Programmers Guide
User Manual: Pdf
Open the PDF directly: View PDF ![]() .
.
Page Count: 92
- Estimators
- Importing Data
- Introduction
- Tensors
- Graphs and Sessions
- Saving and Restoring
- Using GPUs
- Embeddings
- Debugging TensorFlow Programs
- TensorBoard: Visualizing Learning
- TensorBoard: Graph Visualization

Estimators
This document introduces Estimators--a high-level TensorFlow API that greatly simplifies machine learning programming.
Estimators encapsulate the following actions:
training
evaluation
prediction
export for serving
You may either use the pre-made Estimators we provide or write your own custom Estimators. All Estimators--whether pre-
made or custom--are classes based on the tf.estimator.Estimator class.
Note: TensorFlow also includes a deprecated Estimator class at tf.contrib.learn.Estimator, which you should not use.
Advantages of Estimators
Estimators provide the following benefits:
You can run Estimators-based models on a local host or on a distributed multi-server environment without changing your
model. Furthermore, you can run Estimators-based models on CPUs, GPUs, or TPUs without recoding your model.
Estimators simplify sharing implementations between model developers.
You can develop a state of the art model with high-level intuitive code, In short, it is generally much easier to create models
with Estimators than with the low-level TensorFlow APIs.
Estimators are themselves built on tf.layers, which simplifies customization.
Estimators build the graph for you. In other words, you don't have to build the graph.
Estimators provide a safe distributed training loop that controls how and when to:
build the graph
initialize variables
start queues
handle exceptions
create checkpoint files and recover from failures
save summaries for TensorBoard
When writing an application with Estimators, you must separate the data input pipeline from the model. This separation
simplifies experiments with different data sets.
Pre-made Estimators
Pre-made Estimators enable you to work at a much higher conceptual level than the base TensorFlow APIs. You no longer have
to worry about creating the computational graph or sessions since Estimators handle all the "plumbing" for you. That is, pre-
made Estimators create and manage Graph and Session objects for you. Furthermore, pre-made Estimators let you
experiment with different model architectures by making only minimal code changes. DNNClassifier, for example, is a pre-
made Estimator class that trains classification models through dense, feed-forward neural networks.
Structure of a pre-made Estimators program

A TensorFlow program relying on a pre-made Estimator typically consists of the following four steps:
1. Write one or more dataset importing functions. For example, you might create one function to import the training set and
another function to import the test set. Each dataset importing function must return two objects:
a dictionary in which the keys are feature names and the values are Tensors (or SparseTensors) containing the
corresponding feature data
a Tensor containing one or more labels
For example, the following code illustrates the basic skeleton for an input function:
(See Importing Data for full details.)
2. Define the feature columns. Each tf.feature_column identifies a feature name, its type, and any input pre-processing.
For example, the following snippet creates three feature columns that hold integer or floating-point data. The first two
feature columns simply identify the feature's name and type. The third feature column also specifies a lambda the program
will invoke to scale the raw data:
3. Instantiate the relevant pre-made Estimator. For example, here's a sample instantiation of a pre-made Estimator named
LinearClassifier:
4. Call a training, evaluation, or inference method. For example, all Estimators provide a train method, which trains a
model.
Benefits of pre-made Estimators
Pre-made Estimators encode best practices, providing the following benefits:
Best practices for determining where different parts of the computational graph should run, implementing strategies on a
single machine or on a cluster.
Best practices for event (summary) writing and universally useful summaries.
If you don't use pre-made Estimators, you must implement the preceding features yourself.
Custom Estimators
def input_fn(dataset):
... # manipulate dataset, extracting feature names and the label
return feature_dict, label
# Define three numeric feature columns.
population = tf.feature_column.numeric_column('population')
crime_rate = tf.feature_column.numeric_column('crime_rate')
median_education = tf.feature_column.numeric_column('median_education',
normalizer_fn='lambda x: x - global_education_mean')
# Instantiate an estimator, passing the feature columns.
estimator = tf.estimator.Estimator.LinearClassifier(
feature_columns=[population, crime_rate, median_education],
)
# my_training_set is the function created in Step 1
estimator.train(input_fn=my_training_set, steps=2000)

The heart of every Estimator--whether pre-made or custom--is its model function, which is a method that builds graphs for
training, evaluation, and prediction. When you are using a pre-made Estimator, someone else has already implemented the
model function. When relying on a custom Estimator, you must write the model function yourself. A companion document
explains how to write the model function.
Recommended workflow
We recommend the following workflow:
1. Assuming a suitable pre-made Estimator exists, use it to build your first model and use its results to establish a baseline.
2. Build and test your overall pipeline, including the integrity and reliability of your data with this pre-made Estimator.
3. If suitable alternative pre-made Estimators are available, run experiments to determine which pre-made Estimator produces
the best results.
4. Possibly, further improve your model by building your own custom Estimator.
Creating Estimators from Keras models
You can convert existing Keras models to Estimators. Doing so enables your Keras model to access Estimator's strengths, such
as distributed training. Call tf.keras.estimator.model_to_estimator as in the following sample:
Note that the names of feature columns and labels of a keras estimator come from the corresponding compiled keras model.
For example, the input key names for train_input_fn above can be obtained from keras_inception_v3.input_names, and
similarly, the predicted output names can be obtained fromkeras_inception_v3.output_names.
For more details, please refer to the documentation for tf.keras.estimator.model_to_estimator.
Importing Data
# Instantiate a Keras inception v3 model.
keras_inception_v3 = tf.keras.applications.inception_v3.InceptionV3(weights=None)
# Compile model with the optimizer, loss, and metrics you'd like to train with.
keras_inception_v3.compile(optimizer=tf.keras.optimizers.SGD(lr=0.0001, momentum=0.9),
loss='categorical_crossentropy',
metric='accuracy')
# Create an Estimator from the compiled Keras model. Note the initial model
# state of the keras model is preserved in the created Estimator.
est_inception_v3 = tf.keras.estimator.model_to_estimator(keras_model=keras_inception_v3)
# Treat the derived Estimator as you would with any other Estimator.
# First, recover the input name(s) of Keras model, so we can use them as the
# feature column name(s) of the Estimator input function:
keras_inception_v3.input_names # print out: ['input_1']
# Once we have the input name(s), we can create the input function, for example,
# for input(s) in the format of numpy ndarray:
train_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"input_1": train_data},
y=train_labels,
num_epochs=1,
shuffle=False)
# To train, we call Estimator's train function:
est_inception_v3.train(input_fn=train_input_fn, steps=2000)

The tf.data API enables you to build complex input pipelines from simple, reusable pieces. For example, the pipeline for an
image model might aggregate data from files in a distributed file system, apply random perturbations to each image, and
merge randomly selected images into a batch for training. The pipeline for a text model might involve extracting symbols from
raw text data, converting them to embedding identifiers with a lookup table, and batching together sequences of different
lengths. The tf.data API makes it easy to deal with large amounts of data, different data formats, and complicated
transformations.
The tf.data API introduces two new abstractions to TensorFlow:
A tf.data.Dataset represents a sequence of elements, in which each element contains one or more Tensor objects. For
example, in an image pipeline, an element might be a single training example, with a pair of tensors representing the image
data and a label. There are two distinct ways to create a dataset:
Creating a source (e.g. Dataset.from_tensor_slices()) constructs a dataset from one or more tf.Tensor objects.
Applying a transformation (e.g. Dataset.batch()) constructs a dataset from one or more tf.data.Dataset objects.
A tf.data.Iterator provides the main way to extract elements from a dataset. The operation returned by
Iterator.get_next() yields the next element of a Dataset when executed, and typically acts as the interface between
input pipeline code and your model. The simplest iterator is a "one-shot iterator", which is associated with a particular
Dataset and iterates through it once. For more sophisticated uses, theIterator.initializer operation enables you to
reinitialize and parameterize an iterator with different datasets, so that you can, for example, iterate over training and
validation data multiple times in the same program.
Basic mechanics
This section of the guide describes the fundamentals of creating different kinds of Dataset and Iteratorobjects, and how to
extract data from them.
To start an input pipeline, you must define a source. For example, to construct a Dataset from some tensors in memory, you
can use tf.data.Dataset.from_tensors() or tf.data.Dataset.from_tensor_slices(). Alternatively, if your input data
are on disk in the recommended TFRecord format, you can construct atf.data.TFRecordDataset.
Once you have a Dataset object, you can transform it into a new Dataset by chaining method calls on the tf.data.Dataset
object. For example, you can apply per-element transformations such as Dataset.map()(to apply a function to each element),
and multi-element transformations such as Dataset.batch(). See the documentation for tf.data.Dataset for a complete
list of transformations.
The most common way to consume values from a Dataset is to make an iterator object that provides access to one element of
the dataset at a time (for example, by calling Dataset.make_one_shot_iterator()). Atf.data.Iterator provides two
operations: Iterator.initializer, which enables you to (re)initialize the iterator's state; and Iterator.get_next(), which
returns tf.Tensor objects that correspond to the symbolic next element. Depending on your use case, you might choose a
different type of iterator, and the options are outlined below.
Dataset structure
A dataset comprises elements that each have the same structure. An element contains one or more tf.Tensor objects, called
components. Each component has a tf.DType representing the type of elements in the tensor, and a tf.TensorShape
representing the (possibly partially specified) static shape of each element. The Dataset.output_types and
Dataset.output_shapes properties allow you to inspect the inferred types and shapes of each component of a dataset
element. The nested structure of these properties map to the structure of an element, which may be a single tensor, a tuple of
tensors, or a nested tuple of tensors. For example:

It is often convenient to give names to each component of an element, for example if they represent different features of a
training example. In addition to tuples, you can use collections.namedtuple or a dictionary mapping strings to tensors to
represent a single element of a Dataset.
The Dataset transformations support datasets of any structure. When using the Dataset.map(), Dataset.flat_map(), and
Dataset.filter() transformations, which apply a function to each element, the element structure determines the arguments
of the function:
Creating an iterator
Once you have built a Dataset to represent your input data, the next step is to create an Iterator to access elements from that
dataset. The tf.data API currently supports the following iterators, in increasing level of sophistication:
one-shot,
initializable,
reinitializable, and
feedable.
A one-shot iterator is the simplest form of iterator, which only supports iterating once through a dataset, with no need for
explicit initialization. One-shot iterators handle almost all of the cases that the existing queue-based input pipelines support,
but they do not support parameterization. Using the example of Dataset.range():
dataset1 = tf.data.Dataset.from_tensor_slices(tf.random_uniform([4, 10]))
print(dataset1.output_types) # ==> "tf.float32"
print(dataset1.output_shapes) # ==> "(10,)"
dataset2 = tf.data.Dataset.from_tensor_slices(
(tf.random_uniform([4]),
tf.random_uniform([4, 100], maxval=100, dtype=tf.int32)))
print(dataset2.output_types) # ==> "(tf.float32, tf.int32)"
print(dataset2.output_shapes) # ==> "((), (100,))"
dataset3 = tf.data.Dataset.zip((dataset1, dataset2))
print(dataset3.output_types) # ==> (tf.float32, (tf.float32, tf.int32))
print(dataset3.output_shapes) # ==> "(10, ((), (100,)))"
dataset = tf.data.Dataset.from_tensor_slices(
{"a": tf.random_uniform([4]),
"b": tf.random_uniform([4, 100], maxval=100, dtype=tf.int32)})
print(dataset.output_types) # ==> "{'a': tf.float32, 'b': tf.int32}"
print(dataset.output_shapes) # ==> "{'a': (), 'b': (100,)}"
dataset1 = dataset1.map(lambda x: ...)
dataset2 = dataset2.flat_map(lambda x, y: ...)
# Note: Argument destructuring is not available in Python 3.
dataset3 = dataset3.filter(lambda x, (y, z): ...)

Note: Currently, one-shot iterators are the only type that is easily usable with an Estimator.
An initializable iterator requires you to run an explicit iterator.initializer operation before using it. In exchange for this
inconvenience, it enables you to parameterize the definition of the dataset, using one or more tf.placeholder() tensors that
can be fed when you initialize the iterator. Continuing the Dataset.range()example:
A reinitializable iterator can be initialized from multiple different Dataset objects. For example, you might have a training
input pipeline that uses random perturbations to the input images to improve generalization, and a validation input pipeline
that evaluates predictions on unmodified data. These pipelines will typically use different Dataset objects that have the same
structure (i.e. the same types and compatible shapes for each component).
dataset = tf.data.Dataset.range(100)
iterator = dataset.make_one_shot_iterator()
next_element = iterator.get_next()
for i in range(100):
value = sess.run(next_element)
assert i == value
max_value = tf.placeholder(tf.int64, shape=[])
dataset = tf.data.Dataset.range(max_value)
iterator = dataset.make_initializable_iterator()
next_element = iterator.get_next()
# Initialize an iterator over a dataset with 10 elements.
sess.run(iterator.initializer, feed_dict={max_value: 10})
for i in range(10):
value = sess.run(next_element)
assert i == value
# Initialize the same iterator over a dataset with 100 elements.
sess.run(iterator.initializer, feed_dict={max_value: 100})
for i in range(100):
value = sess.run(next_element)
assert i == value

A feedable iterator can be used together with tf.placeholder to select what Iterator to use in each call to
tf.Session.run, via the familiar feed_dict mechanism. It offers the same functionality as a reinitializable iterator, but it
does not require you to initialize the iterator from the start of a dataset when you switch between iterators. For example, using
the same training and validation example from above, you can usetf.data.Iterator.from_string_handle to define a
feedable iterator that allows you to switch between the two datasets:
# Define training and validation datasets with the same structure.
training_dataset = tf.data.Dataset.range(100).map(
lambda x: x + tf.random_uniform([], -10, 10, tf.int64))
validation_dataset = tf.data.Dataset.range(50)
# A reinitializable iterator is defined by its structure. We could use the
# `output_types` and `output_shapes` properties of either `training_dataset`
# or `validation_dataset` here, because they are compatible.
iterator = tf.data.Iterator.from_structure(training_dataset.output_types,
training_dataset.output_shapes)
next_element = iterator.get_next()
training_init_op = iterator.make_initializer(training_dataset)
validation_init_op = iterator.make_initializer(validation_dataset)
# Run 20 epochs in which the training dataset is traversed, followed by the
# validation dataset.
for _ in range(20):
# Initialize an iterator over the training dataset.
sess.run(training_init_op)
for _ in range(100):
sess.run(next_element)
# Initialize an iterator over the validation dataset.
sess.run(validation_init_op)
for _ in range(50):
sess.run(next_element)

Consuming values from an iterator
The Iterator.get_next() method returns one or more tf.Tensor objects that correspond to the symbolic next element of
an iterator. Each time these tensors are evaluated, they take the value of the next element in the underlying dataset. (Note that,
like other stateful objects in TensorFlow, calling Iterator.get_next() does not immediately advance the iterator. Instead
you must use the returned tf.Tensor objects in a TensorFlow expression, and pass the result of that expression to
tf.Session.run() to get the next elements and advance the iterator.)
If the iterator reaches the end of the dataset, executing the Iterator.get_next() operation will raise a
tf.errors.OutOfRangeError. After this point the iterator will be in an unusable state, and you must initialize it again if you
want to use it further.
# Define training and validation datasets with the same structure.
training_dataset = tf.data.Dataset.range(100).map(
lambda x: x + tf.random_uniform([], -10, 10, tf.int64)).repeat()
validation_dataset = tf.data.Dataset.range(50)
# A feedable iterator is defined by a handle placeholder and its structure. We
# could use the `output_types` and `output_shapes` properties of either
# `training_dataset` or `validation_dataset` here, because they have
# identical structure.
handle = tf.placeholder(tf.string, shape=[])
iterator = tf.data.Iterator.from_string_handle(
handle, training_dataset.output_types, training_dataset.output_shapes)
next_element = iterator.get_next()
# You can use feedable iterators with a variety of different kinds of iterator
# (such as one-shot and initializable iterators).
training_iterator = training_dataset.make_one_shot_iterator()
validation_iterator = validation_dataset.make_initializable_iterator()
# The `Iterator.string_handle()` method returns a tensor that can be evaluated
# and used to feed the `handle` placeholder.
training_handle = sess.run(training_iterator.string_handle())
validation_handle = sess.run(validation_iterator.string_handle())
# Loop forever, alternating between training and validation.
while True:
# Run 200 steps using the training dataset. Note that the training dataset is
# infinite, and we resume from where we left off in the previous `while` loop
# iteration.
for _ in range(200):
sess.run(next_element, feed_dict={handle: training_handle})
# Run one pass over the validation dataset.
sess.run(validation_iterator.initializer)
for _ in range(50):
sess.run(next_element, feed_dict={handle: validation_handle})

