
Utils 152 245 252 198 130
While there seems to be an adequate number of
observations in Shops, it is important to note that
there are approximately 60% more observations con-
tributing to the mean return of the lowest quantile
than to the mean return of the highest quantile, 870
versus 548. Overall, we should be more confident in
results for Manuf and Money due to their larger sam-
ple sizes. We might want to examine the result for
HiTec more closely, however, since there are more
than twice the number of observations in the high-
est quantile than the lowest.
by.var can also be numeric, as in this backtest us-
ing cap.usd:
> bt <- backtest(starmine,
+ in.var = "smi", ret.var = "fwd.ret.1m",
+ by.var = "cap.usd",
+ buckets = c(5, 10))
> summary(bt)
Backtest conducted with:
1 in.var: smi;
1 ret.var: fwd.ret.1m;
and by.var: cap.usd.
low 2 3 4 high spread
low 0.0105 0.0139 0.0236 0.028 0.038 0.028
2 0.0078 0.0093 0.0216 0.025 0.046 0.038
3 0.0186 0.0072 0.0167 0.031 0.034 0.016
4 0.0124 0.0142 0.0139 0.013 0.038 0.026
5 0.0080 0.0124 0.0087 0.010 0.025 0.017
6 0.0126 0.0121 0.0191 0.021 0.026 0.013
7 0.0080 0.0070 0.0160 0.019 0.034 0.026
8 0.0050 0.0181 0.0101 0.014 0.027 0.022
9 0.0104 0.0153 0.0167 0.014 0.028 0.018
high 0.0156 0.0207 0.0133 0.023 0.026 0.011
Since cap.usd is numeric, the observations are
now split by two sets of quantiles. Those listed
across the top are, as before, the input variable quan-
tiles of smi. The row names are the quantiles of
cap.usd. The buckets parameter of backtest con-
trols the number of quantiles. The higher returns
in the lower quantiles of cap.usd suggests that smi
performs better in small cap stocks than in large cap
stocks.
Multiple return horizons
Using backtest we can also analyse the performance
of a signal relative to multiple return horizons. Be-
low is a backtest that considers one month and six
month forward returns together:
> bt <- backtest(starmine, in.var = "smi",
+ buckets = 4, ret.var = c("fwd.ret.1m",
+ "fwd.ret.6m"))
> summary(bt)
Backtest conducted with:
1 in.var: smi;
2 ret.vars: fwd.ret.1m, fwd.ret.6m;
and no by.var.
low 2 3 high spread
fwd.ret.1m 0.011 0.015 0.018 0.03 0.019
fwd.ret.6m 0.112 0.121 0.142 0.17 0.059
The performance of smi over these two return
horizons tells us that the power of the signal de-
grades after the first month. Using six month for-
ward return, fwd.ret.6m, the spread is 6%. This
is only 3 times larger than the 2% spread return in
the first month despite covering a period which is 6
times longer. In other words, the model produces 2%
spread returns in the first month but only 4% in the 5
months which follow.
R News ISSN 1609-3631

Vol. 7/1, April 2007 41
Conclusion
The backtest package provides a simple collection of
tools for performing portfolio-based tests of financial
conjectures. A much more complex package, port-
folioSim, provides facilities for historical portfolio
performance analysis using more realistic assump-
tions. Built on the framework of the portfolio2pack-
age, portfolioSim tackles the issues of risk exposures
and liquidity constraints, as well as arbitrary port-
folio construction and trading rules. Above all, the
flexibility of R itself allows users to extend and mod-
ify these packages to suit their own needs. Before
reaching that level of complexity, however, backtest
provides a good starting point for testing a new con-
jecture.
Bibliography
J. Enos and D. Kane. Analysing equity portfolios
in R. R News, 6(2):13–19, MAY 2006. URL http:
//CRAN.R-project.org/doc/Rnews.41
Kyle Campbell, Jeff Enos, Daniel Gerlanc and David Kane
Kane Capital Management
Cambridge, Massachusetts, USA
Kyle.W.Campbell@williams.edu,jeff@kanecap.com,
dgerlanc@gmail.com and david@kanecap.com
Review of John Verzani’s Book
Using R for Introductory Statistics
Andy Liaw
To the best of my knowledge, this
book is the first of its kind: a stan-
dalone introductory statistics text-
book that integrates R throughout.
The advantages should be obvi-
ous: Students would not need to
purchase a supplement that cov-
ers the software, in addition to the
main textbook (although the au-
thor states in the Preface that the
book should also be useful as an accompaniment for
a standard introductory text). That the software is
freely available is a big bonus. Moreover, the book
covers basic descriptive statistics before any proba-
bility models are mentioned. For students that are
less mathematically inclined, this should make mate-
rials easier to absorb. (The author states in the Pref-
ace that the book aims at classes that are based on
pre-calculus skills.)
The book contains 12 chapters. The first four
chapters of the book cover descriptive statistics, both
numerical and graphical, from general introduction
(Chapter 1), through univariate and bivariate data
(Chapters 2 and 3) to multivariate data (Chapter 4).
Each chapter covers both categorical and numeri-
cal data. The author chose to treat two indepen-
dent samples as bivariate data and several indepen-
dent samples as multivariate data, which I think is
a bit unusual. Chapter 5 covers probability models.
Chapter 6 covers simulations, setting up for the top-
ics on inference in the chapters that follow. Chap-
ters 7 and 8 cover confidence intervals and signif-
icance tests, respectively. Chapter 9 discusses the
χ2tests for the multinomial distribution, the test
for independence, and goodness-of-fit tests such as
Kolmogorov-Smirnov and Shapiro-Wilk. Chapter 10
covers both simple and multiple linear regression.
Chapter 11 covers one- and two-way ANOVA as well
as ANCOVA. Chapter 12 covers logistic and nonlin-
ear regression. There are also five appendices that
cover various aspects of R (installation, GUI, teach-
ing with R, graphics, programming). Throughout the
book, examples of R usage are interspersed among
the main text, and some sections devoted to R top-
ics are introduced as the need arises (e.g., in Chap-
ter 6, Simulations, Section 6.2 covers for() loops).
Data used as examples were drawn from a wide va-
riety of areas. Exercises are given at the end of sec-
tions (rather than chapters). The book also has an
accompanying add-on package, UsingR (available on
CRAN), which contains data sets and some functions
used in the book. The book also has a web site that
contains answers to selected problems, the UsingR
package for various platforms (including one for S-
PLUS), as well as errata.
Several ideas presented in the book deserve acco-
lades (e.g., covering EDA before introducing proba-
bility models, coverage of robust/resistant methods,
thorough integration of R into the materials). How-
ever, there are also drawbacks. The most glaring one
is the fact that many rather technical terms are used
before they are introduced or explained, and some
are not sufficiently elaborated. (E.g., “density” is first
used to show how a kernel density estimate can be
added to a histogram, but no explanation was given
for what a density is or what it means.) In my teach-
ing experience, one of the most difficult (but abso-
lutely essential) concepts for students to grasp is the
2See Enos and Kane (2006) for an introduction to the portfolio package.
R News ISSN 1609-3631