A common pattern is to wrap the "training loop" in a try-except block:
If each element of the dataset has a nested structure, the return value of Iterator.get_next() will be one or more
tf.Tensor objects in the same nested structure:
Note that evaluating any of next1, next2, or next3 will advance the iterator for all components. A typical consumer of an
iterator will include all components in a single expression.
Reading input data
Consuming NumPy arrays
If all of your input data fit in memory, the simplest way to create a Dataset from them is to convert them to tf.Tensor objects
and use Dataset.from_tensor_slices().
dataset = tf.data.Dataset.range(5)
iterator = dataset.make_initializable_iterator()
next_element = iterator.get_next()
# Typically `result` will be the output of a model, or an optimizer's
# training operation.
result = tf.add(next_element, next_element)
sess.run(iterator.initializer)
print(sess.run(result)) # ==> "0"
print(sess.run(result)) # ==> "2"
print(sess.run(result)) # ==> "4"
print(sess.run(result)) # ==> "6"
print(sess.run(result)) # ==> "8"
try:
sess.run(result)
except tf.errors.OutOfRangeError:
print("End of dataset") # ==> "End of dataset"
sess.run(iterator.initializer)
while True:
try:
sess.run(result)
except tf.errors.OutOfRangeError:
break
dataset1 = tf.data.Dataset.from_tensor_slices(tf.random_uniform([4, 10]))
dataset2 = tf.data.Dataset.from_tensor_slices((tf.random_uniform([4]), tf.random_uniform([4,
100])))
dataset3 = tf.data.Dataset.zip((dataset1, dataset2))
iterator = dataset3.make_initializable_iterator()
sess.run(iterator.initializer)
next1, (next2, next3) = iterator.get_next()

Note that the above code snippet will embed the features and labels arrays in your TensorFlow graph as tf.constant()
operations. This works well for a small dataset, but wastes memory---because the contents of the array will be copied multiple
times---and can run into the 2GB limit for the tf.GraphDef protocol buffer.
As an alternative, you can define the Dataset in terms of tf.placeholder() tensors, and feed the NumPy arrays when you
initialize an Iterator over the dataset.
Consuming TFRecord data
The tf.data API supports a variety of file formats so that you can process large datasets that do not fit in memory. For
example, the TFRecord file format is a simple record-oriented binary format that many TensorFlow applications use for
training data. The tf.data.TFRecordDataset class enables you to stream over the contents of one or more TFRecord files as
part of an input pipeline.
The filenames argument to the TFRecordDataset initializer can either be a string, a list of strings, or a tf.Tensor of strings.
Therefore if you have two sets of files for training and validation purposes, you can use atf.placeholder(tf.string) to
represent the filenames, and initialize an iterator from the appropriate filenames:
# Load the training data into two NumPy arrays, for example using `np.load()`.
with np.load("/var/data/training_data.npy") as data:
features = data["features"]
labels = data["labels"]
# Assume that each row of `features` corresponds to the same row as `labels`.
assert features.shape[0] == labels.shape[0]
dataset = tf.data.Dataset.from_tensor_slices((features, labels))
# Load the training data into two NumPy arrays, for example using `np.load()`.
with np.load("/var/data/training_data.npy") as data:
features = data["features"]
labels = data["labels"]
# Assume that each row of `features` corresponds to the same row as `labels`.
assert features.shape[0] == labels.shape[0]
features_placeholder = tf.placeholder(features.dtype, features.shape)
labels_placeholder = tf.placeholder(labels.dtype, labels.shape)
dataset = tf.data.Dataset.from_tensor_slices((features_placeholder, labels_placeholder))
# [Other transformations on `dataset`...]
dataset = ...
iterator = dataset.make_initializable_iterator()
sess.run(iterator.initializer, feed_dict={features_placeholder: features,
labels_placeholder: labels})
# Creates a dataset that reads all of the examples from two files.
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)

Consuming text data
Many datasets are distributed as one or more text files. The tf.data.TextLineDataset provides an easy way to extract lines
from one or more text files. Given one or more filenames, a TextLineDataset will produce one string-valued element per line
of those files. Like a TFRecordDataset, TextLineDataset accepts filenamesas a tf.Tensor, so you can parameterize it by
passing a tf.placeholder(tf.string).
By default, a TextLineDataset yields every line of each file, which may not be desirable, for example if the file starts with a
header line, or contains comments. These lines can be removed using the Dataset.skip() andDataset.filter()
transformations. To apply these transformations to each file separately, we use Dataset.flat_map() to create a nested
Dataset for each file.
Preprocessing data with Dataset.map()
filenames = tf.placeholder(tf.string, shape=[None])
dataset = tf.data.TFRecordDataset(filenames)
dataset = dataset.map(...) # Parse the record into tensors.
dataset = dataset.repeat() # Repeat the input indefinitely.
dataset = dataset.batch(32)
iterator = dataset.make_initializable_iterator()
# You can feed the initializer with the appropriate filenames for the current
# phase of execution, e.g. training vs. validation.
# Initialize `iterator` with training data.
training_filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
sess.run(iterator.initializer, feed_dict={filenames: training_filenames})
# Initialize `iterator` with validation data.
validation_filenames = ["/var/data/validation1.tfrecord", ...]
sess.run(iterator.initializer, feed_dict={filenames: validation_filenames})
filenames = ["/var/data/file1.txt", "/var/data/file2.txt"]
dataset = tf.data.TextLineDataset(filenames)
filenames = ["/var/data/file1.txt", "/var/data/file2.txt"]
dataset = tf.data.Dataset.from_tensor_slices(filenames)
# Use `Dataset.flat_map()` to transform each file as a separate nested dataset,
# and then concatenate their contents sequentially into a single "flat" dataset.
# * Skip the first line (header row).
# * Filter out lines beginning with "#" (comments).
dataset = dataset.flat_map(
lambda filename: (
tf.data.TextLineDataset(filename)
.skip(1)
.filter(lambda line: tf.not_equal(tf.substr(line, 0, 1), "#"))))

The Dataset.map(f) transformation produces a new dataset by applying a given function f to each element of the input
dataset. It is based on the map() function that is commonly applied to lists (and other structures) in functional programming
languages. The function f takes the tf.Tensor objects that represent a single element in the input, and returns the tf.Tensor
objects that will represent a single element in the new dataset. Its implementation uses standard TensorFlow operations to
transform one element into another.
This section covers common examples of how to use Dataset.map().
Parsing tf.Example protocol buffer messages
Many input pipelines extract tf.train.Example protocol buffer messages from a TFRecord-format file (written, for example,
using tf.python_io.TFRecordWriter). Each tf.train.Example record contains one or more "features", and the input
pipeline typically converts these features into tensors.
Decoding image data and resizing it
When training a neural network on real-world image data, it is often necessary to convert images of different sizes to a
common size, so that they may be batched into a fixed size.
# Transforms a scalar string `example_proto` into a pair of a scalar string and
# a scalar integer, representing an image and its label, respectively.
def _parse_function(example_proto):
features = {"image": tf.FixedLenFeature((), tf.string, default_value=""),
"label": tf.FixedLenFeature((), tf.int32, default_value=0)}
parsed_features = tf.parse_single_example(example_proto, features)
return parsed_features["image"], parsed_features["label"]
# Creates a dataset that reads all of the examples from two files, and extracts
# the image and label features.
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
dataset = dataset.map(_parse_function)
# Reads an image from a file, decodes it into a dense tensor, and resizes it
# to a fixed shape.
def _parse_function(filename, label):
image_string = tf.read_file(filename)
image_decoded = tf.image.decode_image(image_string)
image_resized = tf.image.resize_images(image_decoded, [28, 28])
return image_resized, label
# A vector of filenames.
filenames = tf.constant(["/var/data/image1.jpg", "/var/data/image2.jpg", ...])
# `labels[i]` is the label for the image in `filenames[i].
labels = tf.constant([0, 37, ...])
dataset = tf.data.Dataset.from_tensor_slices((filenames, labels))
dataset = dataset.map(_parse_function)

Applying arbitrary Python logic with tf.py_func()
For performance reasons, we encourage you to use TensorFlow operations for preprocessing your data whenever possible.
However, it is sometimes useful to call upon external Python libraries when parsing your input data. To do so, invoke, the
tf.py_func() operation in a Dataset.map() transformation.
Batching dataset elements
Simple batching
The simplest form of batching stacks n consecutive elements of a dataset into a single element. The Dataset.batch()
transformation does exactly this, with the same constraints as the tf.stack() operator, applied to each component of the
elements: i.e. for each component i, all elements must have a tensor of the exact same shape.
Batching tensors with padding
import cv2
# Use a custom OpenCV function to read the image, instead of the standard
# TensorFlow `tf.read_file()` operation.
def _read_py_function(filename, label):
image_decoded = cv2.imread(image_string, cv2.IMREAD_GRAYSCALE)
return image_decoded, label
# Use standard TensorFlow operations to resize the image to a fixed shape.
def _resize_function(image_decoded, label):
image_decoded.set_shape([None, None, None])
image_resized = tf.image.resize_images(image_decoded, [28, 28])
return image_resized, label
filenames = ["/var/data/image1.jpg", "/var/data/image2.jpg", ...]
labels = [0, 37, 29, 1, ...]
dataset = tf.data.Dataset.from_tensor_slices((filenames, labels))
dataset = dataset.map(
lambda filename, label: tuple(tf.py_func(
_read_py_function, [filename, label], [tf.uint8, label.dtype])))
dataset = dataset.map(_resize_function)
inc_dataset = tf.data.Dataset.range(100)
dec_dataset = tf.data.Dataset.range(0, -100, -1)
dataset = tf.data.Dataset.zip((inc_dataset, dec_dataset))
batched_dataset = dataset.batch(4)
iterator = batched_dataset.make_one_shot_iterator()
next_element = iterator.get_next()
print(sess.run(next_element)) # ==> ([0, 1, 2, 3], [ 0, -1, -2, -3])
print(sess.run(next_element)) # ==> ([4, 5, 6, 7], [-4, -5, -6, -7])
print(sess.run(next_element)) # ==> ([8, 9, 10, 11], [-8, -9, -10, -11])

The above recipe works for tensors that all have the same size. However, many models (e.g. sequence models) work with input
data that can have varying size (e.g. sequences of different lengths). To handle this case, theDataset.padded_batch()
transformation enables you to batch tensors of different shape by specifying one or more dimensions in which they may be
padded.
The Dataset.padded_batch() transformation allows you to set different padding for each dimension of each component,
and it may be variable-length (signified by None in the example above) or constant-length. It is also possible to override the
padding value, which defaults to 0.
Training workflows
Processing multiple epochs
The tf.data API offers two main ways to process multiple epochs of the same data.
The simplest way to iterate over a dataset in multiple epochs is to use the Dataset.repeat() transformation. For example, to
create a dataset that repeats its input for 10 epochs:
Applying the Dataset.repeat() transformation with no arguments will repeat the input indefinitely. The
Dataset.repeat() transformation concatenates its arguments without signaling the end of one epoch and the beginning of
the next epoch.
If you want to receive a signal at the end of each epoch, you can write a training loop that catches the
tf.errors.OutOfRangeError at the end of a dataset. At that point you might collect some statistics (e.g. the validation error)
for the epoch.
dataset = tf.data.Dataset.range(100)
dataset = dataset.map(lambda x: tf.fill([tf.cast(x, tf.int32)], x))
dataset = dataset.padded_batch(4, padded_shapes=[None])
iterator = dataset.make_one_shot_iterator()
next_element = iterator.get_next()
print(sess.run(next_element)) # ==> [[0, 0, 0], [1, 0, 0], [2, 2, 0], [3, 3, 3]]
print(sess.run(next_element)) # ==> [[4, 4, 4, 4, 0, 0, 0],
# [5, 5, 5, 5, 5, 0, 0],
# [6, 6, 6, 6, 6, 6, 0],
# [7, 7, 7, 7, 7, 7, 7]]
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
dataset = dataset.map(...)
dataset = dataset.repeat(10)
dataset = dataset.batch(32)

Randomly shuffling input data
The Dataset.shuffle() transformation randomly shuffles the input dataset using a similar algorithm to
tf.RandomShuffleQueue: it maintains a fixed-size buffer and chooses the next element uniformly at random from that buffer.
Using high-level APIs
The tf.train.MonitoredTrainingSession API simplifies many aspects of running TensorFlow in a distributed setting.
MonitoredTrainingSession uses the tf.errors.OutOfRangeError to signal that training has completed, so to use it with
the tf.data API, we recommend usingDataset.make_one_shot_iterator(). For example:
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
dataset = dataset.map(...)
dataset = dataset.batch(32)
iterator = dataset.make_initializable_iterator()
next_element = iterator.get_next()
# Compute for 100 epochs.
for _ in range(100):
sess.run(iterator.initializer)
while True:
try:
sess.run(next_element)
except tf.errors.OutOfRangeError:
break
# [Perform end-of-epoch calculations here.]
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
dataset = dataset.map(...)
dataset = dataset.shuffle(buffer_size=10000)
dataset = dataset.batch(32)
dataset = dataset.repeat()

To use a Dataset in the input_fn of a tf.estimator.Estimator, we also recommend using
Dataset.make_one_shot_iterator(). For example:
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
dataset = dataset.map(...)
dataset = dataset.shuffle(buffer_size=10000)
dataset = dataset.batch(32)
dataset = dataset.repeat(num_epochs)
iterator = dataset.make_one_shot_iterator()
next_example, next_label = iterator.get_next()
loss = model_function(next_example, next_label)
training_op = tf.train.AdagradOptimizer(...).minimize(loss)
with tf.train.MonitoredTrainingSession(...) as sess:
while not sess.should_stop():
sess.run(training_op)
def dataset_input_fn():
filenames = ["/var/data/file1.tfrecord", "/var/data/file2.tfrecord"]
dataset = tf.data.TFRecordDataset(filenames)
# Use `tf.parse_single_example()` to extract data from a `tf.Example`
# protocol buffer, and perform any additional per-record preprocessing.
def parser(record):
keys_to_features = {
"image_data": tf.FixedLenFeature((), tf.string, default_value=""),
"date_time": tf.FixedLenFeature((), tf.int64, default_value=""),
"label": tf.FixedLenFeature((), tf.int64,
default_value=tf.zeros([], dtype=tf.int64)),
}
parsed = tf.parse_single_example(record, keys_to_features)
# Perform additional preprocessing on the parsed data.
image = tf.image.decode_jpeg(parsed["image_data"])
image = tf.reshape(image, [299, 299, 1])
label = tf.cast(parsed["label"], tf.int32)
return {"image_data": image, "date_time": parsed["date_time"]}, label
# Use `Dataset.map()` to build a pair of a feature dictionary and a label
# tensor for each example.
dataset = dataset.map(parser)
dataset = dataset.shuffle(buffer_size=10000)
dataset = dataset.batch(32)
dataset = dataset.repeat(num_epochs)
iterator = dataset.make_one_shot_iterator()
# `features` is a dictionary in which each value is a batch of values for
# that feature; `labels` is a batch of labels.
features, labels = iterator.get_next()
return features, labels

Introduction
This guide gets you started programming in the low-level TensorFlow APIs (TensorFlow Core), showing you how to:
Manage your own TensorFlow program (a tf.Graph) and TensorFlow runtime (a tf.Session), instead of relying on
Estimators to manage them.
Run TensorFlow operations, using a tf.Session.
Use high level components (datasets, layers, and feature_columns) in this low level environment.
Build your own training loop, instead of using the one provided by Estimators.
We recommend using the higher level APIs to build models when possible. Knowing TensorFlow Core is valuable for the
following reasons:
Experimentation and debugging are both more straight forward when you can use low level TensorFlow operations directly.
It gives you a mental model of how things work internally when using the higher level APIs.
Setup
Before using this guide, install TensorFlow.
To get the most out of this guide, you should know the following:
How to program in Python.
At least a little bit about arrays.
Ideally, something about machine learning.
Feel free to launch python and follow along with this walkthrough. Run the following lines to set up your Python
environment:
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import numpy as np
import tensorflow as tf
Tensor Values
The central unit of data in TensorFlow is the tensor. A tensor consists of a set of primitive values shaped into an array of any
number of dimensions. A tensor's rank is its number of dimensions, while its shape is a tuple of integers specifying the array's
length along each dimension. Here are some examples of tensor values:
3. # a rank 0 tensor; a scalar with shape [],
[1., 2., 3.] # a rank 1 tensor; a vector with shape [3]
[[1., 2., 3.], [4., 5., 6.]] # a rank 2 tensor; a matrix with shape [2, 3]
[[[1., 2., 3.]], [[7., 8., 9.]]] # a rank 3 tensor with shape [2, 1, 3]
TensorFlow uses numpy arrays to represent tensor values.
TensorFlow Core Walkthrough

You might think of TensorFlow Core programs as consisting of two discrete sections:
1. Building the computational graph (a tf.Graph).
2. Running the computational graph (using a tf.Session).
Graph
A computational graph is a series of TensorFlow operations arranged into a graph. The graph is composed of two types of
objects.
Operations (or "ops"): The nodes of the graph. Operations describe calculations that consume and produce tensors.
Tensors: The edges in the graph. These represent the values that will flow through the graph. Most TensorFlow functions
return tf.Tensors.
Important: tf.Tensors do not have values, they are just handles to elements in the computation graph.
Let's build a simple computational graph. The most basic operation is a constant. The Python function that builds the operation
takes a tensor value as input. The resulting operation takes no inputs. When run, it outputs the value that was passed to the
constructor. We can create two floating point constants a and b as follows:
a = tf.constant(3.0, dtype=tf.float32)
b = tf.constant(4.0) # also tf.float32 implicitly
total = a + b
print(a)
print(b)
print(total)
The print statements produce:
Tensor("Const:0", shape=(), dtype=float32)
Tensor("Const_1:0", shape=(), dtype=float32)
Tensor("add:0", shape=(), dtype=float32)
Notice that printing the tensors does not output the values 3.0, 4.0, and 7.0 as you might expect. The above statements only
build the computation graph. These tf.Tensor objects just represent the results of the operations that will be run.
Each operation in a graph is given a unique name. This name is independent of the names the objects are assigned to in
Python. Tensors are named after the operation that produces them followed by an output index, as in "add:0" above.
TensorBoard
TensorFlow provides a utility called TensorBoard. One of TensorBoard's many capabilities is visualizing a computation graph.
You can easily do this with a few simple commands.
First you save the computation graph to a TensorBoard summary file as follows:
writer = tf.summary.FileWriter('.')
writer.add_graph(tf.get_default_graph())
This will produce an event file in the current directory with a name in the following format:

events.out.tfevents.{timestamp}.{hostname}
Now, in a new terminal, launch TensorBoard with the following shell command:
tensorboard --logdir .
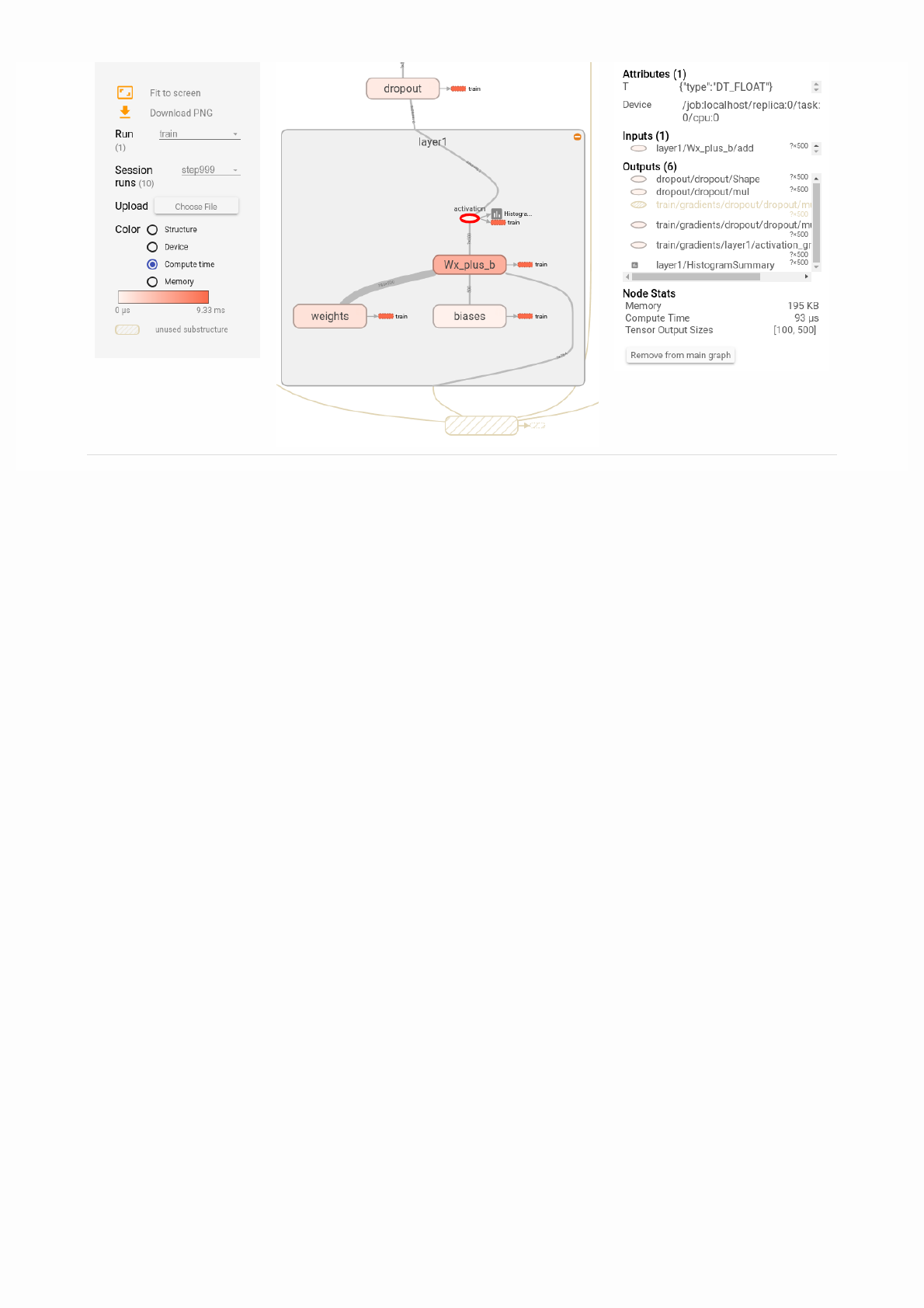
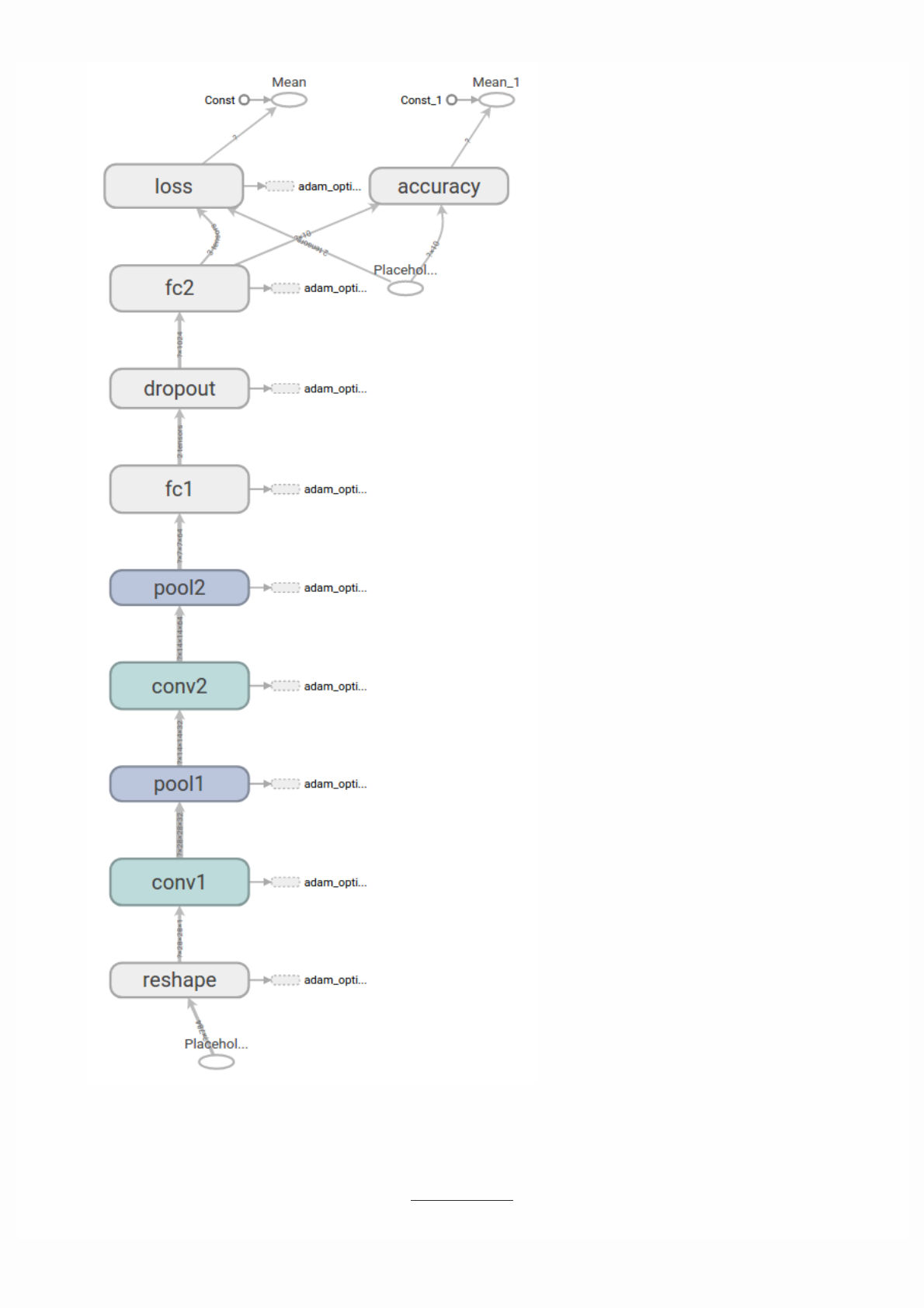
Then open TensorBoard's graphs page in your browser, and you should see a graph similar to the following:
For more about TensorBoard's graph visualization tools see TensorBoard: Graph Visualization.
Session
To evaluate tensors, instantiate a tf.Session object, informally known as a session. A session encapsulates the state of the
TensorFlow runtime, and runs TensorFlow operations. If a tf.Graph is like a .py file, a tf.Session is like the python
executable.
The following code creates a tf.Session object and then invokes its run method to evaluate the totaltensor we created
above:
sess = tf.Session()
print(sess.run(total))
When you request the output of a node with Session.run TensorFlow backtracks through the graph and runs all the nodes
that provide input to the requested output node. So this prints the expected value of 7.0:
7.0
You can pass multiple tensors to tf.Session.run. The run method transparently handles any combination of tuples or
dictionaries, as in the following example:
print(sess.run({'ab':(a, b), 'total':total}))
which returns the results in a structure of the same layout:
{'total': 7.0, 'ab': (3.0, 4.0)}
During a call to tf.Session.run any tf.Tensor only has a single value. For example, the following code calls
tf.random_uniform to produce a tf.Tensor that generates a random 3-element vector (with values in [0,1)):

vec = tf.random_uniform(shape=(3,))
out1 = vec + 1
out2 = vec + 2
print(sess.run(vec))
print(sess.run(vec))
print(sess.run((out1, out2)))
The result shows a different random value on each call to run, but a consistent value during a single run(out1 and out2
receive the same random input):
[ 0.52917576 0.64076328 0.68353939]
[ 0.66192627 0.89126778 0.06254101]
(
array([ 1.88408756, 1.87149239, 1.84057522], dtype=float32),
array([ 2.88408756, 2.87149239, 2.84057522], dtype=float32)
)
Some TensorFlow functions return tf.Operations instead of tf.Tensors. The result of calling run on an Operation is None.
You run an operation to cause a side-effect, not to retrieve a value. Examples of this include the initialization, and training ops
demonstrated later.
Feeding
As it stands, this graph is not especially interesting because it always produces a constant result. A graph can be parameterized
to accept external inputs, known as placeholders. A placeholder is a promise to provide a value later, like a function argument.
x = tf.placeholder(tf.float32)
y = tf.placeholder(tf.float32)
z = x + y
The preceding three lines are a bit like a function in which we define two input parameters (x and y) and then an operation on
them. We can evaluate this graph with multiple inputs by using the feed_dict argument of the run method to feed concrete
values to the placeholders:
print(sess.run(z, feed_dict={x: 3, y: 4.5}))
print(sess.run(z, feed_dict={x: [1, 3], y: [2, 4]}))
This results in the following output:
7.5
[ 3. 7.]
Also note that the feed_dict argument can be used to overwrite any tensor in the graph. The only difference between
placeholders and other tf.Tensors is that placeholders throw an error if no value is fed to them.
Datasets
Placeholders work for simple experiments, but Datasets are the preferred method of streaming data into a model.

To get a runnable tf.Tensor from a Dataset you must first convert it to a tf.data.Iterator, and then call the Iterator's
get_next method.
The simplest way to create an Iterator is with the make_one_shot_iterator method. For example, in the following code the
next_item tensor will return a row from the my_data array on each run call:
my_data = [
[0, 1,],
[2, 3,],
[4, 5,],
[6, 7,],
]
slices = tf.data.Dataset.from_tensor_slices(my_data)
next_item = slices.make_one_shot_iterator().get_next()
Reaching the end of the data stream causes Dataset to throw an OutOfRangeError. For example, the following code reads the
next_item until there is no more data to read:
while True:
try:
print(sess.run(next_item))
except tf.errors.OutOfRangeError:
break
For more details on Datasets and Iterators see: Importing Data.
Layers
A trainable model must modify the values in the graph to get new outputs with the same input. Layers are the preferred way
to add trainable parameters to a graph.
Layers package together both the variables and the operations that act on them, . For example a densely-connected layer
performs a weighted sum across all inputs for each output and applies an optional activation function. The connection weights
and biases are managed by the layer object.
Creating Layers
The following code creates a Dense layer that takes a batch of input vectors, and produces a single output value for each. To
apply a layer to an input, call the layer as if it were a function. For example:
x = tf.placeholder(tf.float32, shape=[None, 3])
linear_model = tf.layers.Dense(units=1)
y = linear_model(x)
The layer inspects its input to determine sizes for its internal variables. So here we must set the shape of the xplaceholder so
that the layer can build a weight matrix of the correct size.
Now that we have defined the calculation of the output, y, there is one more detail we need to take care of before we run the
calculation.