Vol. 7/1, April 2007 42
idea of the sampling distribution of a statistic, yet
this did not receive nearly the attention I believe it
deserves. The discussion of scatterplot smoothing
(Section 3.4.7, “Trend lines”) gave a minimal descrip-
tion of what smoothers are available in base R, such
as smoothing splines, loess, and Friedman’s super-
smoother. I would be surprised if students in intro
stat courses are not completely bewildered by such
a minimal description. This will make it harder for
some students to follow the text along on their own.
Some people might be interested in learning how
to do basic data analysis using R. As these people
are not among the intended audience, this book may
not serve them as nicely as others, because the R-
specific topics are scattered throughout the book in
bits and pieces, each making its entrance as the sta-
tistical topic being covered requires it.
Now, some real nit-picking on more esoteRic
things: The author seems to use “library” and “pack-
age” interchangeably, which could make some R
users cringe. Also, on page 8, the virtue of using <-
is touted, but the author still decides to use =for the
book, without explanation. I also found the mention
of the (evil?) <<- wholely unnecessary: The author
said that it may be useful in programming R, yet per-
sonally I have not used it in the years I have been
programming R. At the level the book is intended,
I believe the students would be better served by its
exclusion.
In summary, I like the structure of the book very
much. However, the various problems mentioned
above keep me from giving it a whole-hearted rec-
ommendation as a standalone text. It may serve well
as a supplementary text for a more standard intro-
ductory Statistics textbook (a la Peter Dalgaard’s “In-
troductory Statistics with R”).
Andy Liaw
Merck Research Laboratories
andy_liaw@merck.com
DSC 2007
by Hadley Wickham
The fifth “Directions in Statistical Computing” con-
ference was held in sunny Auckland, New Zealand,
8–10 February 2007. Despite the distant location,
over 80 attendees from around the world made it to
the conference, to enjoy the interesting talks, fabu-
lous weather and the great food and drink of Auck-
land, including a fantastic ocean-side conference din-
ner.
The five key note speakers presented five quite
different looks at the future of statistical computing:
• Ross Ihaka (re-)explored the use of LISP as a
computational backend to R. He said, repeat-
edly, that this was just for fun, and will not be
the next R(Q?)
• L. Fraser Jackson demonstrated the use of J(an
open source grand child of APL) for statistics.
A very concise and powerful language.
• Patrick Wessa gave an interesting talk about
costs of the current scientific publishing frame-
work and how we can do better.
• Olivia Lau discussed zelig, a modeling frame-
work for R, which provides a common struc-
ture for modeling functions, making it easier
for users and developers.
• Duncan Temple Lang talked how about a more
composable object orientated Rcore (the code,
not the people) would make it easier to exper-
iment and explore new ideas, and continue to
keep Rrelevant for the future.
A complete programme and abstracts are avail-
able on the conference website, http://www.stat.
auckland.ac.nz/dsc-2007/. Conference papers
will be published in a special issue of Computational
Statistics.
A big round of thanks goes to the organizing
committee, Paul Murrell, Ross Ihaka, David Scott
and Thomas Yee, for such a great conference, as well
as to Sharon Walker and Suryashobha Herle for feed-
ing the hungry attendees.
Hadley Wickham
Department of Statistics
Iowa State University
Ames, Iowa, U.S.A.
h.wickham@gmail.org
R News ISSN 1609-3631

Vol. 7/1, April 2007 43
New Journal: Annals of Applied Statistics
Bradley Efron, Stanford University, USA
The Annals of Applied Statistics, newest journal
from the Institute of Mathematical Statistics, began
accepting papers on November 1, 2006. Computa-
tional statistics is an area of particular interest. Please
visit the journal website at http://www.imstat.org/
aoas for further information.
Forthcoming Events: useR! 2007
by Dianne Cook
The first North American useR! will be held at Iowa
State University, Ames, Iowa, August 8–10, 2007,
which will be a week after JSM’07.
This follows successful meetings in Vienna, Aus-
tria, in 2006 and 2004, and also Directions in Sta-
tistical Computing (DSC) meetings in Auckland,
NZ (Feb 2007), Seattle (2005), and Vienna (1999,
2001, 2003). Information about the meeting can be
found at http://www.useR2007.org/ or by emailing
useR2007@iastate.edu.
The important dates to remember are:
• April 23, 2007: Deadline for paper submission,
with referees comments returned by April 30.
• May 1, 2007: Early registration ends
• June 30: Poster abstract submission ends
• August 8, 2007: useR! 2007 begins!
We look forward to meet you in Ames!
On behalf of the organizing committee:
Diane Cook
Department of Statistics
Iowa State University
Ames, Iowa, U.S.A.
dicook@iastate.edu
Changes in R 2.5.0
by the R Core Team
User-visible changes
•apropos(x) and find(x) now both only work
for character x, and hence drop all non-
standard evaluation behaviour.
• Data frames can have ‘automatic’ row names
which are not converted to dimnames by
as.matrix(). (Consequently, e.g., t() for such
data frames has NULL column names.) This
change leads to memory reductions in several
places, but can break code which assumes char-
acter dimnames for data frames derived from
matrices.
No existing R object is regarded as having ‘au-
tomatic’ row names, and it may be beneficial
to recreate such objects via read.table() or
data.frame().
• Using $on an atomic vector now raises a warn-
ing, as does use on an S4 class for which a
method has not been defined.
• The Unix-alike readline terminal interface now
does command-completion for R objects, incor-
porating the functionality formerly in package
rcompletion by Deepayan Sarkar. This can be
disabled by setting the environment variable
R_COMPLETION to FALSE when starting R (e.g.,
in ‘∼/.Renviron’).
New features
•abbreviate() no longer has an 8191 byte limit
on the size of strings it can handle.
•abs(x) now returns integer for integer or logi-
cal arguments.
•apropos() has a new argument ignore.case
which defaults to TRUE, potentially matching
more than previously, thanks to a suggestion
by Seth Falcon.
•args(),str() and print() now give the argu-
ment lists of primitive functions.
•as.matrix() gains the ... argument that sev-
eral packages have assumed it always had (and
S-PLUS has).
• Manipulation of integers as roman numerals
via as.roman() in package utils.
R News ISSN 1609-3631

Vol. 7/1, April 2007 44
•attr() no longer treats name = NA_character_
as meaning name = "NA".
•binom.test() now allows a ‘fuzz’ for calcu-
lated integer values in its xand narguments.
•boxplot(*, notch = TRUE) now warns when
notches are outside hinges; related to PR#7690.
• New function callCC() providing a
downward-only version of Scheme’s call with
current continuation.
•capabilities() now has a profmem entry in-
dicating whether R has been compiled with
memory profiling.
•colnames<-() and rownames<-() now handle
data frames explicitly, so calling colnames<- on
a data frame no longer alters the representation
of the row names.
•commandArgs() has a new trailingOnly argu-
ment to be used in conjunction with ‘--args’.
•contour() now passes graphical parameters in
... to axis() and box().
• New data set crimtab on Student (1908)’s 3000
criminals.
•cut.default() has a new argument
ordered_result.
•.deparseOpts() has two new options: keepNA
to ensure that different types (logical, integer,
double, character and complex) of NAs are dis-
tinguished, and S_compatible to suppress the
use of R-specific features such as 123L and to
deparse integer values of a double vector with
a trailing decimal point.
The keepInteger option now uses the suffix L
rather than as.integer() where possible (un-
less all entries are NA or S_compatible is also
set).
Other deparse options can now be added to all
(which has not for some time actually switched
on all options).
Integer sequences m:n are now deparsed in that
form.
•deparse() and dput() now include
keepInteger and keepNA in their defaults for
the control argument.
•detach() now takes another argument,
unload, which indicates whether or not to un-
load the package and then only cleans up the S4
methods if the package successfully unloads.
• There are new constants NA_integer_,
NA_real_,NA_complex_ and NA_character_
to denote NAs of those types, and they
will be used in deparsing in place of
as.integer(NA) etc unless .deparseOpts()
includes S_compatible.
•dev.print() now recognizes screen devices
as all those with an enabled display list, rather
than a hard-coded set.
• Objects of class difftime are now handled
more flexibly. The units of such objects can
now be accessed via a units() function, which
also has a replacement form, and there are con-
version methods to and from numeric, which
also allow the specification of units. Objects of
this class can also be stored in data frames now.
Aformat() method has been added, and the
print method was revised.
• New function environmentName() to give
the print name of environments such as
‘namespace:base’. This is now used by str().
• New function env.profile() provides R level
access to summary statistics on environments.
In a related patch, new.env() now allows the
user to specify an initial size for a hashed envi-
ronment.
•file() can read the X11 clipboard selection
as "X11_clipboard" on suitable X11-using sys-
tems.
•file("stdin") is now recognized, and refers
to the process’s stdin file stream whereas
stdin() refers to the console. These may dif-
fer, for example for a GUI console, an embed-
ded application of R or if ‘--file=’ has been
used.
•file_test() is now also available in package
utils. (It is now private in package tools.)
•file.show() gains an encoding argument.
• New functions formatUL() and formatOL()
in package utils for formatting unordered
(itemize) and ordered (enumerate) lists.
• The statistics reported when gcinfo(TRUE) are
now of the amounts used (in Mb) and not of
the amounts free (which are not really relevant
when there are no hard limits, only gc trigger
points).
• New function get_all_vars() to retrieve all
the (untransformed) variables that the default
method of model.frame() would use to create
the model frame.
•interaction() has a new argument
lex.order.
R News ISSN 1609-3631

Vol. 7/1, April 2007 45
•initialize() (in methods) now tries to be
smarter about updating the new instance in
place, thereby reducing copying.
•install.packages(dependencies = NA) is a
new default, which is to install essential de-
pendencies when installing from repositories
to a single library. As a result of this change,
update.packages() will install any new de-
pendencies of the packages it is updating
(alongside the package in the same library
tree).
If lib is not specified or is specified of length
one and the chosen location is not a writable
directory, install.packages() offers to create
a personal library directory for you if one does
not already exist, and to install there.
•is.atomic,is.call,is.character,
is.complex,is.double (= is.real),
is.environment,is.expression,is.function,
is.integer,is.list,is.logical,is.null,
is.object,is.pairlist,is.recursive,
is.single and is.symbol (= is.name) are no
longer internally S3 generic, nor can S4 meth-
ods be written for them.
The factor methods of is.integer and
is.numeric have been replaced by internal
code.
• Added is.raw() for completeness.
•l10n_info() also reports if the current locale is
Latin-1.
•levels<-(),names() and names<-() now dis-
patch internally for efficiency and so no longer
have S3 default methods.
•.libPaths() now does both tilde and glob ex-
pansion.
• Functions lm(),glm() loess(),xtabs() and
the default method of model.frame() coerce
their formula argument (if supplied) to a for-
mula.
•max(),min() and range() now work with
character vectors.
•message() has a new argument appendLF
to handle messages with and without new-
lines. There is a new message class
packageStartupMessage that can be sup-
pressed separately.
• A new function, method.skeleton() writes a
skeleton version of a call to setMethod() to a
file, with correct arguments and format, given
the name of the function and the method sig-
nature.
•mode<- and storage.mode<- do slightly less
copying.
•nls.control(* , printEval = FALSE,
warnOnly = FALSE) are two new options
to help better analyze (non-)convergence of
nls(), thanks to Kate Mullen.
nls() and summary(nls()) now contain more
information and also print information about
convergence.
•options(device = ) now accepts a function
object as well as the name of a function.
•pdf() supports new values for paper of ‘"US"’
(same as ‘"letter"’), ‘"a4r"’ and ‘"USr"’
(the latter two meaning rotated to landscape).
postscript() also accepts paper = "US".
•persp() now respects the graphical pars
cex.axis,cex.lab,font.axis and font.lab.
• New faster internal functions pmax.int() and
pmin.int() for inputs which are atomic vec-
tors without classes (called by pmax/pmin
where applicable).
pmin/pmax are now more likely to work with
classed objects: they work with POSIXlt date-
times, for example.
•postscript() now by default writes grey col-
ors (including black and white) via setgray,
which gives more widely acceptable output.
There are options to write pure RGB, CMYK or
gray via the new argument colormodel.
•rbind.data.frame() now ignores all zero-row
inputs, as well as zero-column inputs (which
it used to do, undocumented). This is because
read.table() can create zero-row data frames
with NULL columns, and those cannot be ex-
tended.
•readChar() and writeChar() can now work
with a raw vector.
•read.table(),write.table() and allies have
been moved to package utils.
•rgb() now accepts the red, green and blue
components in a single matrix or data frame.
• New utility function RShowDoc() in package
utils to find and display manuals and other
documentation files.
• New .row_names_info() utility function finds
the number of rows efficiently for data frames;
consequently, dim.data.frame() has become
very fast for large data frames with ‘automatic’
row names.
•RSiteSearch() now also allows to search post-
ings of the ‘R-devel’ mailing list.
R News ISSN 1609-3631

Vol. 7/1, April 2007 46
•screeplot() is now (S3) generic with a default
method, thanks to a patch from Gavin Simp-
son.
• Experimental verbose argument for
selectMethod(). Might be replaced later by
a better interface for method selection inspec-
tion.
• Added links to source files to the parsing rou-
tines, so that source() can now echo the orig-
inal source and comments (rather than depars-
ing). This affects example() and Sweave() as
well.
•stack() and unstack() have been moved to
package utils.
•strptime() now sets the ‘tzone’ attribute on
the result if tz != "".
•str.default() typically prints fewer entries of
logical vectors.
• The RweaveLatex driver for Sweave() now
supports two new options: expand=FALSE, to
show chunk references in the output, and
concordance=TRUE, to output the concordance
between input and output lines.
•system() now takes the same set of arguments
on all platforms, with those which are not ap-
plicable being ignored with a warning. Unix-
alikes gain input and wait, and Windows
gains ignore.stderr.
•system.time() and proc.time() now return
an object of class proc_time with a print()
method that returns a POSIX-like format with
names.
•Sys.getenv() has a new argument unset to al-
low unset and set to "" to be distinguished (if
the OS does). The results of Sys.getenv() are
now sorted (by name).
• New function Sys.glob(), a wrapper for the
POSIX.2 function glob(3) to do wildcard ex-
pansion (on systems which have it, plus an em-
ulation on Windows).
•Sys.setenv() is a new (and preferred) syn-
onym for Sys.putenv(). The internal C code
uses the POSIX-preferred setenv rather than
putenv where the former is available.
• New function Sys.unsetenv() to remove envi-
ronment variables (on systems where unsetenv
is implemented or putenv can remove vari-
ables, such as on Windows).
•text(),mtext(),strheight(),strwidth(),
legend(),axis(),title(),pie(),
grid.text() and textGrob() all attempt to
coerce non-language annotation objects (in the
sense of is.object) to character vectors. This
is principally intended to cover factors and
POSIXt and Date objects, and is done via the
new utility function as.graphicsAnnot() in
package grDevices.
•tk_select.list() in package tcltk now
chooses the width to fit the widest item.
•retracemem() and untracemem() are now
primitives for efficiency and so migrate from
utils to base.
•union(),intersect(),setdiff() and
setequal() now coerce their arguments to be
vectors (and they were documented only to
apply to vectors).
•uniroot() now works if the zero occurs at one
of the ends of the interval (suggestion of Tamas
Papp).
• There is a new function View() for viewing
matrix-like objects in a spreadsheet, which can
be left up whilst R is running.
• New function withVisible() allows R level
access to the visibility flag.
•zip.file.extract() has been moved to pack-
age utils.
• A few more cases of subassignment work,
e.g., raw[] <- list and vector[] <- expres-
sion, with suitable coercion of the LHS.
• There is a warning if a ‘\’ is used unnecessarily
in a string when being parsed., e.g., ‘\.’ where
probably ‘\\.’ was intended. (‘\.’ is valid, but
the same as ‘.’.) Thanks to Bill Dunlap for the
suggestion.
• Introduced the suffix Lfor integer literals to
create integer rather than numeric values, e.g.,
100L,0x10L,1e2L.
• Set the parser to give verbose error messages in
case of syntax errors.
• The class LinearMethodsList has been ex-
tended and will be used to create list ver-
sions of methods, derived from the methods
tables (environments). The older recursive
MethodsList class will be deprecated (by the
release of 2.5.0 if possible).
• There are more flexible ways to specify
the default library search path. In ad-
dition to R_LIBS and .Library, there are
.Library.site (defaults to ‘R HOME/site-
library’) and R_LIBS_USER (defaults to a
platform- and version-specific directory in
‘∼/R’). See ?.libPaths for details.
R News ISSN 1609-3631