Initializing Layers
The layer contains variables that must be initialized before they can be used. While it is possible to initialize variables
individually, you can easily initialize all the variables in a TensorFlow graph as follows:
init = tf.global_variables_initializer()
sess.run(init)
Important: Calling tf.global_variables_initializer only creates and returns a handle to a TensorFlow operation. That
op will initialize all the global variables when we run it with tf.Session.run.
Also note that this global_variables_initializer only initializes variables that existed in the graph when the initializer
was created. So the initializer should be one of the last things added during graph construction.
Executing Layers
Now that the layer is initialized, we can evaluate the linear_model's output tensor as we would any other tensor. For
example, the following code:
print(sess.run(y, {x: [[1, 2, 3],[4, 5, 6]]}))
will generate a two-element output vector such as the following:
[[-3.41378999]
[-9.14999008]]
Layer Function shortcuts
For each layer class (like tf.layers.Dense) TensorFlow also supplies a shortcut function (like tf.layers.dense). The only
difference is that the shortcut function versions create and run the layer in a single call. For example, the following code is
equivalent to the earlier version:
x = tf.placeholder(tf.float32, shape=[None, 3])
y = tf.layers.dense(x, units=1)
init = tf.global_variables_initializer()
sess.run(init)
print(sess.run(y, {x: [[1, 2, 3], [4, 5, 6]]}))
While convenient, this approach allows no access to the tf.layers.Layer object. This makes introspection and debugging
more difficult, and layer reuse impossible.
Feature columns

The easiest way to experiment with feature columns is using the tf.feature_column.input_layer function. This function
only accepts dense columns as inputs, so to view the result of a categorical column you must wrap it in an
tf.feature_column.indicator_column. For example:
features = {
'sales' : [[5], [10], [8], [9]],
'department': ['sports', 'sports', 'gardening', 'gardening']}
department_column = tf.feature_column.categorical_column_with_vocabulary_list(
'department', ['sports', 'gardening'])
department_column = tf.feature_column.indicator_column(department_column)
columns = [
tf.feature_column.numeric_column('sales'),
department_column
]
inputs = tf.feature_column.input_layer(features, columns)
Running the inputs tensor will parse the features into a batch of vectors.
Feature columns can have internal state, like layers, so they often need to be initialized. Categorical columns use lookup tables
internally and these require a separate initialization op, tf.tables_initializer.
var_init = tf.global_variables_initializer()
table_init = tf.tables_initializer()
sess = tf.Session()
sess.run((var_init, table_init))
Once the internal state has been initialized you can run inputs like any other tf.Tensor:
print(sess.run(inputs))
This shows how the feature columns have packed the input vectors, with the one-hot "department" as the first two indices and
"sales" as the third.
[[ 1. 0. 5.]
[ 1. 0. 10.]
[ 0. 1. 8.]
[ 0. 1. 9.]]
Training
Now that you're familiar with the basics of core TensorFlow, let's train a small regression model manually.
Define the data
First let's define some inputs, x, and the expected output for each input, y_true:

x = tf.constant([[1], [2], [3], [4]], dtype=tf.float32)
y_true = tf.constant([[0], [-1], [-2], [-3]], dtype=tf.float32)
Define the model
Next, build a simple linear model, with 1 output:
linear_model = tf.layers.Dense(units=1)
y_pred = linear_model(x)
You can evaluate the predictions as follows:
sess = tf.Session()
init = tf.global_variables_initializer()
sess.run(init)
print(sess.run(y_pred))
The model hasn't yet been trained, so the four "predicted" values aren't very good. Here's what we got; your own output will
almost certainly differ:
[[ 0.02631879]
[ 0.05263758]
[ 0.07895637]
[ 0.10527515]]
loss
To optimize a model, you first need to define the loss. We'll use the mean square error, a standard loss for regression problems.
While you could do this manually with lower level math operations, the tf.losses module provides a set of common loss
functions. You can use it to calculate the mean square error as follows:
loss = tf.losses.mean_squared_error(labels=y_true, predictions=y_pred)
print(sess.run(loss))
This will produce a loss value, something like:
2.23962
Training
TensorFlow provides optimizers implementing standard optimization algorithms. These are implemented as sub-classes of
tf.train.Optimizer. They incrementally change each variable in order to minimizethe loss. The simplest optimization
algorithm is gradient descent, implemented by tf.train.GradientDescentOptimizer. It modifies each variable according
to the magnitude of the derivative of loss with respect to that variable. For example:
optimizer = tf.train.GradientDescentOptimizer(0.01)
train = optimizer.minimize(loss)
This code builds all the graph components necessary for the optimization, and returns a training operation. When run, the
training op will update variables in the graph. You might run it as follows:
for i in range(100):
_, loss_value = sess.run((train, loss))
print(loss_value)
Since train is an op, not a tensor, it doesn't return a value when run. To see the progression of the loss during training, we run
the loss tensor at the same time, producing output like the following:
1.35659
1.00412
0.759167
0.588829
0.470264
0.387626
0.329918
0.289511
0.261112
0.241046
...
Complete program
x = tf.constant([[1], [2], [3], [4]], dtype=tf.float32)
y_true = tf.constant([[0], [-1], [-2], [-3]], dtype=tf.float32)
linear_model = tf.layers.Dense(units=1)
y_pred = linear_model(x)
loss = tf.losses.mean_squared_error(labels=y_true, predictions=y_pred)
optimizer = tf.train.GradientDescentOptimizer(0.01)
train = optimizer.minimize(loss)
init = tf.global_variables_initializer()
sess = tf.Session()
sess.run(init)
for i in range(100):
_, loss_value = sess.run((train, loss))
print(loss_value)
print(sess.run(y_pred))

Rank Math entity
0 Scalar (magnitude only)
Next steps
To learn more about building models with TensorFlow consider the following:
Custom Estimators, to learn how to build customized models with TensorFlow. Your knowledge of TensorFlow Core will
help you understand and debug your own models.
If you want to learn more about the inner workings of TensorFlow consider the following documents, which go into more
depth on many of the topics discussed here:
Graphs and Sessions
Tensors
Variables
Tensors
TensorFlow, as the name indicates, is a framework to define and run computations involving tensors. A tensoris a
generalization of vectors and matrices to potentially higher dimensions. Internally, TensorFlow represents tensors as n-
dimensional arrays of base datatypes.
When writing a TensorFlow program, the main object you manipulate and pass around is the tf.Tensor. A tf.Tensor object
represents a partially defined computation that will eventually produce a value. TensorFlow programs work by first building a
graph of tf.Tensor objects, detailing how each tensor is computed based on the other available tensors and then by running
parts of this graph to achieve the desired results.
A tf.Tensor has the following properties:
a data type (float32, int32, or string, for example)
a shape
Each element in the Tensor has the same data type, and the data type is always known. The shape (that is, the number of
dimensions it has and the size of each dimension) might be only partially known. Most operations produce tensors of fully-
known shapes if the shapes of their inputs are also fully known, but in some cases it's only possible to find the shape of a
tensor at graph execution time.
Some types of tensors are special, and these will be covered in other units of the Programmer's guide. The main ones are:
tf.Variable
tf.constant
tf.placeholder
tf.SparseTensor
With the exception of tf.Variable, the value of a tensor is immutable, which means that in the context of a single execution
tensors only have a single value. However, evaluating the same tensor twice can return different values; for example that
tensor can be the result of reading data from disk, or generating a random number.
Rank
The rank of a tf.Tensor object is its number of dimensions. Synonyms for rank include order or degree or n-dimension. Note
that rank in TensorFlow is not the same as matrix rank in mathematics. As the following table shows, each rank in TensorFlow
corresponds to a different mathematical entity:

1 Vector (magnitude and direction)
2 Matrix (table of numbers)
3 3-Tensor (cube of numbers)
n n-Tensor (you get the idea)
Rank 0
The following snippet demonstrates creating a few rank 0 variables:
mammal = tf.Variable("Elephant", tf.string)
ignition = tf.Variable(451, tf.int16)
floating = tf.Variable(3.14159265359, tf.float64)
its_complicated = tf.Variable(12.3 - 4.85j, tf.complex64)
Note: A string is treated as a single item in TensorFlow, not as a sequence of characters. It is possible to have scalar strings,
vectors of strings, etc.
Rank 1
To create a rank 1 tf.Tensor object, you can pass a list of items as the initial value. For example:
mystr = tf.Variable(["Hello"], tf.string)
cool_numbers = tf.Variable([3.14159, 2.71828], tf.float32)
first_primes = tf.Variable([2, 3, 5, 7, 11], tf.int32)
its_very_complicated = tf.Variable([12.3 - 4.85j, 7.5 - 6.23j], tf.complex64)
Higher ranks
A rank 2 tf.Tensor object consists of at least one row and at least one column:
mymat = tf.Variable([[7],[11]], tf.int16)
myxor = tf.Variable([[False, True],[True, False]], tf.bool)
linear_squares = tf.Variable([[4], [9], [16], [25]], tf.int32)
squarish_squares = tf.Variable([ [4, 9], [16, 25] ], tf.int32)
rank_of_squares = tf.rank(squarish_squares)
mymatC = tf.Variable([[7],[11]], tf.int32)
Higher-rank Tensors, similarly, consist of an n-dimensional array. For example, during image processing, many tensors of rank
4 are used, with dimensions corresponding to example-in-batch, image width, image height, and color channel.
my_image = tf.zeros([10, 299, 299, 3]) # batch x height x width x color

Rank Shape Dimension number Example
0 [] 0-D A 0-D tensor. A scalar.
1 [D0] 1-D A 1-D tensor with shape [5].
2 [D0, D1] 2-D A 2-D tensor with shape [3, 4].
Getting a tf.Tensor object's rank
To determine the rank of a tf.Tensor object, call the tf.rank method. For example, the following method programmatically
determines the rank of the tf.Tensor defined in the previous section:
r = tf.rank(my_image)
# After the graph runs, r will hold the value 4.
Referring to tf.Tensor slices
Since a tf.Tensor is an n-dimensional array of cells, to access a single cell in a tf.Tensor you need to specify n indices.
For a rank 0 tensor (a scalar), no indices are necessary, since it is already a single number.
For a rank 1 tensor (a vector), passing a single index allows you to access a number:
my_scalar = my_vector[2]
Note that the index passed inside the [] can itself be a scalar tf.Tensor, if you want to dynamically choose an element from
the vector.
For tensors of rank 2 or higher, the situation is more interesting. For a tf.Tensor of rank 2, passing two numbers returns a
scalar, as expected:
my_scalar = my_matrix[1, 2]
Passing a single number, however, returns a subvector of a matrix, as follows:
my_row_vector = my_matrix[2]
my_column_vector = my_matrix[:, 3]
The : notation is python slicing syntax for "leave this dimension alone". This is useful in higher-rank Tensors, as it allows you
to access its subvectors, submatrices, and even other subtensors.
Shape
The shape of a tensor is the number of elements in each dimension. TensorFlow automatically infers shapes during graph
construction. These inferred shapes might have known or unknown rank. If the rank is known, the sizes of each dimension
might be known or unknown.
The TensorFlow documentation uses three notational conventions to describe tensor dimensionality: rank, shape, and
dimension number. The following table shows how these relate to one another:

3 [D0, D1, D2] 3-D A 3-D tensor with shape [1, 4, 3].
n [D0, D1, ... Dn-1] n-D A tensor with shape [D0, D1, ... Dn-1].
Shapes can be represented via Python lists / tuples of ints, or with the tf.TensorShape.
Getting a tf.Tensor object's shape
There are two ways of accessing the shape of a tf.Tensor. While building the graph, it is often useful to ask what is already
known about a tensor's shape. This can be done by reading the shape property of a tf.Tensor object. This method returns a
TensorShape object, which is a convenient way of representing partially-specified shapes (since, when building the graph, not
all shapes will be fully known).
It is also possible to get a tf.Tensor that will represent the fully-defined shape of another tf.Tensor at runtime. This is done
by calling the tf.shape operation. This way, you can build a graph that manipulates the shapes of tensors by building other
tensors that depend on the dynamic shape of the input tf.Tensor.
For example, here is how to make a vector of zeros with the same size as the number of columns in a given matrix:
zeros = tf.zeros(my_matrix.shape[1])
Changing the shape of a tf.Tensor
The number of elements of a tensor is the product of the sizes of all its shapes. The number of elements of a scalar is always 1.
Since there are often many different shapes that have the same number of elements, it's often convenient to be able to change
the shape of a tf.Tensor, keeping its elements fixed. This can be done with tf.reshape.
The following examples demonstrate how to reshape tensors:
rank_three_tensor = tf.ones([3, 4, 5])
matrix = tf.reshape(rank_three_tensor, [6, 10]) # Reshape existing content into
# a 6x10 matrix
matrixB = tf.reshape(matrix, [3, -1]) # Reshape existing content into a 3x20
# matrix. -1 tells reshape to calculate
# the size of this dimension.
matrixAlt = tf.reshape(matrixB, [4, 3, -1]) # Reshape existing content into a
#4x3x5 tensor
# Note that the number of elements of the reshaped Tensors has to match the
# original number of elements. Therefore, the following example generates an
# error because no possible value for the last dimension will match the number
# of elements.
yet_another = tf.reshape(matrixAlt, [13, 2, -1]) # ERROR!
Data types
In addition to dimensionality, Tensors have a data type. Refer to the tf.DataType page in the programmer's guide for a full
list of the data types.