Vol. 7/1, April 2007 47
• LAPACK has been updated to version 3.1.0.
This should cause only small changes to the
output, but do remember that the sign of eigen-
vectors (and principal components) is indeter-
minate.
• PCRE has been updated to version 7.0.
• Several functions handle row names more effi-
ciently:
–read.table() and read.DIF() make use
of integer row names where appropriate,
and avoid at least one copy in assigning
them.
–data.frame() and the standard
as.data.frame() methods avoid gener-
ating long dummy row names and then
discarding them.
–expand.grid() and merge() generate
compact ‘automatic’ row names.
–data.matrix() and
as.matrix.data.frame() have a new ar-
gument ‘rownames.force’ that by default
drops ‘automatic’ row names.
•data_frame[i, j] is substantially more
memory-efficient when only a small part of the
data frame is selected, especially when (part
of) a single column is selected.
• Command-line R (and ‘Rterm.exe’ under Win-
dows) accepts the options ‘-f filename’,
‘--file=filename’ and ‘-e expression’ to fol-
low other script interpreters. These imply
‘--no-save’ unless ‘--save’ is specified.
• Invalid bytes in character strings in an
MBCS now deparse/print in the form ‘\xc1’
rather than ‘<c1>’, which means they can be
parsed/scanned.
• Printing functions (without source attributes)
and expressions now preserves integers (using
the Lsuffix) and NAs (using NA_real_ etc where
necessary).
• The ‘internal’ objects .helpForCall,.tryHelp
and topicName are no longer exported from
utils.
• The internal regex code has been upgraded to
glibc 2.5 (from 2.3.6).
• Text help now attempts to display files which
have an encoding section in the specified en-
coding via file.show().
• R now attempts to keep track of character
strings which are known to be in Latin-1 or
UTF-8 and print or plot them appropriately in
other locales. This is primarily intended to
make it possible to use data in Western Eu-
ropean languages in both Latin-1 and UTF-
8 locales. Currently scan(),read.table(),
readLines(),parse() and source() allow en-
codings to be declared, and console input in
suitable locales is also recognized.
New function Encoding() can read or set the
declared encodings for a character vector.
• There have been numerous performance im-
provements to the data editor on both Win-
dows and X11. In particular, resizing the win-
dow works much better on X11.
Deprecated & defunct
•symbol.C() and symbol.For() are defunct,
and have been replaced by wrappers that give
a warning.
• Calling a builtin function with an empty argu-
ment is now always an error.
• The autoloading of ts() is defunct.
• The undocumented reserved word GLOBAL.ENV
has been removed. (It was yet another way to
get the value of the symbol .GlobalEnv.)
• The deprecated behaviour of structure() in
adding a class when specifying with tsp or
levels attributes is now defunct.
•unix() is now finally defunct, having been
deprecated for at least seven years.
•Sys.putenv() is now deprecated in favour
of Sys.setenv(), following the POSIX recom-
mendation.
• Building R with ‘--without-iconv’ is depre-
cated.
• Using $on an atomic vector is deprecated (it
was previously valid and documented to re-
turn NULL).
• The use of storage.mode<- for other than
standard types (and in particular for value
"single") is deprecated: use mode<- instead.
Installation
• A suitable iconv (e.g., from glibc or GNU
libiconv) is required. For 2.5.x only you
can build R without it by configuring using
‘--without-iconv’.
• There is support again for building on AIX
(tested on 5.2 and 5.3) thanks to Ei-ji Nakama.
R News ISSN 1609-3631

Vol. 7/1, April 2007 48
• Autoconf 2.60 or later is used to create
configure. This makes a number of small
changes, and incorporates the changes to the
detection of a C99-compliant C compiler back-
ported for 2.4.1.
• Detection of a Java development environment
was added such that packages don’t need to
provide their own Java detection.
R CMD javareconf was updated to look for the
corresponding Java tools as well.
• Added workaround for reported non-POSIX
sh on OSF1. (PR#9375)
•make install-strip now works, stripping the
executables and also the shared libraries and
modules on platforms where libtool knows
how to do so.
• Building R as a shared library and standalone
nmath now installs pkg-config files ‘libR.pc’
and ‘libRmath.pc’ respectively.
• Added test for insufficiently complete imple-
mentation of sigaction.
C-level facilities
• Functions str2type,type2char and type2str
are now available in ‘Rinternals.h’.
• Added support for Objective C in R and pack-
ages (if available).
•R_ParseVector() has a new 4th argument
SEXP srcfile allowing source references to be
attached to the returned expression list.
• Added ptr_R_WriteConsoleEx callback which
allows consoles to distinguish between reg-
ular output and errors/warnings. To en-
sure backward compatibility it is only used if
ptr_R_WriteConsole is set to NULL.
Utilities
• Additional Sweave() internal functions are
exported to help writing new drivers, and
RweaveLatexRuncode() is now created using a
helper function (all from a patch submitted by
Seth Falcon).
• The following additional flags are accessible
from R CMD config:OBJC,OBJCFLAGS,JAR,
JAVA,JAVAC,JAVAH,JAVA_HOME,JAVA_LIBS
•R CMD build now takes the package name
from the ‘DESCRIPTION’ file and not from the
directory. (PR#9266)
•checkS3methods() (and hence R CMD check)
now checks agreement with primitive internal
generics, and checks for additional arguments
in methods where the generic does not have a
... argument.
codoc() now knows the argument lists of
primitive functions.
•R CMD INSTALL and R CMD REMOVE now use as
the default library (if ‘-l’ is not specified) the
first library that would be used if R were run
in the current environment (and they run R to
find it).
• There is a new front-end ‘Rscript’ which can
be used for #! scripts and similar tasks. See
help("Rscript") and ‘An Introduction to R’
for further details.
•R CMD BATCH (not Windows) no longer
prepends ’invisible(options(echo =
TRUE))’ to the input script. This was the de-
fault unless ‘--slave’ is specified and the latter
is no longer overridden.
On all OSes it makes use of the ‘-f’ argument
to R, so file("stdin") can be used from BATCH
scripts.
On all OSes it reports proc.time() at the end
of the script unless q() is called with options to
inhibit this.
•R CMD INSTALL now prepends the installation
directory (if specified) to the library search
path.
• Package installation now re-encodes R files and
the ‘NAMESPACE’ file if the ‘DESCRIPTION’
file specifies an encoding, and sets the en-
coding used for reading files in preparing for
LazyData. This will help if a package needs to
be used in (say) both Latin-1 and UTF-8 locales
on different systems.
•R CMD check now reports on non-ASCII strings
in datasets. (These are a portability issue,
which can be alleviated by marking their en-
coding: see ‘Writing R Extensions’.)
•Rdiff now converts CRLF endings in the target
file, and converts UTF-8 single quotes in either
to ASCII quotes.
• New recommended package codetools by
Luke Tierney provides code-analysis tools.
This can optionally be used by R CMD check to
detect problems, especially symbols which are
not visible.
•R CMD config now knows about LIBnn .
R News ISSN 1609-3631