It is not possible to have a tf.Tensor with more than one data type. It is possible, however, to serialize arbitrary data
structures as strings and store those in tf.Tensors.
It is possible to cast tf.Tensors from one datatype to another using tf.cast:
# Cast a constant integer tensor into floating point.
float_tensor = tf.cast(tf.constant([1, 2, 3]), dtype=tf.float32)
To inspect a tf.Tensor's data type use the Tensor.dtype property.
When creating a tf.Tensor from a python object you may optionally specify the datatype. If you don't, TensorFlow chooses a
datatype that can represent your data. TensorFlow converts Python integers to tf.int32 and python floating point numbers to
tf.float32. Otherwise TensorFlow uses the same rules numpy uses when converting to arrays.
Evaluating Tensors
Once the computation graph has been built, you can run the computation that produces a particular tf.Tensorand fetch the
value assigned to it. This is often useful for debugging as well as being required for much of TensorFlow to work.
The simplest way to evaluate a Tensor is using the Tensor.eval method. For example:
constant = tf.constant([1, 2, 3])
tensor = constant * constant
print tensor.eval()
The eval method only works when a default tf.Session is active (see Graphs and Sessions for more information).
Tensor.eval returns a numpy array with the same contents as the tensor.
Sometimes it is not possible to evaluate a tf.Tensor with no context because its value might depend on dynamic information
that is not available. For example, tensors that depend on placeholders can't be evaluated without providing a value for the
placeholder.
p = tf.placeholder(tf.float32)
t = p + 1.0
t.eval() # This will fail, since the placeholder did not get a value.
t.eval(feed_dict={p:2.0}) # This will succeed because we're feeding a value
# to the placeholder.
Note that it is possible to feed any tf.Tensor, not just placeholders.
Other model constructs might make evaluating a tf.Tensor complicated. TensorFlow can't directly evaluate tf.Tensors
defined inside functions or inside control flow constructs. If a tf.Tensor depends on a value from a queue, evaluating the
tf.Tensor will only work once something has been enqueued; otherwise, evaluating it will hang. When working with queues,
remember to call tf.train.start_queue_runners before evaluating any tf.Tensors.
Printing Tensors
For debugging purposes you might want to print the value of a tf.Tensor. While tfdbg provides advanced debugging
support, TensorFlow also has an operation to directly print the value of a tf.Tensor.
Note that you rarely want to use the following pattern when printing a tf.Tensor:

t = <<some tensorflow operation>>
print t # This will print the symbolic tensor when the graph is being built.
# This tensor does not have a value in this context.
This code prints the tf.Tensor object (which represents deferred computation) and not its value. Instead, TensorFlow
provides the tf.Print operation, which returns its first tensor argument unchanged while printing the set of tf.Tensors it is
passed as the second argument.
To correctly use tf.Print its return value must be used. See the example below
t = <<some tensorflow operation>>
tf.Print(t, [t]) # This does nothing
t = tf.Print(t, [t]) # Here we are using the value returned by tf.Print
result = t + 1 # Now when result is evaluated the value of `t` will be printed.
When you evaluate result you will evaluate everything result depends upon. Since result depends upon t, and
evaluating t has the side effect of printing its input (the old value of t), t gets printed.Variables
A TensorFlow variable is the best way to represent shared, persistent state manipulated by your program.
Variables are manipulated via the tf.Variable class. A tf.Variable represents a tensor whose value can be changed by
running ops on it. Unlike tf.Tensor objects, a tf.Variable exists outside the context of a single session.run call.
Internally, a tf.Variable stores a persistent tensor. Specific ops allow you to read and modify the values of this tensor. These
modifications are visible across multiple tf.Sessions, so multiple workers can see the same values for a tf.Variable.
Creating a Variable
The best way to create a variable is to call the tf.get_variable function. This function requires you to specify the Variable's
name. This name will be used by other replicas to access the same variable, as well as to name this variable's value when
checkpointing and exporting models. tf.get_variable also allows you to reuse a previously created variable of the same
name, making it easy to define models which reuse layers.
To create a variable with tf.get_variable, simply provide the name and shape
my_variable = tf.get_variable("my_variable", [1, 2, 3])
This creates a variable named "my_variable" which is a three-dimensional tensor with shape [1, 2, 3]. This variable will, by
default, have the dtype tf.float32 and its initial value will be randomized viatf.glorot_uniform_initializer.
You may optionally specify the dtype and initializer to tf.get_variable. For example:
my_int_variable = tf.get_variable("my_int_variable", [1, 2, 3], dtype=tf.int32,
initializer=tf.zeros_initializer)
TensorFlow provides many convenient initializers. Alternatively, you may initialize a tf.Variable to have the value of a
tf.Tensor. For example:
other_variable = tf.get_variable("other_variable", dtype=tf.int32,
initializer=tf.constant([23, 42]))
Note that when the initializer is a tf.Tensor you should not specify the variable's shape, as the shape of the initializer tensor
will be used.

Variable collections
Because disconnected parts of a TensorFlow program might want to create variables, it is sometimes useful to have a single
way to access all of them. For this reason TensorFlow provides collections, which are named lists of tensors or other objects,
such as tf.Variable instances.
By default every tf.Variable gets placed in the following two collections: * tf.GraphKeys.GLOBAL_VARIABLES --- variables
that can be shared across multiple devices, * tf.GraphKeys.TRAINABLE_VARIABLES--- variables for which TensorFlow will
calculate gradients.
If you don't want a variable to be trainable, add it to the tf.GraphKeys.LOCAL_VARIABLES collection instead. For example,
the following snippet demonstrates how to add a variable named my_local to this collection:
my_local = tf.get_variable("my_local", shape=(),
collections=[tf.GraphKeys.LOCAL_VARIABLES])
Alternatively, you can specify trainable=False as an argument to tf.get_variable:
my_non_trainable = tf.get_variable("my_non_trainable",
shape=(),
trainable=False)
You can also use your own collections. Any string is a valid collection name, and there is no need to explicitly create a
collection. To add a variable (or any other object) to a collection after creating the variable, calltf.add_to_collection. For
example, the following code adds an existing variable named my_local to a collection named my_collection_name:
tf.add_to_collection("my_collection_name", my_local)
And to retrieve a list of all the variables (or other objects) you've placed in a collection you can use:
tf.get_collection("my_collection_name")
Device placement
Just like any other TensorFlow operation, you can place variables on particular devices. For example, the following snippet
creates a variable named v and places it on the second GPU device:
with tf.device("/device:GPU:1"):
v = tf.get_variable("v", [1])
It is particularly important for variables to be in the correct device in distributed settings. Accidentally putting variables on
workers instead of parameter servers, for example, can severely slow down training or, in the worst case, let each worker
blithely forge ahead with its own independent copy of each variable. For this reason we provide
tf.train.replica_device_setter, which can automatically place variables in parameter servers. For example:

cluster_spec = {
"ps": ["ps0:2222", "ps1:2222"],
"worker": ["worker0:2222", "worker1:2222", "worker2:2222"]}
with tf.device(tf.train.replica_device_setter(cluster=cluster_spec)):
v = tf.get_variable("v", shape=[20, 20]) # this variable is placed
# in the parameter server
# by the replica_device_setter
Initializing variables
Before you can use a variable, it must be initialized. If you are programming in the low-level TensorFlow API (that is, you are
explicitly creating your own graphs and sessions), you must explicitly initialize the variables. Most high-level frameworks such
as tf.contrib.slim, tf.estimator.Estimator and Keras automatically initialize variables for you before training a model.
Explicit initialization is otherwise useful because it allows you not to rerun potentially expensive initializers when reloading a
model from a checkpoint as well as allowing determinism when randomly-initialized variables are shared in a distributed
setting.
To initialize all trainable variables in one go, before training starts, calltf.global_variables_initializer(). This function
returns a single operation responsible for initializing all variables in the tf.GraphKeys.GLOBAL_VARIABLES collection.
Running this operation initializes all variables. For example:
session.run(tf.global_variables_initializer())
# Now all variables are initialized.
If you do need to initialize variables yourself, you can run the variable's initializer operation. For example:
session.run(my_variable.initializer)
You can also ask which variables have still not been initialized. For example, the following code prints the names of all
variables which have not yet been initialized:
print(session.run(tf.report_uninitialized_variables()))
Note that by default tf.global_variables_initializer does not specify the order in which variables are initialized.
Therefore, if the initial value of a variable depends on another variable's value, it's likely that you'll get an error. Any time you
use the value of a variable in a context in which not all variables are initialized (say, if you use a variable's value while
initializing another variable), it is best to use variable.initialized_value()instead of variable:
v = tf.get_variable("v", shape=(), initializer=tf.zeros_initializer())
w = tf.get_variable("w", initializer=v.initialized_value() + 1)
Using variables
To use the value of a tf.Variable in a TensorFlow graph, simply treat it like a normal tf.Tensor:

v = tf.get_variable("v", shape=(), initializer=tf.zeros_initializer())
w = v + 1 # w is a tf.Tensor which is computed based on the value of v.
# Any time a variable is used in an expression it gets automatically
# converted to a tf.Tensor representing its value.
To assign a value to a variable, use the methods assign, assign_add, and friends in the tf.Variable class. For example, here
is how you can call these methods:
v = tf.get_variable("v", shape=(), initializer=tf.zeros_initializer())
assignment = v.assign_add(1)
tf.global_variables_initializer().run()
sess.run(assignment) # or assignment.op.run()
Most TensorFlow optimizers have specialized ops that efficiently update the values of variables according to some gradient
descent-like algorithm. See tf.train.Optimizer for an explanation of how to use optimizers.
Because variables are mutable it's sometimes useful to know what version of a variable's value is being used at any point in
time. To force a re-read of the value of a variable after something has happened, you can usetf.Variable.read_value. For
example:
v = tf.get_variable("v", shape=(), initializer=tf.zeros_initializer())
assignment = v.assign_add(1)
with tf.control_dependencies([assignment]):
w = v.read_value() # w is guaranteed to reflect v's value after the
# assign_add operation.
Sharing variables
TensorFlow supports two ways of sharing variables:
Explicitly passing tf.Variable objects around.
Implicitly wrapping tf.Variable objects within tf.variable_scope objects.
While code which explicitly passes variables around is very clear, it is sometimes convenient to write TensorFlow functions
that implicitly use variables in their implementations. Most of the functional layers fromtf.layer use this approach, as well as
all tf.metrics, and a few other library utilities.
Variable scopes allow you to control variable reuse when calling functions which implicitly create and use variables. They also
allow you to name your variables in a hierarchical and understandable way.
For example, let's say we write a function to create a convolutional / relu layer:
def conv_relu(input, kernel_shape, bias_shape):
# Create variable named "weights".
weights = tf.get_variable("weights", kernel_shape,
initializer=tf.random_normal_initializer())
# Create variable named "biases".
biases = tf.get_variable("biases", bias_shape,
initializer=tf.constant_initializer(0.0))
conv = tf.nn.conv2d(input, weights,
strides=[1, 1, 1, 1], padding='SAME')
return tf.nn.relu(conv + biases)

This function uses short names weights and biases, which is good for clarity. In a real model, however, we want many such
convolutional layers, and calling this function repeatedly would not work:
input1 = tf.random_normal([1,10,10,32])
input2 = tf.random_normal([1,20,20,32])
x = conv_relu(input1, kernel_shape=[5, 5, 32, 32], bias_shape=[32])
x = conv_relu(x, kernel_shape=[5, 5, 32, 32], bias_shape = [32]) # This fails.
Since the desired behavior is unclear (create new variables or reuse the existing ones?) TensorFlow will fail. Calling conv_relu
in different scopes, however, clarifies that we want to create new variables:
def my_image_filter(input_images):
with tf.variable_scope("conv1"):
# Variables created here will be named "conv1/weights", "conv1/biases".
relu1 = conv_relu(input_images, [5, 5, 32, 32], [32])
with tf.variable_scope("conv2"):
# Variables created here will be named "conv2/weights", "conv2/biases".
return conv_relu(relu1, [5, 5, 32, 32], [32])
If you do want the variables to be shared, you have two options. First, you can create a scope with the same name using
reuse=True:
with tf.variable_scope("model"):
output1 = my_image_filter(input1)
with tf.variable_scope("model", reuse=True):
output2 = my_image_filter(input2)
You can also call scope.reuse_variables() to trigger a reuse:
with tf.variable_scope("model") as scope:
output1 = my_image_filter(input1)
scope.reuse_variables()
output2 = my_image_filter(input2)
Since depending on exact string names of scopes can feel dangerous, it's also possible to initialize a variable scope based on
another one:
Graphs and Sessions
TensorFlow uses a dataflow graph to represent your computation in terms of the dependencies between individual operations.
This leads to a low-level programming model in which you first define the dataflow graph, then create a TensorFlow session to
run parts of the graph across a set of local and remote devices.
with tf.variable_scope("model") as scope:
output1 = my_image_filter(input1)
with tf.variable_scope(scope, reuse=True):
output2 = my_image_filter(input2)
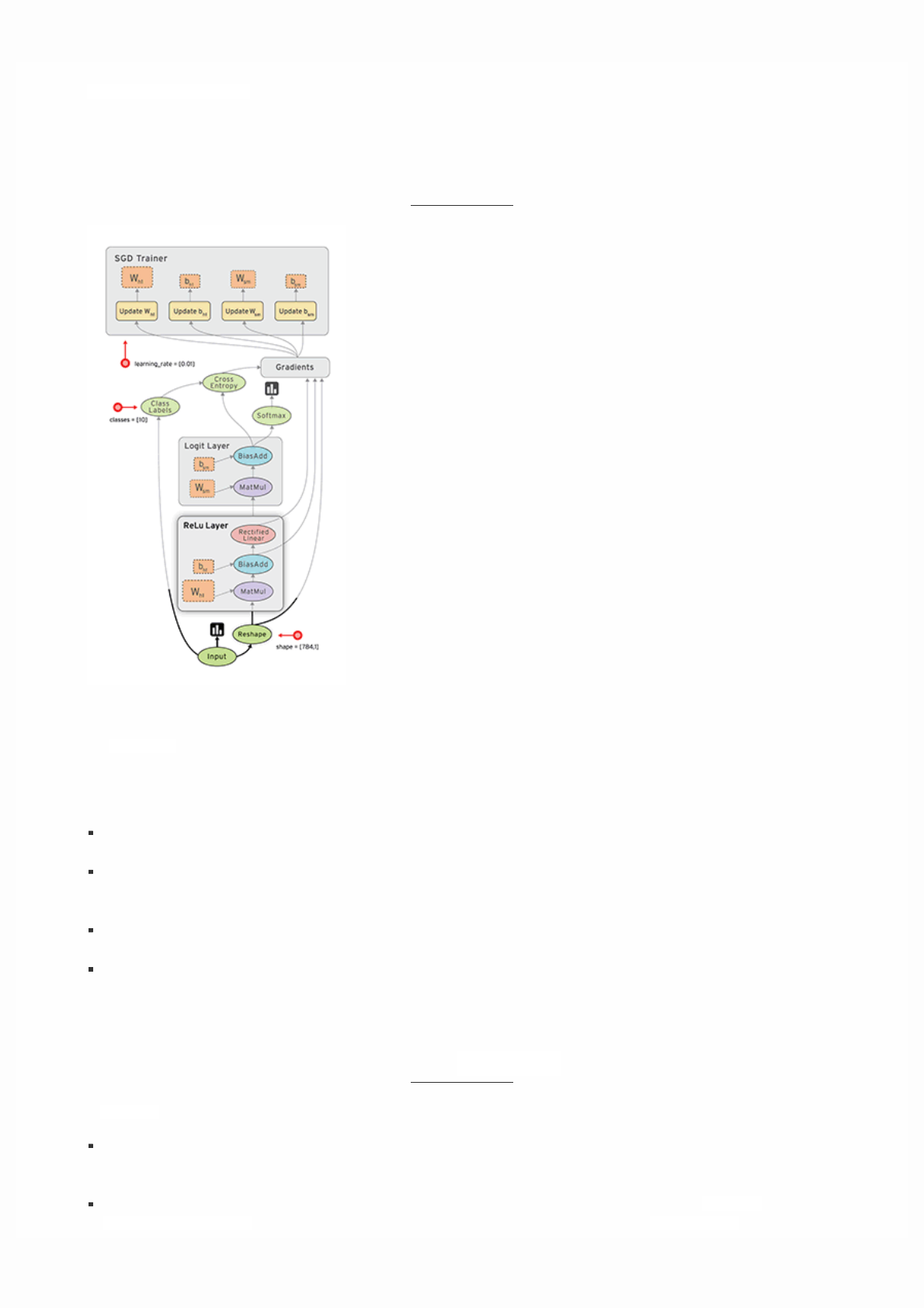
This guide will be most useful if you intend to use the low-level programming model directly. Higher-level APIs such as
tf.estimator.Estimator and Keras hide the details of graphs and sessions from the end user, but this guide may also be
useful if you want to understand how these APIs are implemented.
Why dataflow graphs?
Dataflow is a common programming model for parallel computing. In a dataflow graph, the nodes represent units of
computation, and the edges represent the data consumed or produced by a computation. For example, in a TensorFlow graph,
the tf.matmul operation would correspond to a single node with two incoming edges (the matrices to be multiplied) and one
outgoing edge (the result of the multiplication).
Dataflow has several advantages that TensorFlow leverages when executing your programs:
Parallelism. By using explicit edges to represent dependencies between operations, it is easy for the system to identify
operations that can execute in parallel.
Distributed execution. By using explicit edges to represent the values that flow between operations, it is possible for
TensorFlow to partition your program across multiple devices (CPUs, GPUs, and TPUs) attached to different machines.
TensorFlow inserts the necessary communication and coordination between devices.
Compilation. TensorFlow's XLA compiler can use the information in your dataflow graph to generate faster code, for
example, by fusing together adjacent operations.
Portability. The dataflow graph is a language-independent representation of the code in your model. You can build a
dataflow graph in Python, store it in a SavedModel, and restore it in a C++ program for low-latency inference.
What is a tf.Graph?
A tf.Graph contains two relevant kinds of information:
Graph structure. The nodes and edges of the graph, indicating how individual operations are composed together, but not
prescribing how they should be used. The graph structure is like assembly code: inspecting it can convey some useful
information, but it does not contain all of the useful context that source code conveys.
Graph collections. TensorFlow provides a general mechanism for storing collections of metadata in a tf.Graph. The
tf.add_to_collection function enables you to associate a list of objects with a key (where tf.GraphKeys defines some of