Vol. 7/1, April 2007 49
• New recommended package rcompgen
by Deepayan Sarkar provides support for
command-line completion under the Unix ter-
minal interface (provided readline is enabled)
and the Windows Rgui and Rterm front ends.
Bug fixes
•gc() can now report quantities of ‘Vcells’ in ex-
cess of 16Gb on 64-bit systems (rather than re-
porting NA).
• Assigning class factor to an object now re-
quires it has integer (and not say double) codes.
•structure() ensures that objects with added
class factor have integer codes.
• The formula and outer attributes of datasets
ChickWeight,CO2,DNase,Indometh,Loblolly,
Orange and Theoph now have an empty en-
vironment and not the environment used to
dump the datasets in the package.
• Dataset Seatbelts now correctly has class
c("mts", "ts").
•str() now labels classes on data frames more
coherently.
• Several ‘special’ primitives and .Internals could
return invisibly if the evaluation of an argu-
ment led to the visibility flag being turned off.
These included as.character(),as.vector(),
call(),dim(),dimnames(),lapply(),rep(),
seq() and seq_along(). Others (e.g., dput()
and print.default()) could return visibly
when this was not intended.
• Several primitives such as dim() were not
checking the number of arguments supplied
before method dispatch.
• Tracing of primitive functions has been cor-
rected. It should now be the case that tracing
either works or is not allowed for all primi-
tive functions. (Problems remain if you make a
primitive into a generic when it is being traced.
To be fixed later.)
•max.col() now omits infinite values in deter-
mining the relative tolerance.
•R CMD Sweave and R CMD Stangle now re-
spond to ‘--help’ and ‘--version’ like other
utilities.
•.libPaths() adds only existing directories (as
it was documented to, but could add non-
directories).
•setIs() and setClassUnion() failed to find
some existing subclasses and produced spuri-
ous warnings, now fixed.
•data.frame() ignored row.names for 0-column
data frames, and no longer treats an explicit
row.names=NULL differently from the default
value.
•identical() looked at the internal structure of
the row.names attribute, and not the value visi-
ble at R level.
•abline(reg) now also correctly works with
intercept-only lm models, and abline() warns
more when it’s called illogically.
•warning() was truncating messages at
getOption("warning.length") - 1 (not as doc-
umented), with no indication. It now appends
[... truncated].
• Stangle/Sweave were throwing spurious
warnings if options result or strip.white
were unset.
•all.equal() was ignoring check.attributes
for list and expression targets, and checking
only attributes on raw vectors. Logical vectors
were being compared as if they were numeric,
(with a mean difference being quoted).
• Calculating the number of significant digits in
a number was itself subject to rounding er-
rors for digits ≥16. The calculation has
been changed to err on the side of slightly
too few significant digits (but still at least 15)
rather than far too many. (An example is
print(1.001, digits=16).)
•unlink() on Unix-alikes failed for paths con-
taining spaces.
•substr() and friends treated NA start or stop
incorrectly.
•merge(x, y, all.y = TRUE) would some-
times incorrectly return logical columns for
columns only in ywhen there were no com-
mon rows.
•read.table(fn, col.names=) on an empty
file returned NULL columns, rather than
logical(0) columns (which is what results
from reading a file with just a header).
•grid.[xy]axis(label=logical(0)) failed.
•expression() was unnecessarily duplicating
arguments.
•as.expression(list)returned a single-
element expression vector, which was not com-
patible with S: it now copies lists element-by-
element.
R News ISSN 1609-3631

Vol. 7/1, April 2007 50
•supsmu(periodic = TRUE) could segfault.
(PR#9502, detection and patch by Bill Dunlap.)
•pmax/pmin called with only logical arguments
did not coerce to numeric, although they were
documented to do so (as max/min do).
•methods() did not know that cbind() and
rbind() are internally generic.
•dim(x) <- NULL removed the names of x, but
this was always undocumented. It is not clear
that it is desirable but it is S-compatible and re-
lied on, so is now documented.
•which(x, arr.ind = TRUE) did not return a
matrix (as documented) if xwas an array of
length 0.
• C-level duplicate() truncated CHARSXPs with
embedded nuls.
• Partial matching of attributes was not work-
ing as documented in some cases if there were
more than two partial matches or if ‘names’
was involved.
•data(package=character(0)) was not looking
in ‘./data’ as documented.
•summary.mlm() failed if some response names
were "" (as can easily happen if cbind() is
used).
• The postscript() and pdf() drivers shared
an encoding list but used slightly different for-
mats. This caused problems if both were used
with the same non-default encoding in the
same session. (PR#9517)
• The data editor was not allowing Inf,NA and
NaN to be entered in numerical columns. It was
intended to differentiate between empty cells
and NAs, but did not do so: it now does so for
strings.
•supsmu() could segfault if all cases had non-
finite values. (PR#9519)
•plnorm(x, lower.tail=FALSE) was returning
the wrong tail for x≤0. (PR#9520)
•which.min() would not report a minimum
of +Inf, and analogously for which.max().
(PR#9522)
•R CMD check could fail with an unhelpful er-
ror when checking Rd files for errors if there
was only one file and that had a serious error.
(PR#9459)
•try() has been reimplemented using
tryCatch() to solve two problems with the
original implementation: (i) try() would run
non-NULL options("error") expressions for
errors within a try, and (ii) try() would catch
user interrupts.
•str(obj) could fail when obj contained a den-
drogram.
• Using data_frame[, last_column] <- NULL
failed (PR#9565).
•choose(n, k) could return non-integer values
for integer nand small kon some platforms.
•nclass.scott(x) and nclass.FD(x) no longer
return NaN when var(x) or IQR(x) (respec-
tively) is zero.
hist() now allows breaks = 1 (which the
above patch will return), but not breaks = Inf
(which gave an obscure error).
•strptime("%j") now also works for the first
days of Feb-Dec. (PR#9577)
•write.table() now recovers better if file is
an unopened connection. (It used to open it for
both the column names and the data.)
• Fixed bug in mosaicplot(sort=) introduced
by undocumented change in R 2.4.1 (changeset
r39655).
•contr.treatment(n=0) failed with a spurious
error message. (It remains an error.)
•as.numeric() was incorrectly documented: it
is identical to as.double().
•jitter(rep(-1, 3)) gave NaNs. (PR#9580)
•max.col() was not random for a row of zeroes.
(PR#9542)
•ansari.test(conf.int=TRUE, exact=FALSE)
failed.
•trace() now works on S3 registered methods,
by modifying the version in the S3 methods ta-
ble.
•rep(length=1, each=0) segfaulted.
•postscript() could overflow a buffer if used
with a long command argument.
• The internal computations to copy complete at-
tribute lists did not copy the flag marking S4
objects, so the copies no longer behaved like S4
objects.
• The C code of nlminb() was altering a vari-
able without duplicating it. (This did not af-
fect nlminb() but would have if the code was
called from a different wrapper.)
•smooth(kind = "3RS3R") (the current default)
used .C(DUP = FALSE) but altered its input ar-
gument. (This was masked by duplication in
as.double().)
R News ISSN 1609-3631