the standard keys), and tf.get_collection enables you to look up all objects associated with a key. Many parts of the
TensorFlow library use this facility: for example, when you create a tf.Variable, it is added by default to collections
representing "global variables" and "trainable variables". When you later come to create a tf.train.Saver or
tf.train.Optimizer, the variables in these collections are used as the default arguments.
Building a tf.Graph
Most TensorFlow programs start with a dataflow graph construction phase. In this phase, you invoke TensorFlow API
functions that construct new tf.Operation (node) and tf.Tensor (edge) objects and add them to a tf.Graph instance.
TensorFlow provides a default graph that is an implicit argument to all API functions in the same context. For example:
Calling tf.constant(42.0) creates a single tf.Operation that produces the value 42.0, adds it to the default graph, and
returns a tf.Tensor that represents the value of the constant.
Calling tf.matmul(x, y) creates a single tf.Operation that multiplies the values of tf.Tensorobjects x and y, adds it to
the default graph, and returns a tf.Tensor that represents the result of the multiplication.
Executing v = tf.Variable(0) adds to the graph a tf.Operation that will store a writeable tensor value that persists
between tf.Session.run calls. The tf.Variable object wraps this operation, and can be used like a tensor, which will
read the current value of the stored value. The tf.Variable object also has methods such as assign and assign_add that
create tf.Operation objects that, when executed, update the stored value. (See Variables for more information about
variables.)
Calling tf.train.Optimizer.minimize will add operations and tensors to the default graph that calculate gradients, and
return a tf.Operation that, when run, will apply those gradients to a set of variables.
Most programs rely solely on the default graph. However, see Dealing with multiple graphs for more advanced use cases.
High-level APIs such as the tf.estimator.Estimator API manage the default graph on your behalf, and--for example--may
create different graphs for training and evaluation.
Note: Calling most functions in the TensorFlow API merely adds operations and tensors to the default graph, but does not
perform the actual computation. Instead, you compose these functions until you have a tf.Tensor or tf.Operation that
represents the overall computation--such as performing one step of gradient descent--and then pass that object to a
tf.Session to perform the computation. See the section "Executing a graph in a tf.Session" for more details.
Naming operations
A tf.Graph object defines a namespace for the tf.Operation objects it contains. TensorFlow automatically chooses a unique
name for each operation in your graph, but giving operations descriptive names can make your program easier to read and
debug. The TensorFlow API provides two ways to override the name of an operation:
Each API function that creates a new tf.Operation or returns a new tf.Tensor accepts an optional name argument. For
example, tf.constant(42.0, name="answer") creates a new tf.Operationnamed "answer" and returns a tf.Tensor
named "answer:0". If the default graph already contained an operation named "answer", the TensorFlow would append
"_1", "_2", and so on to the name, in order to make it unique.
The tf.name_scope function makes it possible to add a name scope prefix to all operations created in a particular context.
The current name scope prefix is a "/"-delimited list of the names of all active tf.name_scope context managers. If a name
scope has already been used in the current context, TensorFlow appens "_1", "_2", and so on. For example:

c_0 = tf.constant(0, name="c") # => operation named "c"
# Already-used names will be "uniquified".
c_1 = tf.constant(2, name="c") # => operation named "c_1"
# Name scopes add a prefix to all operations created in the same context.
with tf.name_scope("outer"):
c_2 = tf.constant(2, name="c") # => operation named "outer/c"
# Name scopes nest like paths in a hierarchical file system.
with tf.name_scope("inner"):
c_3 = tf.constant(3, name="c") # => operation named "outer/inner/c"
# Exiting a name scope context will return to the previous prefix.
c_4 = tf.constant(4, name="c") # => operation named "outer/c_1"
# Already-used name scopes will be "uniquified".
with tf.name_scope("inner"):
c_5 = tf.constant(5, name="c") # => operation named "outer/inner_1/c"
The graph visualizer uses name scopes to group operations and reduce the visual complexity of a graph. See Visualizing your
graph for more information.
Note that tf.Tensor objects are implicitly named after the tf.Operation that produces the tensor as output. A tensor name
has the form "<OP_NAME>:<i>" where:
"<OP_NAME>" is the name of the operation that produces it.
"<i>" is an integer representing the index of that tensor among the operation's outputs.
Placing operations on different devices
If you want your TensorFlow program to use multiple different devices, the tf.device function provides a convenient way to
request that all operations created in a particular context are placed on the same device (or type of device).
A device specification has the following form:
/job:<JOB_NAME>/task:<TASK_INDEX>/device:<DEVICE_TYPE>:<DEVICE_INDEX>
where:
<JOB_NAME> is an alpha-numeric string that does not start with a number.
<DEVICE_TYPE> is a registered device type (such as GPU or CPU).
<TASK_INDEX> is a non-negative integer representing the index of the task in the job named <JOB_NAME>. See
tf.train.ClusterSpec for an explanation of jobs and tasks.
<DEVICE_INDEX> is a non-negative integer representing the index of the device, for example, to distinguish between
different GPU devices used in the same process.
You do not need to specify every part of a device specification. For example, if you are running in a single-machine
configuration with a single GPU, you might use tf.device to pin some operations to the CPU and GPU:

# Operations created outside either context will run on the "best possible"
# device. For example, if you have a GPU and a CPU available, and the operation
# has a GPU implementation, TensorFlow will choose the GPU.
weights = tf.random_normal(...)
with tf.device("/device:CPU:0"):
# Operations created in this context will be pinned to the CPU.
img = tf.decode_jpeg(tf.read_file("img.jpg"))
with tf.device("/device:GPU:0"):
# Operations created in this context will be pinned to the GPU.
result = tf.matmul(weights, img)
If you are deploying TensorFlow in a @{$deploy/distributed$typical distributed configuration}, you might specify the job
name and task ID to place variables on a task in the parameter server job ("/job:ps"), and the other operations on task in the
worker job ("/job:worker"):
with tf.device("/job:ps/task:0"):
weights_1 = tf.Variable(tf.truncated_normal([784, 100]))
biases_1 = tf.Variable(tf.zeroes([100]))
with tf.device("/job:ps/task:1"):
weights_2 = tf.Variable(tf.truncated_normal([100, 10]))
biases_2 = tf.Variable(tf.zeroes([10]))
with tf.device("/job:worker"):
layer_1 = tf.matmul(train_batch, weights_1) + biases_1
layer_2 = tf.matmul(train_batch, weights_2) + biases_2
tf.device gives you a lot of flexibility to choose placements for individual operations or broad regions of a TensorFlow graph.
In many cases, there are simple heuristics that work well. For example, thetf.train.replica_device_setter API can be
used with tf.device to place operations for data-parallel distributed training. For example, the following code fragment
shows how tf.train.replica_device_setter applies different placement policies to tf.Variable objects and other
operations:
with tf.device(tf.train.replica_device_setter(ps_tasks=3)):
# tf.Variable objects are, by default, placed on tasks in "/job:ps" in a
# round-robin fashion.
w_0 = tf.Variable(...) # placed on "/job:ps/task:0"
b_0 = tf.Variable(...) # placed on "/job:ps/task:1"
w_1 = tf.Variable(...) # placed on "/job:ps/task:2"
b_1 = tf.Variable(...) # placed on "/job:ps/task:0"
input_data = tf.placeholder(tf.float32) # placed on "/job:worker"
layer_0 = tf.matmul(input_data, w_0) + b_0 # placed on "/job:worker"
layer_1 = tf.matmul(layer_0, w_1) + b_1 # placed on "/job:worker"
Tensor-like objects
Many TensorFlow operations take one or more tf.Tensor objects as arguments. For example, tf.matmultakes two
tf.Tensor objects, and tf.add_n takes a list of n tf.Tensor objects. For convenience, these functions will accept a tensor-
like object in place of a tf.Tensor, and implicitly convert it to a tf.Tensorusing the tf.convert_to_tensor method.
Tensor-like objects include elements of the following types:
tf.Tensor

tf.Variable
numpy.ndarray
list (and lists of tensor-like objects)
Scalar Python types: bool, float, int, str
You can register additional tensor-like types using tf.register_tensor_conversion_function.
Note: By default, TensorFlow will create a new tf.Tensor each time you use the same tensor-like object. If the tensor-like
object is large (e.g. a numpy.ndarray containing a set of training examples) and you use it multiple times, you may run out of
memory. To avoid this, manually call tf.convert_to_tensor on the tensor-like object once and use the returned tf.Tensor
instead.
Executing a graph in a tf.Session
TensorFlow uses the tf.Session class to represent a connection between the client program---typically a Python program,
although a similar interface is available in other languages---and the C++ runtime. A tf.Session object provides access to
devices in the local machine, and remote devices using the distributed TensorFlow runtime. It also caches information about
your tf.Graph so that you can efficiently run the same computation multiple times.
Creating a tf.Session
If you are using the low-level TensorFlow API, you can create a tf.Session for the current default graph as follows:
# Create a default in-process session.
with tf.Session() as sess:
# ...
# Create a remote session.
with tf.Session("grpc://example.org:2222"):
# ...
Since a tf.Session owns physical resources (such as GPUs and network connections), it is typically used as a context
manager (in a with block) that automatically closes the session when you exit the block. It is also possible to create a session
without using a with block, but you should explicitly call tf.Session.closewhen you are finished with it to free the
resources.
Note: Higher-level APIs such as tf.train.MonitoredTrainingSession or tf.estimator.Estimator will create and
manage a tf.Session for you. These APIs accept optional target and config arguments (either directly, or as part of a
tf.estimator.RunConfig object), with the same meaning as described below.
tf.Session.**init** accepts three optional arguments:
target. If this argument is left empty (the default), the session will only use devices in the local machine. However, you may
also specify a grpc:// URL to specify the address of a TensorFlow server, which gives the session access to all devices on
machines that this server controls. See tf.train.Server for details of how to create a TensorFlow server. For example, in
the common between-graph replicationconfiguration, the tf.Session connects to a tf.train.Server in the same process
as the client. The distributed TensorFlow deployment guide describes other common scenarios.
graph. By default, a new tf.Session will be bound to---and only able to run operations in---the current default graph. If
you are using multiple graphs in your program (see Programming with multiple graphs for more details), you can specify
an explicit tf.Graph when you construct the session.
config. This argument allows you to specify a tf.ConfigProto that controls the behavior of the session. For example, some
of the configuration options include:
allow_soft_placement. Set this to True to enable a "soft" device placement algorithm, which ignores tf.device
annotations that attempt to place CPU-only operations on a GPU device, and places them on the CPU instead.
cluster_def. When using distributed TensorFlow, this option allows you to specify what machines to use in the

computation, and provide a mapping between job names, task indices, and network addresses. See
tf.train.ClusterSpec.as_cluster_def for details.
graph_options.optimizer_options. Provides control over the optimizations that TensorFlow performs on your graph
before executing it.
gpu_options.allow_growth. Set this to True to change the GPU memory allocator so that it gradually increases the
amount of memory allocated, rather than allocating most of the memory at startup.
Using tf.Session.run to execute operations
The tf.Session.run method is the main mechanism for running a tf.Operation or evaluating a tf.Tensor. You can pass
one or more tf.Operation or tf.Tensor objects to tf.Session.run, and TensorFlow will execute the operations that are
needed to compute the result.
tf.Session.run requires you to specify a list of fetches, which determine the return values, and may be a tf.Operation, a
tf.Tensor, or a tensor-like type such as tf.Variable. These fetches determine what subgraph of the overall tf.Graph must
be executed to produce the result: this is the subgraph that contains all operations named in the fetch list, plus all operations
whose outputs are used to compute the value of the fetches. For example, the following code fragment shows how different
arguments to tf.Session.run cause different subgraphs to be executed:
x = tf.constant([[37.0, -23.0], [1.0, 4.0]])
w = tf.Variable(tf.random_uniform([2, 2]))
y = tf.matmul(x, w)
output = tf.nn.softmax(y)
init_op = w.initializer
with tf.Session() as sess:
# Run the initializer on `w`.
sess.run(init_op)
# Evaluate `output`. `sess.run(output)` will return a NumPy array containing
# the result of the computation.
print(sess.run(output))
# Evaluate `y` and `output`. Note that `y` will only be computed once, and its
# result used both to return `y_val` and as an input to the `tf.nn.softmax()`
# op. Both `y_val` and `output_val` will be NumPy arrays.
y_val, output_val = sess.run([y, output])
tf.Session.run also optionally takes a dictionary of feeds, which is a mapping from tf.Tensor objects (typically
tf.placeholder tensors) to values (typically Python scalars, lists, or NumPy arrays) that will be substituted for those tensors
in the execution. For example:

# Define a placeholder that expects a vector of three floating-point values,
# and a computation that depends on it.
x = tf.placeholder(tf.float32, shape=[3])
y = tf.square(x)
with tf.Session() as sess:
# Feeding a value changes the result that is returned when you evaluate `y`.
print(sess.run(y, {x: [1.0, 2.0, 3.0]})) # => "[1.0, 4.0, 9.0]"
print(sess.run(y, {x: [0.0, 0.0, 5.0]})) # => "[0.0, 0.0, 25.0]"
# Raises `tf.errors.InvalidArgumentError`, because you must feed a value for
# a `tf.placeholder()` when evaluating a tensor that depends on it.
sess.run(y)
# Raises `ValueError`, because the shape of `37.0` does not match the shape
# of placeholder `x`.
sess.run(y, {x: 37.0})
tf.Session.run also accepts an optional options argument that enables you to specify options about the call, and an
optional run_metadata argument that enables you to collect metadata about the execution. For example, you can use these
options together to collect tracing information about the execution:
y = tf.matmul([[37.0, -23.0], [1.0, 4.0]], tf.random_uniform([2, 2]))
with tf.Session() as sess:
# Define options for the `sess.run()` call.
options = tf.RunOptions()
options.output_partition_graphs = True
options.trace_level = tf.RunOptions.FULL_TRACE
# Define a container for the returned metadata.
metadata = tf.RunMetadata()
sess.run(y, options=options, run_metadata=metadata)
# Print the subgraphs that executed on each device.
print(metadata.partition_graphs)
# Print the timings of each operation that executed.
print(metadata.step_stats)
Visualizing your graph
TensorFlow includes tools that can help you to understand the code in a graph. The graph visualizer is a component of
TensorBoard that renders the structure of your graph visually in a browser. The easiest way to create a visualization is to pass a
tf.Graph when creating the tf.summary.FileWriter:

# Build your graph.
x = tf.constant([[37.0, -23.0], [1.0, 4.0]])
w = tf.Variable(tf.random_uniform([2, 2]))
y = tf.matmul(x, w)
# ...
loss = ...
train_op = tf.train.AdagradOptimizer(0.01).minimize(loss)
with tf.Session() as sess:
# `sess.graph` provides access to the graph used in a `tf.Session`.
writer = tf.summary.FileWriter("/tmp/log/...", sess.graph)
# Perform your computation...
for i in range(1000):
sess.run(train_op)
# ...
writer.close()
Note: If you are using a tf.estimator.Estimator, the graph (and any summaries) will be logged automatically to the
model_dir that you specified when creating the estimator.
You can then open the log in tensorboard, navigate to the "Graph" tab, and see a high-level visualization of your graph's
structure. Note that a typical TensorFlow graph---especially training graphs with automatically computed gradients---has too
many nodes to visualize at once. The graph visualizer makes use of name scopes to group related operations into "super"
nodes. You can click on the orange "+" button on any of these super nodes to expand the subgraph inside.