Vol. 7/1, April 2007 51
• The signature for the predefined S4 method for
as.character() was missing ... .
•readBin(raw_vector)could read beyond the
end of the vector when size-changing was in-
volved.
• The C entry point PrintValue (designed to em-
ulate auto-printing) would not find show() for
use on S4 objects, and did not have the same
search path (for show(),print() and print()
methods) as auto-printing. Also, auto-printing
and print() of S4 objects would fail to find
show if the methods namespace was loaded but
the package was not attached (or otherwise not
in the search path).
•print() (and auto-printing) now recognize S4
objects even when methods is not loaded, and
print a short summary rather than dump the
internal structure.
Changes on CRAN
by Kurt Hornik
New contributed packages
ARES Allelic richness estimation, with extrapola-
tion beyond the sample size. Generates an al-
lelic richness accumulation curve. This curve
shows the expected number of unique alleles
in a population when taking a sample of indi-
viduals. The function aresCalc takes a binary
data matrix as input, showing the presence of
alleles per individual, and gives an accumula-
tion curve (mean with 95% confidence bounds)
back. The function aresPlot can be used to
plot the output from aresCalc. By Emiel van
Loon and Scott Davis.
BayHaz Bayesian Hazard Rate Estimation: a suite of
R functions for Bayesian estimation of smooth
hazard rates via Compound Poisson Process
(CPP) priors. By Luca La Rocca.
BiasedUrn Biased Urn model distributions. Statis-
tical models of biased sampling in the form
of univariate and multivariate non-central hy-
pergeometric distributions, including those of
Wallenius and Fisher (also called extended hy-
pergeometric distribution). By Agner Fog.
BootCL Bootstrapping test for chromosomal local-
ization. By Eun-Kyung Lee, Samsun Sung, and
Heebal Kim.
Brobdingnag Very large numbers in R. Real num-
bers are held using their natural logarithms,
plus a logical flag indicating sign. The package
includes a vignette that gives a step-by-step in-
troduction to using S4 methods. By Robin K. S.
Hankin.
CCA Canonical correlation analysis. Provides a set
of functions that extend the cancor function
with new numerical and graphical outputs.
It also include a regularized extension of the
canonical correlation analysis to deal with data
sets with more variables than observations. By
Ignacio González and Sébastien Déjean.
CreditMetrics Functions for calculating the Credit-
Metrics risk model. By Andreas Wittmann.
DAAGxtras Data sets and functions additional to
DAAG, used in additional exercises for the
book “Data Analysis and Graphics Using R”
by J. H. Maindonald and W. J. Brain (2007, 2nd
edn.), and for laboratory exercises prepared for
a ‘Data Mining’ course. By John Maindonald.
GenABEL genome-wide association analysis be-
tween quantitative or binary traits and SNPs.
By Yurii Aulchenko.
GeoXp Interactive exploratory spatial data analy-
sis. A tool for researchers in spatial statis-
tics, spatial econometrics, geography, ecol-
ogy etc., allowing to link dynamically statis-
tical plots with elementary maps. By Chris-
tine Thomas-Agnan, Yves Aragon, Anne Ruiz-
Gazen, Thibault Laurent, and Laurianne Ro-
bidou.
HydroMe Estimation of the parameters in infiltra-
tion and water retention models by curve-
fitting method. The models considered are
those that are commonly used in soil science.
By Christian Thine Omuto.
IPSUR Data sets and functions accompanying the
book “Introduction to Probability and Statistics
Using R” by G. Andy Chang and G. Jay Kerns
(in progress). By G. Jay Kerns with contribu-
tions by Theophilius Boye, adapted from the
work of John Fox et al.
InfNet Simulation of epidemics in a network of con-
tacts. The simulations consider SIR epidemics
with events in continuous time (exponential
inter-event times). It can consider a structure
of local networks and has an option to visual-
ize it with the animator called SoNIA (http:
//www.stanford.edu/group/sonia/). By Lilia
Ramirez Ramirez and Mary Thompson.
R News ISSN 1609-3631

Vol. 7/1, April 2007 52
JointModeling Joint modeling of mean and disper-
sion through two interlinked GLMs or GAMs.
By Mathieu Ribatet and Bertrand Iooss.
NMMAPSlite NMMAPS Data Lite. Provides re-
mote access to daily mortality, weather, and
air pollution data from the National Morbid-
ity, Mortality, and Air Pollution Study for 108
U.S. cities (1987–2000); data are obtained from
the Internet-based Health and Air Pollution
Surveillance System (iHAPSS). By Roger D.
Peng.
NRAIA Data sets from the book “Nonlinear Regres-
sion Analysis and Its Applications” by D. M.
Bates and D. G. Watts (1998), John Wiley and
Sons. By Douglas Bates.
PKtools Unified computational interfaces for pop
PK. By M. Suzette Blanchard.
PhySim Phylogenetic tree simulation based on a
virth death model. Functions are provided to
model a lag-time to speciation and extract sister
species ages. By Jason T. Weir, Dolph Schluter.
PresenceAbsence Presence-absence model evalua-
tion. Includes functions for calculating thresh-
old dependent measures such as confusion ma-
trices, pcc, sensitivity, specificity, and Kappa,
and produces plots of each measure as the
threshold is varied. Can also optimize thresh-
old choice, and plot the threshold independent
ROC curves along with the associated AUC By
Elizabeth Freeman.
RJDBC Provides access to databases through the
JDBC interface, implementing R’s DBI interface
using JDBC as a back-end. This allows R to con-
nect to any DBMS that has a JDBC driver. By
Simon Urbanek.
RLadyBug Analysis of small scale infectious disease
data using stochastic Susceptible-Exposed-
Infectious-Recovered (SEIR) models. By
Michael Hoehle and Ulrike Feldmann.
RaschSampler A Rasch sampler for sampling bi-
nary matrices with fixed margins. By Reinhold
Hatzinger, Patrick Mair, and Norman Verhelst.
Rcmdr.HH Rcmdr support for the introductory
course at Temple University, which spends
time on several topics that are not yet in the
R Commander. By Richard M. Heiberger, with
contributions from Burt Holland.
Rserve A binary R server which acts as a socket
server (TCP/IP or local sockets) allowing bi-
nary requests to be sent to R. Every connection
has a separate workspace and working direc-
tory. Client-side implementations are available
for popular languages such as C/C++ and Java,
allowing any application to use facilities of R
without the need of linking to R code. Supports
remote connection, user authentication and file
transfer. A simple R client is included as well.
By Simon Urbanek.
SLmisc Miscellaneous functions for analysis of
gene expression data at SIRS-Lab GmbH. By
Matthias Kohl.
SMPracticals Data sets and a few functions for use
with the practicals outlined in Appendix A
of the book “Statistical Models” by A. Davi-
son (2003), Cambridge University Press. The
practicals themselves can be found at http://
statwww.epfl.ch/davison/SM/. By Anthony
Davison.
SNPmaxsel Asymptotic methods related to maxi-
mally selected statistics, with applications to
SNP data. By Anne-Laure Boulesteix.
Snowball Snowball stemmers. By Kurt Hornik.
SpherWave Spherical wavelets and SW-based spa-
tially adaptive methods. Carries out spherical
wavelet transform developed Li (1999) and Oh
(1999), and implements wavelet thresholding
approaches proposed by Oh and Li (2004). By
Hee-Seok Oh and Donghoh Kim.
Synth Causal inference using the synthetic con-
trol group method of Abadie and Gardeaza-
bal (2003) and Abadie, Diamond, Hainmueller
(2007). By Alexis Diamond and Jens Hain-
mueller.
TRAMPR Matching of terminal restriction frag-
ment length polymorphism (TRFLP) profiles
between unknown samples and a database of
knowns. Facilitates the analysis of many un-
known profiles at once, and provides tools for
working directly with electrophoresis output
through to generating summaries suitable for
community analyses with R’s rich set of statisti-
cal functions. Also resolves the issues of multi-
ple TRFLP profiles within a species, and shared
TRFLP profiles across species. By Rich FitzJohn
and Ian Dickie.
VGAM Vector generalized linear and additive mod-
els, and associated models (Reduced-Rank
VGLMs, Quadratic RR-VGLMs, Reduced-Rank
VGAMs). Fits many models and distribution
by maximum likelihood estimation (MLE) or
penalized MLE. Also fits constrained ordina-
tion models in ecology. By Thomas W. Yee.
WWGbook Functions and data sets for the book
“Linear Mixed Models: A Practical Guide Us-
ing Statistical Software” by B. T. West, K. B.
Welch, and A. T. Galecki (2006), Chapman
R News ISSN 1609-3631