Note: When training a model, a common way of organizing your code is to use one graph for training your model, and a
separate graph for evaluating or performing inference with a trained model. In many cases, the inference graph will be
different from the training graph: for example, techniques like dropout and batch normalization use different operations in
each case. Furthermore, by default utilities like tf.train.Saver use the names of tf.Variable objects (which have names
based on an underlying tf.Operation) to identify each variable in a saved checkpoint. When programming this way, you can
either use completely separate Python processes to build and execute the graphs, or you can use multiple graphs in the same
process. This section describes how to use multiple graphs in the same process.
As noted above, TensorFlow provides a "default graph" that is implicitly passed to all API functions in the same context. For
many applications, a single graph is sufficient. However, TensorFlow also provides methods for manipulating the default
graph, which can be useful in more advanced used cases. For example:
A tf.Graph defines the namespace for tf.Operation objects: each operation in a single graph must have a unique name.
TensorFlow will "uniquify" the names of operations by appending "_1", "_2", and so on to their names if the requested
name is already taken. Using multiple explicitly created graphs gives you more control over what name is given to each
operation.
The default graph stores information about every tf.Operation and tf.Tensor that was ever added to it. If your program
creates a large number of unconnected subgraphs, it may be more efficient to use a different tf.Graph to build each
subgraph, so that unrelated state can be garbage collected.
You can install a different tf.Graph as the default graph, using the tf.Graph.as_default context manager:
g_1 = tf.Graph()
with g_1.as_default():
# Operations created in this scope will be added to `g_1`.
c = tf.constant("Node in g_1")
# Sessions created in this scope will run operations from `g_1`.
sess_1 = tf.Session()
g_2 = tf.Graph()
with g_2.as_default():
# Operations created in this scope will be added to `g_2`.
d = tf.constant("Node in g_2")
# Alternatively, you can pass a graph when constructing a `tf.Session`:
# `sess_2` will run operations from `g_2`.
sess_2 = tf.Session(graph=g_2)
assert c.graph is g_1
assert sess_1.graph is g_1
assert d.graph is g_2
assert sess_2.graph is g_2
To inspect the current default graph, call tf.get_default_graph, which returns a tf.Graph object:
Saving and Restoring
This document explains how to save and restore variables and models.
# Print all of the operations in the default graph.
g = tf.get_default_graph()
print(g.get_operations())

Saving and restoring variables
A TensorFlow variable provides the best way to represent shared, persistent state manipulated by your program. (See Variables
for details.) This section explains how to save and restore variables. Note that Estimators automatically saves and restores
variables (in the model_dir).
The tf.train.Saver class provides methods for saving and restoring models. The tf.train.Saverconstructor adds save
and restore ops to the graph for all, or a specified list, of the variables in the graph. The Saver object provides methods to run
these ops, specifying paths for the checkpoint files to write to or read from.
The saver will restore all variables already defined in your model. If you're loading a model without knowing how to build its
graph (for example, if you're writing a generic program to load models), then read the Overview of saving and restoring
models section later in this document.
TensorFlow saves variables in binary checkpoint files that, roughly speaking, map variable names to tensor values.
Saving variables
Create a Saver with tf.train.Saver() to manage all variables in the model. For example, the following snippet
demonstrates how to call the tf.train.Saver.save method to save variables to checkpoint files:
# Create some variables.
v1 = tf.get_variable("v1", shape=[3], initializer = tf.zeros_initializer)
v2 = tf.get_variable("v2", shape=[5], initializer = tf.zeros_initializer)
inc_v1 = v1.assign(v1+1)
dec_v2 = v2.assign(v2-1)
# Add an op to initialize the variables.
init_op = tf.global_variables_initializer()
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
# Later, launch the model, initialize the variables, do some work, and save the
# variables to disk.
with tf.Session() as sess:
sess.run(init_op)
# Do some work with the model.
inc_v1.op.run()
dec_v2.op.run()
# Save the variables to disk.
save_path = saver.save(sess, "/tmp/model.ckpt")
print("Model saved in path: %s" % save_path)
Restoring variables
The tf.train.Saver object not only saves variables to checkpoint files, it also restores variables. Note that when you restore
variables you do not have to initialize them beforehand. For example, the following snippet demonstrates how to call the
tf.train.Saver.restore method to restore variables from the checkpoint files:

tf.reset_default_graph()
# Create some variables.
v1 = tf.get_variable("v1", shape=[3])
v2 = tf.get_variable("v2", shape=[5])
# Add ops to save and restore all the variables.
saver = tf.train.Saver()
# Later, launch the model, use the saver to restore variables from disk, and
# do some work with the model.
with tf.Session() as sess:
# Restore variables from disk.
saver.restore(sess, "/tmp/model.ckpt")
print("Model restored.")
# Check the values of the variables
print("v1 : %s" % v1.eval())
print("v2 : %s" % v2.eval())
Notes:
There is not a physical file called "/tmp/model.ckpt". It is the prefix of filenames created for the checkpoint. Users only
interact with the prefix instead of physical checkpoint files.
Choosing which variables to save and restore
If you do not pass any arguments to tf.train.Saver(), the saver handles all variables in the graph. Each variable is saved
under the name that was passed when the variable was created.
It is sometimes useful to explicitly specify names for variables in the checkpoint files. For example, you may have trained a
model with a variable named "weights" whose value you want to restore into a variable named"params".
It is also sometimes useful to only save or restore a subset of the variables used by a model. For example, you may have trained
a neural net with five layers, and you now want to train a new model with six layers that reuses the existing weights of the five
trained layers. You can use the saver to restore the weights of just the first five layers.
You can easily specify the names and variables to save or load by passing to the tf.train.Saver()constructor either of the
following:
A list of variables (which will be stored under their own names).
A Python dictionary in which keys are the names to use and the values are the variables to manage.
Continuing from the save/restore examples shown earlier:

tf.reset_default_graph()
# Create some variables.
v1 = tf.get_variable("v1", [3], initializer = tf.zeros_initializer)
v2 = tf.get_variable("v2", [5], initializer = tf.zeros_initializer)
# Add ops to save and restore only `v2` using the name "v2"
saver = tf.train.Saver({"v2": v2})
# Use the saver object normally after that.
with tf.Session() as sess:
# Initialize v1 since the saver will not.
v1.initializer.run()
saver.restore(sess, "/tmp/model.ckpt")
print("v1 : %s" % v1.eval())
print("v2 : %s" % v2.eval())
Notes:
You can create as many Saver objects as you want if you need to save and restore different subsets of the model variables.
The same variable can be listed in multiple saver objects; its value is only changed when the Saver.restore() method is
run.
If you only restore a subset of the model variables at the start of a session, you have to run an initialize op for the other
variables. See tf.variables_initializer for more information.
To inspect the variables in a checkpoint, you can use the inspect_checkpoint library, particularly the
print_tensors_in_checkpoint_file function.
By default, Saver uses the value of the tf.Variable.name property for each variable. However, when you create a Saver
object, you may optionally choose names for the variables in the checkpoint files.
Inspect variables in a checkpoint
We can quickly inspect variables in a checkpoint with the inspect_checkpoint library.
Continuing from the save/restore examples shown earlier:

# import the inspect_checkpoint library
from tensorflow.python.tools import inspect_checkpoint as chkp
# print all tensors in checkpoint file
chkp.print_tensors_in_checkpoint_file("/tmp/model.ckpt", tensor_name='', all_tensors=True)
# tensor_name: v1
# [ 1. 1. 1.]
# tensor_name: v2
# [-1. -1. -1. -1. -1.]
# print only tensor v1 in checkpoint file
chkp.print_tensors_in_checkpoint_file("/tmp/model.ckpt", tensor_name='v1', all_tensors=False)
# tensor_name: v1
# [ 1. 1. 1.]
# print only tensor v2 in checkpoint file
chkp.print_tensors_in_checkpoint_file("/tmp/model.ckpt", tensor_name='v2', all_tensors=False)
# tensor_name: v2
# [-1. -1. -1. -1. -1.]
Overview of saving and restoring models
When you want to save and load variables, the graph, and the graph's metadata--basically, when you want to save or restore
your model--we recommend using SavedModel. SavedModel is a language-neutral, recoverable, hermetic serialization format.
SavedModel enables higher-level systems and tools to produce, consume, and transform TensorFlow models. TensorFlow
provides several mechanisms for interacting with SavedModel, including tf.saved_model APIs, Estimator APIs and a CLI.
APIs to build and load a SavedModel
This section focuses on the APIs for building and loading a SavedModel, particularly when using lower-level TensorFlow APIs.
Building a SavedModel
We provide a Python implementation of the SavedModel builder. The SavedModelBuilder class provides functionality to
save multiple MetaGraphDefs. A MetaGraph is a dataflow graph, plus its associated variables, assets, and signatures. A
MetaGraphDef is the protocol buffer representation of a MetaGraph. A signature is the set of inputs to and outputs from a
graph.
If assets need to be saved and written or copied to disk, they can be provided when the first MetaGraphDef is added. If
multiple MetaGraphDefs are associated with an asset of the same name, only the first version is retained.
Each MetaGraphDef added to the SavedModel must be annotated with user-specified tags. The tags provide a means to
identify the specific MetaGraphDef to load and restore, along with the shared set of variables and assets. These tags typically
annotate a MetaGraphDef with its functionality (for example, serving or training), and optionally with hardware-specific
aspects (for example, GPU).
For example, the following code suggests a typical way to use SavedModelBuilder to build a SavedModel:

export_dir = ...
...
builder = tf.saved_model.builder.SavedModelBuilder(export_dir)
with tf.Session(graph=tf.Graph()) as sess:
...
builder.add_meta_graph_and_variables(sess,
[tag_constants.TRAINING],
signature_def_map=foo_signatures,
assets_collection=foo_assets)
...
# Add a second MetaGraphDef for inference.
with tf.Session(graph=tf.Graph()) as sess:
...
builder.add_meta_graph([tag_constants.SERVING])
...
builder.save()
Loading a SavedModel in Python
The Python version of the SavedModel loader provides load and restore capability for a SavedModel. The loadoperation
requires the following information:
The session in which to restore the graph definition and variables.
The tags used to identify the MetaGraphDef to load.
The location (directory) of the SavedModel.
Upon a load, the subset of variables, assets, and signatures supplied as part of the specific MetaGraphDef will be restored into
the supplied session.
export_dir = ...
...
with tf.Session(graph=tf.Graph()) as sess:
tf.saved_model.loader.load(sess, [tag_constants.TRAINING], export_dir)
...
Loading a Savedmodel in C++
The C++ version of the SavedModel loader provides an API to load a SavedModel from a path, while
allowingSessionOptions and RunOptions. You have to specify the tags associated with the graph to be loaded. The loaded
version of SavedModel is referred to as SavedModelBundle and contains the MetaGraphDef and the session within which it is
loaded.
const string export_dir = ...
SavedModelBundle bundle;
...
LoadSavedModel(session_options, run_options, export_dir, {kSavedModelTagTrain},
&bundle);

Standard constants
SavedModel offers the flexibility to build and load TensorFlow graphs for a variety of use-cases. For the most common use-
cases, SavedModel's APIs provide a set of constants in Python and C++ that are easy to reuse and share across tools
consistently.
Standard MetaGraphDef tags
You may use sets of tags to uniquely identify a MetaGraphDef saved in a SavedModel. A subset of commonly used tags is
specified in:
Python
C++
Standard SignatureDef constants
A SignatureDef is a protocol buffer that defines the signature of a computation supported by a graph. Commonly used input
keys, output keys, and method names are defined in:
Python
C++
Using SavedModel with Estimators
After training an Estimator model, you may want to create a service from that model that takes requests and returns a result.
You can run such a service locally on your machine or deploy it scalably in the cloud.
To prepare a trained Estimator for serving, you must export it in the standard SavedModel format. This section explains how
to:
Specify the output nodes and the corresponding APIs that can be served (Classify, Regress, or Predict).
Export your model to the SavedModel format.
Serve the model from a local server and request predictions.
Preparing serving inputs
During training, an input_fn() ingests data and prepares it for use by the model. At serving time, similarly,
aserving_input_receiver_fn() accepts inference requests and prepares them for the model. This function has the following
purposes:
To add placeholders to the graph that the serving system will feed with inference requests.
To add any additional ops needed to convert data from the input format into the feature Tensors expected by the model.
The function returns a tf.estimator.export.ServingInputReceiver object, which packages the placeholders and the
resulting feature Tensors together.
A typical pattern is that inference requests arrive in the form of serialized tf.Examples, so the
serving_input_receiver_fn() creates a single string placeholder to receive them. The serving_input_receiver_fn() is
then also responsible for parsing the tf.Examples by adding a tf.parse_example op to the graph.
When writing such a serving_input_receiver_fn(), you must pass a parsing specification to tf.parse_example to tell the
parser what feature names to expect and how to map them to Tensors. A parsing specification takes the form of a dict from
feature names to tf.FixedLenFeature, tf.VarLenFeature, and tf.SparseFeature. Note this parsing specification should
not include any label or weight columns, since those will not be available at serving time—in contrast to a parsing specification

used in the input_fn() at training time.
In combination, then:
feature_spec = {'foo': tf.FixedLenFeature(...),
'bar': tf.VarLenFeature(...)}
def serving_input_receiver_fn():
"""An input receiver that expects a serialized tf.Example."""
serialized_tf_example = tf.placeholder(dtype=tf.string,
shape=[default_batch_size],
name='input_example_tensor')
receiver_tensors = {'examples': serialized_tf_example}
features = tf.parse_example(serialized_tf_example, feature_spec)
return tf.estimator.export.ServingInputReceiver(features, receiver_tensors)
The tf.estimator.export.build_parsing_serving_input_receiver_fn utility function provides that input receiver for
the common case.
Note: when training a model to be served using the Predict API with a local server, the parsing step is not needed because
the model will receive raw feature data.
Even if you require no parsing or other input processing—that is, if the serving system will feed feature Tensors directly—you
must still provide a serving_input_receiver_fn() that creates placeholders for the featureTensors and passes them
through. The tf.estimator.export.build_raw_serving_input_receiver_fnutility provides for this.
If these utilities do not meet your needs, you are free to write your own serving_input_receiver_fn(). One case where this
may be needed is if your training input_fn() incorporates some preprocessing logic that must be recapitulated at serving
time. To reduce the risk of training-serving skew, we recommend encapsulating such processing in a function which is then
called from both input_fn() and serving_input_receiver_fn().
Note that the serving_input_receiver_fn() also determines the input portion of the signature. That is, when writing a
serving_input_receiver_fn(), you must tell the parser what signatures to expect and how to map them to your model's
expected inputs. By contrast, the output portion of the signature is determined by the model.
Performing the export
To export your trained Estimator, call tf.estimator.Estimator.export_savedmodel with the export base path and the
serving_input_receiver_fn.
estimator.export_savedmodel(export_dir_base, serving_input_receiver_fn)
This method builds a new graph by first calling the serving_input_receiver_fn() to obtain feature Tensors, and then
calling this Estimator's model_fn() to generate the model graph based on those features. It starts a fresh Session, and, by
default, restores the most recent checkpoint into it. (A different checkpoint may be passed, if needed.) Finally it creates a time-
stamped export directory below the givenexport_dir_base (i.e., export_dir_base/<timestamp>), and writes a
SavedModel into it containing a single MetaGraphDef saved from this Session.
Note: It is your responsibility to garbage-collect old exports. Otherwise, successive exports will accumulate under
export_dir_base.
Specifying the outputs of a custom model