Vol. 7/1, April 2007 53
Hall / CRC Press. By Shu Chen and Andrzej
Galecki.
WeedMap Map of weed intensity. Compute spatial
predictions from exact count and a covariate.
By Gilles Guillot.
ZIGP Adapt and analyze Zero Inflated Generalized
Poisson (ZIGP) regression models. By Vinzenz
Erhardt.
adimpro Adaptive smoothing of digital images
via the Propagation Separation approach by
Polzehl and Spokoiny (2006). By Karsten Tabe-
low and Joerg Polzehl.
agricolae Statistical Procedures for Agricultural Re-
search. These functions are currently utilized
by the International Potato Center Research
(CIP), the Statistics and Informatics Instruc-
tors and the Students of the Universidad Na-
cional Agraria La Molina Peru, and the Special-
ized Master in “Bosques y Gestión de Recur-
sos Forestales” (Forest Resource Management).
Contains functionality for the statistical anal-
ysis of experimental designs applied specially
for field experiments in agriculture and plant
breeding. By Felipe de Mendiburu.
analogue Analogue methods for palaeoecology. Fits
modern analogue technique transfer function
models for prediction of environmental data
from species data. Also performs analogue
matching, a related technique used in palaeoe-
cological restoration. By Gavin L. Simpson.
arm Data analysis using regression and multi-
level/hierarchical models. By Andrew Gel-
man, Yu-Sung Su, Jennifer Hill, Maria Grazia
Pittau, Jouni Kerman, and Tian Zheng.
arrayMissPattern Exploratory analysis of missing
patterns for microarray data. By Eun-kyung
Lee and Taesung Park.
bbmle Modifications and extensions of the stats4
mle code. By Ben Bolker, modifying code by
Peter Dalgaard and other R-core members.
bcp Bayesian Change Point. An implementation of
the Barry and Hartigan (1993) product parti-
tion model for the standard change point prob-
lem using Markov Chain Monte Carlo. By
Chandra Erdman.
binGroup Evaluation and experimental design
for binomial group testing. By Frank
Schaarschmidt.
cgh Analysis of microarray comparative genome
hybridization data using the Smith-Waterman
algorithm. By Tom Price.
classifly Exploration of classification models in high
dimensions, implementing methods for under-
standing the division of space between groups.
See http://had.co.nz/classifly for more
details. By Hadley Wickham.
clustTool GUI for clustering data with spatial infor-
mation. By Matthias Templ.
clusterGeneration Random cluster generation (with
specified degree of separation). By Weiliang
Qiu and Harry Joe.
clusterSim Searching for optimal clustering proce-
dure for a data set. By Marek Walesiak Andrzej
Dudek.
cobs99 Constrained B-splines – outdated 1999 ver-
sion. By Pin T. Ng and Xuming He; R port by
Martin Maechler.
corrgram Plot correlograms. By Kevin Wright.
distrDoc Documentation for packages distr,dis-
trEx,distrSim, and distrTEst. By Florian Cam-
phausen, Matthias Kohl, Peter Ruckdeschel,
and Thomas Stabla.
drm Likelihood-based marginal regression and as-
sociation modeling for repeated, or otherwise
clustered, categorical responses using depen-
dence ratio as a measure of the association. By
Jukka Jokinen.
elrm Exact Logistic Regression via MCMC. By
David Zamar, Jinko Graham, and Brad Mc-
Neney.
emdbook Data sets and auxiliary functions for the
book “Ecological Models and Data” by B.
Bolker (in progress). By Ben Bolker.
epicalc Epidemiological calculator. By Virasakdi
Chongsuvivatwong.
fEcofin Selected economic and financial data for
teaching financial engineering and computa-
tional finance. By Diethelm Wuertz and many
others.
fame Interface for the FAME time series database.
Includes FAME storage and retrieval function,
as well as functions and S3 classes for time in-
dexes and time indexed series, which are com-
patible with FAME frequencies. By Jeff Hall-
man.
filehashSQLite Simple key-value database using
SQLite as the backend. By Roger D. Peng.
gamlss.dist Extra distributions for GAMLSS mod-
eling. By Mikis Stasinopoulos and Bob Rigby
with contributions from Calliope Akantziliotou
and Raydonal Ospina.
R News ISSN 1609-3631

Vol. 7/1, April 2007 54
glmc Fits generalized linear models where the pa-
rameters are subject to linear constraints. The
model is specified by giving a symbolic de-
scription of the linear predictor, a description
of the error distribution, and a matrix of con-
straints on the parameters. By Sanjay Chaud-
huri, Mark S. Handcock, and Michael S. Ren-
dall.
heplots Visualizing Tests in Multivariate Linear
Models. Represents sums-of-squares-and-
products matrices for linear hypotheses and for
error using ellipses (in two dimensions) and el-
lipsoids (in three dimensions). By John Fox,
Michael Friendly, and Georges Monette.
homtest A collection of homogeneity tests and other
stuff for hydrological applications. By Alberto
Viglione.
ibdreg A method to test genetic linkage with covari-
ates by regression methods with response IBD
sharing for relative pairs. Accounts for correla-
tions of IBD statistics and covariates for relative
pairs within the same pedigree. By Jason P. Sin-
nwell and Daniel J. Schaid.
kernelPop Spatially explicit population genetic sim-
ulations. Creates an individual-based popula-
tion genetic simulation in discrete time, where
individuals have spatial coordinates, with dis-
persal governed by mixtures of Weibull and
normal pdfs. Simulates diploid and haploid
inheritance. Allows the development of null
distributions of genotypes for complex demo-
graphic scenarios. By Allan Strand and James
Niehaus.
klin Implements efficient ways to evaluate and
solve equations of the form Ax =b, where Ais
a Kronecker product of matrices. Also includes
functions to solve least squares problems of this
type. By Tamas K Papp.
knnflex A more flexible KNN implementation
which allows the specification of the distance
used to calculate nearest neighbors (euclidean,
binary, etc.), the aggregation method used
to summarize response (majority class, mean,
etc.) and the method of handling ties (all,
random selection, etc.). Additionally, continu-
ous responses are handled. By Atina Dunlap
Brooks.
languageR Data sets and functions for the book
“Analyzing Linguistic Data: A practical intro-
duction to Statistics” by R. H. Baayen (2006),
Cambridge University Press. By R. H. Baayen.
lss Accelerated failure time model to right censored
data based on least-squares principle. By Lin
Huang and Zhezhen Jin.
mFilter Several time series filters useful for smooth-
ing and extracting trend and cyclical compo-
nents of a time series. Currently, Christiano-
Fitzgerald, Baxter-King, Hodrick-Prescott, But-
terworth, and trigonometric regression filters
are included in the package. By Mehmet Bal-
cilar.
meboot Maximum entropy density based depen-
dent data bootstrap for time series. An algo-
rithm is provided to create a population of time
series (ensemble) without assuming stationar-
ity. By Hrishikesh D. Vinod and Javier López-
de-Lacalle.
memisc Miscellaneous tools for data management,
including tools for preparing (especially so-
cial science) survey data, conducting simula-
tion studies, and presentation of results of sta-
tistical analyses. By Martin Elff.
mixstock Functions for mixed stock analysis. By
Ben Bolker.
mixtools Tools for analyzing mixture models. By
Ryan Elmore, Tom Hettmansperger, David
Hunter, Hoben Thomas, Fengjuan Xuan, and
Derek Young.
mlCopulaSelection Use numerical maximum likeli-
hood to choose and fit a bivariate copula model
(from a library of 40 models) to the data. By Je-
sus Garcia and Veronica Gonzalez-Lopez.
msgcop Semiparametric Bayesian Gaussian copula
estimation, treating the univariate marginal
distributions as nuisance parameters in Hoff
(2006). Also provides a semiparametric impu-
tation procedure for missing multivariate data.
By Peter Hoff.
muS2RC S-PLUS to R compatibility for package
muStat. By Knut M. Wittkowski and Tingting
Song.
muStat Prentice Rank Sum Test and McNemar Test.
Performs Wilcox rank sum test, Kruskal rank
sum test, Friedman rank sum test and McNe-
mar test. By Knut M. Wittkowski and Tingting
Song.
muUtil A collection of utility functions for package
muStat. By Knut M. Wittkowski and Tingting
Song.
np Nonparametric kernel smoothing methods for
mixed data types. By Tristen Hayfield and Jef-
frey S. Racine.
nsRFA Tools for objective (non-supervised) applica-
tions of the Regional Frequency Analysis meth-
ods in hydrology. By Alberto Viglione.
R News ISSN 1609-3631

Vol. 7/1, April 2007 55
pARtial (Partial) attributable risk estimates, corre-
sponding variance estimates and confidence
intervals. By Andrea Lehnert-Batar.
pmml Generate PMML for various models. The
Predictive Modeling Markup Language
(PMML) is a language for representing mod-
els, in an application independent way. Such
models can then be loaded into other appli-
cations supporting PMML, such as Teradata
Warehouse Miner and IBM’s DB2. By Graham
Williams.
popbio Construct and analyze projection matrix
models from a demography study of marked
individuals classified by age or stage. Covers
methods described in the books “Matrix Popu-
lation Models” by Caswell (2001) and “Quan-
titative Conservation Biology” by Morris and
Doak (2002). By Chris Stubben and Brook Mil-
ligan.
portfolioSim Framework for simulating equity
portfolio strategies. By Jeff Enos and David
Kane, with contributions from Kyle Campbell.
prim Patient Rule Induction Method (PRIM) for
estimating highest density regions in high-
dimensional data. By Tarn Duong.
qgen Analysis of quantitative genetic data, espe-
cially helpful to perform parametric resam-
pling of quantitative genetic data sets. By
Thomas Fabbro.
rcdk Interface to the CDK libraries, a Java frame-
work for cheminformatics. Allows to load
molecules, evaluate fingerprints, calculate
molecular descriptors and so on. By Rajarshi
Guha.
rcompgen Generates potential completions for a
given command snippet based on circumstan-
tial evidence. By Deepayan Sarkar.
rcompletion Pseudo-intelligent TAB completion for
R when run from a terminal, using the GNU
Readline library. By Deepayan Sarkar.
relaxo Relaxed Lasso, a generalization of the Lasso
shrinkage technique for linear regression. Both
variable selection and parameter estimation is
achieved by regular Lasso, yet both steps do
not necessarily use the same penalty parame-
ter. By Nicolai Meinshausen.
rggm Fitting Gaussian Graphical Models with ro-
bustified methods. By Masashi Miyamura.
rjacobi Compute Jacobi polynomials and Gauss-
Jacobi quadrature related operations. By Paulo
Jabardo.
sciplot Scientific graphs with error bars for data col-
lected from one-way or higher factorial de-
signs. By Manuel Morales.
signal A set of generally Matlab/Octave-compatible
signal processing functions. Includes filter gen-
eration utilities, filtering functions, resampling
routines, and visualization of filter models. By
Tom Short.
smoothtail Smooth Estimation of the Generalized
Pareto distribution shape parameter. By Kas-
par Rufibach and Samuel Mueller.
snp.plotter Creates plots of p-values using single
SNP and/or haplotype data, with options to
display a linkage disequilibrium (LD) plot and
the ability to plot multiple data sets simulta-
neously. By Augustin Luna and Kristin K.
Nicodemus.
spsurvey Algorithms required for design and anal-
ysis of probability surveys such as those uti-
lized by the U.S. Environmental Protection
Agency’s Environmental Monitoring and As-
sessment Program (EMAP). By Tom Kincaid
and Tony Olsen, with contributions from Don
Stevens, Christian Platt, Denis White, and
Richard Remington.
staRt Inferenza classica con TI-83 Plus. Una libre-
ria per utilizzare con semplicità le tecniche di
statistica inferenziale presenti sulla calcolatrice
scientifica grafica TI-83 Plus. By Fabio Frascati.
stashR A set of tools for administering shared repos-
itories. By Sandy Eckel, and Roger D. Peng.
stochmod Stochastic Modeling: learning and infer-
ence algorithms for a variety of probabilistic
models. By Artem Sokolov.
stream.net Functions with example data for creat-
ing, importing, attributing, analyzing, and dis-
playing stream networks represented as bi-
nary trees. Capabilities include upstream
and downstream distance matrices, stochastic
network generation, segmentation of network
into reaches, adding attributes to reaches with
specified statistical distributions, interpolating
reach attributes from sparse data, analyzing
auto-correlation of reach attributes, and creat-
ing maps with legends of attribute data. Tar-
get applications include dynamic fish model-
ing. By Denis White.
svcm Fir 2-d and 3-d space-varying coefficient mod-
els to regular grid data using either a full B-
spline tensor product approach or a sequen-
tial approximation. By Susanne Heim, with
support from Paul Eilers, Thomas Kneib, and
Michael Kobl.
R News ISSN 1609-3631

Vol. 7/1, April 2007 56
tdm A tool for therapeutic drug monitoring. Can es-
timate individual pharmacokinetic parameters
with one or more drug serum/plasma concen-
trations obtained from a single subject or mul-
tiple subjects, calculate a suggested dose with
the target drug concentration or calculate a pre-
dicted drug concentration with a given dose.
By Miou-Ting Chen, Yung-Jin Lee.
titan GUI to analyze mass spectrometric data on the
relative abundance of two substances from a
titration series. By Tom Price.
tkrgl TK widget tools for package rgl. By Duncan
Murdoch and Ming Chen.
tm A framework for text mining applications within
R. By Ingo Feinerer.
trip Access and manipulate spatial data for animal
tracking. By Michael D. Sumner.
wombsoft Wombling Computation. Analyses indi-
vidually geo-referenced multilocus genotypes
for the inferences of genetic boundaries be-
tween populations. Based on the wombling
method which estimates the systemic function
by looking for the local variation of the allele
frequencies. By Ameline Crida.
yaImpute Perform k-NN imputation. By Nicholas
L. Crookston and Andrew O. Finley.
Other changes
• Package micEcdat was moved from the main
CRAN section to the Archive.
• Package SAGx was removed from CRAN.
Kurt Hornik
Wirtschaftsuniversität Wien, Austria
Kurt.Hornik@R-project.org
R Foundation News
by Kurt Hornik
Donations and new members
Donations
Peter M Maksymuk (USA)
New benefactors
Schröder Investment Management Ltd., London,
GB
Prediction Company, Santa Fe, New Mexico, USA
New supporting institutions
Ef-prime Inc., Tokyo, Japan
New supporting members
David Vonka (CZ)
Olivier Celhay (France)
Katarzyna Kopczewska (Poland)
Kurt Hornik
Wirtschaftsuniversität Wien, Austria
Kurt.Hornik@R-project.org
R News Referees 2006
by Torsten Hothorn
R News articles are peer-reviewed by external and
independent experts. The editorial board mem-
bers would like to take the opportunity to thank
all referees who read and commented on submitted
manuscripts during the last year. Much of the qual-
ity of R News publications is due to their invaluable
and timely service. Thank you!
• Murray Aitkin
• Doug Bates
• Axel Benner
• Adrian Bowman
• Patrick Burns
• Philip Dixon
• Romain Francois
• Thomas Gerds
• Jos Hageman
• Frank Harrell
• David James
• Sigbert Klinke
R News ISSN 1609-3631

Vol. 7/1, April 2007 57
• Eric Lecoutre
• Fritz Leisch
• Martin Maechler
• Andrew Martin
• Georges Monette
• Martyn Plummer
• Matt Pocernich
• Christina Rabe
• Tony Rossini
• Peter Ruckdeschl
• Alec Stephenson
• Simon Urbanek
• Bill Venables
• Ron Wehrens
• Keith Worsley
• Achim Zeileis
Torsten Hothorn
Friedrich–Alexander–Universität Erlangen–Nürnberg
Germany
Torsten.Hothorn@R-project.org
R News ISSN 1609-3631

Vol. 7/1, April 2007 58
Editor-in-Chief:
Torsten Hothorn
Institut für Statistik
Ludwigs–Maximilians–Universität München
Ludwigstraße 33, D-80539 München
Germany
Editorial Board:
John Fox and Vincent Carey.
Editor Programmer’s Niche:
Bill Venables
Editor Help Desk:
Uwe Ligges
Email of editors and editorial board:
firstname.lastname @R-project.org
R News is a publication of the R Foundation for Sta-
tistical Computing. Communications regarding this
publication should be addressed to the editors. All
articles are copyrighted by the respective authors.
Please send submissions to regular columns to the
respective column editor and all other submissions
to the editor-in-chief or another member of the edi-
torial board. More detailed submission instructions
can be found on the R homepage.
R Project Homepage:
http://www.R-project.org/
This newsletter is available online at
http://CRAN.R-project.org/doc/Rnews/
R News ISSN 1609-3631