When writing a custom model_fn, you must populate the export_outputs element of the tf.estimator.EstimatorSpec
return value. This is a dict of {name: output} describing the output signatures to be exported and used during serving.
In the usual case of making a single prediction, this dict contains one element, and the name is immaterial. In a multi-headed
model, each head is represented by an entry in this dict. In this case the name is a string of your choice that can be used to
request a specific head at serving time.
Each output value must be an ExportOutput object such astf.estimator.export.ClassificationOutput,
tf.estimator.export.RegressionOutput, ortf.estimator.export.PredictOutput.
These output types map straightforwardly to the TensorFlow Serving APIs, and so determine which request types will be
honored.
Note: In the multi-headed case, a SignatureDef will be generated for each element of the export_outputs dict returned from
the model_fn, named using the same keys. These SignatureDefs differ only in their outputs, as provided by the
corresponding ExportOutput entry. The inputs are always those provided by the serving_input_receiver_fn. An
inference request may specify the head by name. One head must be named using
signature_constants.DEFAULT_SERVING_SIGNATURE_DEF_KEY indicating which SignatureDef will be served when an
inference request does not specify one.
Serving the exported model locally
For local deployment, you can serve your model using TensorFlow Serving, an open-source project that loads a SavedModel
and exposes it as a gRPC service.
First, install TensorFlow Serving.
Then build and run the local model server, substituting $export_dir_base with the path to the SavedModel you exported
above:
bazel build //tensorflow_serving/model_servers:tensorflow_model_server
bazel-bin/tensorflow_serving/model_servers/tensorflow_model_server --port=9000 --
model_base_path=$export_dir_base
Now you have a server listening for inference requests via gRPC on port 9000!
Requesting predictions from a local server
The server responds to gRPC requests according to the PredictionService gRPC API service definition. (The nested protocol
buffers are defined in various neighboring files).
From the API service definition, the gRPC framework generates client libraries in various languages providing remote access to
the API. In a project using the Bazel build tool, these libraries are built automatically and provided via dependencies like these
(using Python for example):
deps = [
"//tensorflow_serving/apis:classification_proto_py_pb2",
"//tensorflow_serving/apis:regression_proto_py_pb2",
"//tensorflow_serving/apis:predict_proto_py_pb2",
"//tensorflow_serving/apis:prediction_service_proto_py_pb2"
]
Python client code can then import the libraries thus:

from tensorflow_serving.apis import classification_pb2
from tensorflow_serving.apis import regression_pb2
from tensorflow_serving.apis import predict_pb2
from tensorflow_serving.apis import prediction_service_pb2
Note: prediction_service_pb2 defines the service as a whole and so is always required. However a typical client will
need only one of classification_pb2, regression_pb2, and predict_pb2, depending on the type of requests being
made.
Sending a gRPC request is then accomplished by assembling a protocol buffer containing the request data and passing it to the
service stub. Note how the request protocol buffer is created empty and then populated via thegenerated protocol buffer API.
from grpc.beta import implementations
channel = implementations.insecure_channel(host, int(port))
stub = prediction_service_pb2.beta_create_PredictionService_stub(channel)
request = classification_pb2.ClassificationRequest()
example = request.input.example_list.examples.add()
example.features.feature['x'].float_list.value.extend(image[0].astype(float))
result = stub.Classify(request, 10.0) # 10 secs timeout
The returned result in this example is a ClassificationResponse protocol buffer.
This is a skeletal example; please see the Tensorflow Serving documentation and examples for more details.
Note: ClassificationRequest and RegressionRequest contain a tensorflow.serving.Input protocol buffer, which
in turn contains a list of tensorflow.Example protocol buffers. PredictRequest, by contrast, contains a mapping from
feature names to values encoded via TensorProto. Correspondingly: When using the Classify and Regress APIs,
TensorFlow Serving feeds serialized tf.Examples to the graph, so yourserving_input_receiver_fn() should include
a tf.parse_example() Op. When using the generic PredictAPI, however, TensorFlow Serving feeds raw feature data to
the graph, so a pass through serving_input_receiver_fn() should be used.
CLI to inspect and execute SavedModel
You can use the SavedModel Command Line Interface (CLI) to inspect and execute a SavedModel. For example, you can use
the CLI to inspect the model's SignatureDefs. The CLI enables you to quickly confirm that the input Tensor dtype and shape
match the model. Moreover, if you want to test your model, you can use the CLI to do a sanity check by passing in sample
inputs in various formats (for example, Python expressions) and then fetching the output.
Installing the SavedModel CLI
Broadly speaking, you can install TensorFlow in either of the following two ways:
By installing a pre-built TensorFlow binary.
By building TensorFlow from source code.
If you installed TensorFlow through a pre-built TensorFlow binary, then the SavedModel CLI is already installed on your
system at pathname bin\saved_model_cli.
If you built TensorFlow from source code, you must run the following additional command to build saved_model_cli:
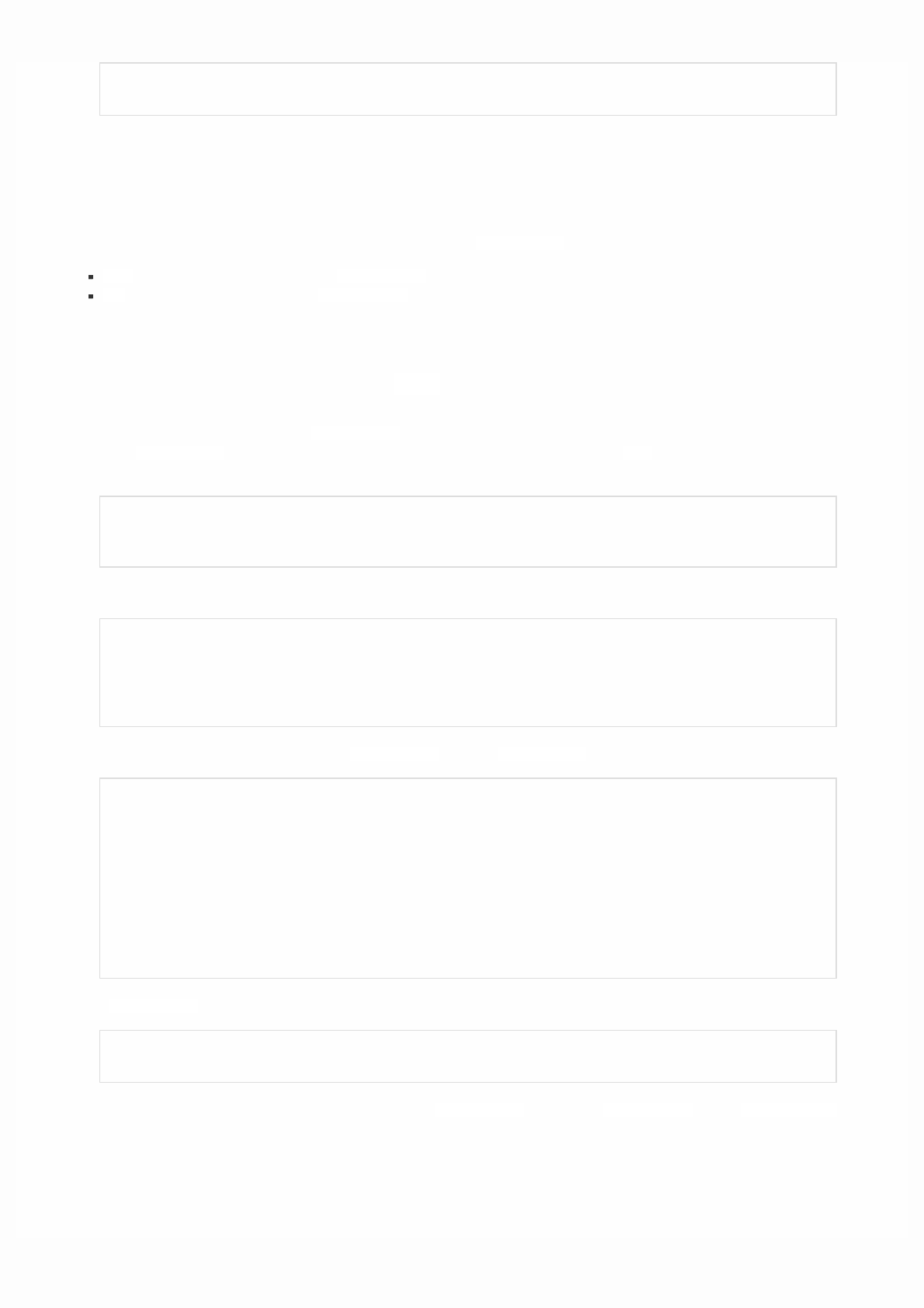
$ bazel build tensorflow/python/tools:saved_model_cli
Overview of commands
The SavedModel CLI supports the following two commands on a MetaGraphDef in a SavedModel:
show, which shows a computation on a MetaGraphDef in a SavedModel.
run, which runs a computation on a MetaGraphDef.
show command
A SavedModel contains one or more MetaGraphDefs, identified by their tag-sets. To serve a model, you might wonder what
kind of SignatureDefs are in each model, and what are their inputs and outputs. The showcommand let you examine the
contents of the SavedModel in hierarchical order. Here's the syntax:
usage: saved_model_cli show [-h] --dir DIR [--all]
[--tag_set TAG_SET] [--signature_def SIGNATURE_DEF_KEY]
For example, the following command shows all available MetaGraphDef tag-sets in the SavedModel:
$ saved_model_cli show --dir /tmp/saved_model_dir
The given SavedModel contains the following tag-sets:
serve
serve, gpu
The following command shows all available SignatureDef keys in a MetaGraphDef:
$ saved_model_cli show --dir /tmp/saved_model_dir --tag_set serve
The given SavedModel `MetaGraphDef` contains `SignatureDefs` with the
following keys:
SignatureDef key: "classify_x2_to_y3"
SignatureDef key: "classify_x_to_y"
SignatureDef key: "regress_x2_to_y3"
SignatureDef key: "regress_x_to_y"
SignatureDef key: "regress_x_to_y2"
SignatureDef key: "serving_default"
If a MetaGraphDef has multiple tags in the tag-set, you must specify all tags, each tag separated by a comma. For example:
$ saved_model_cli show --dir /tmp/saved_model_dir --tag_set serve,gpu
To show all inputs and outputs TensorInfo for a specific SignatureDef, pass in the SignatureDef key to signature_def
option. This is very useful when you want to know the tensor key value, dtype and shape of the input tensors for executing the
computation graph later. For example:

$ saved_model_cli show --dir \
/tmp/saved_model_dir --tag_set serve --signature_def serving_default
The given SavedModel SignatureDef contains the following input(s):
inputs['x'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: x:0
The given SavedModel SignatureDef contains the following output(s):
outputs['y'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: y:0
Method name is: tensorflow/serving/predict
To show all available information in the SavedModel, use the --all option. For example:
$ saved_model_cli show --dir /tmp/saved_model_dir --all
MetaGraphDef with tag-set: 'serve' contains the following SignatureDefs:
signature_def['classify_x2_to_y3']:
The given SavedModel SignatureDef contains the following input(s):
inputs['inputs'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: x2:0
The given SavedModel SignatureDef contains the following output(s):
outputs['scores'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: y3:0
Method name is: tensorflow/serving/classify
...
signature_def['serving_default']:
The given SavedModel SignatureDef contains the following input(s):
inputs['x'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: x:0
The given SavedModel SignatureDef contains the following output(s):
outputs['y'] tensor_info:
dtype: DT_FLOAT
shape: (-1, 1)
name: y:0
Method name is: tensorflow/serving/predict
run command
Invoke the run command to run a graph computation, passing inputs and then displaying (and optionally saving) the outputs.
Here's the syntax:

usage: saved_model_cli run [-h] --dir DIR --tag_set TAG_SET --signature_def
SIGNATURE_DEF_KEY [--inputs INPUTS]
[--input_exprs INPUT_EXPRS] [--outdir OUTDIR]
[--overwrite] [--tf_debug]
The run command provides the following two ways to pass inputs to the model:
--inputs option enables you to pass numpy ndarray in files.
--input_exprs option enables you to pass Python expressions.
--inputs
To pass input data in files, specify the --inputs option, which takes the following general format:
--inputs <INPUTS>
where INPUTS is either of the following formats:
<input_key>=<filename>
<input_key>=<filename>[<variable_name>]
You may pass multiple INPUTS. If you do pass multiple inputs, use a semicolon to separate each of the INPUTS.
saved_model_cli uses numpy.load to load the filename. The filename may be in any of the following formats:
.npy
.npz
pickle format
A .npy file always contains a numpy ndarray. Therefore, when loading from a .npy file, the content will be directly assigned to
the specified input tensor. If you specify a variable_name with that .npy file, thevariable_name will be ignored and a warning will
be issued.
When loading from a .npz (zip) file, you may optionally specify a variable_name to identify the variable within the zip file to
load for the input tensor key. If you don't specify a variable_name, the SavedModel CLI will check that only one file is included
in the zip file and load it for the specified input tensor key.
When loading from a pickle file, if no variable_name is specified in the square brackets, whatever that is inside the pickle file
will be passed to the specified input tensor key. Otherwise, the SavedModel CLI will assume a dictionary is stored in the pickle
file and the value corresponding to the variable_name will be used.
--inputs_exprs
To pass inputs through Python expressions, specify the --input_exprs option. This can be useful for when you don't have
data files lying around, but still want to sanity check the model with some simple inputs that match the dtype and shape of the
model's SignatureDefs. For example:
`input_key=[[1], [2], [3]]`
In addition to Python expressions, you may also pass numpy functions. For example:
input_key=np.ones((32, 32, 3))
(Note that the numpy module is already available to you as np.)

Save Output
By default, the SavedModel CLI writes output to stdout. If a directory is passed to --outdir option, the outputs will be saved
as npy files named after output tensor keys under the given directory.
Use --overwrite to overwrite existing output files.
TensorFlow Debugger (tfdbg) Integration
If --tf_debug option is set, the SavedModel CLI will use the TensorFlow Debugger (tfdbg) to watch the intermediate Tensors
and runtime graphs or subgraphs while running the SavedModel.
Full examples of run
Given:
Your model simply adds x1 and x2 to get output y.
All tensors in the model have shape (-1, 1).
You have two npy files:
/tmp/my_data1.npy, which contains a numpy ndarray [[1], [2], [3]].
/tmp/my_data2.npy, which contains another numpy ndarray [[0.5], [0.5], [0.5]].
To run these two npy files through the model to get output y, issue the following command:
$ saved_model_cli run --dir /tmp/saved_model_dir --tag_set serve \
--signature_def x1_x2_to_y --inputs x1=/tmp/my_data1.npy;x2=/tmp/my_data2.npy \
--outdir /tmp/out
Result for output key y:
[[ 1.5]
[ 2.5]
[ 3.5]]
Let's change the preceding example slightly. This time, instead of two .npy files, you now have an .npz file and a pickle file.
Furthermore, you want to overwrite any existing output file. Here's the command:
$ saved_model_cli run --dir /tmp/saved_model_dir --tag_set serve \
--signature_def x1_x2_to_y \
--inputs x1=/tmp/my_data1.npz[x];x2=/tmp/my_data2.pkl --outdir /tmp/out \
--overwrite
Result for output key y:
[[ 1.5]
[ 2.5]
[ 3.5]]
You may specify python expression instead of an input file. For example, the following command replaces input x2 with a
Python expression:
$ saved_model_cli run --dir /tmp/saved_model_dir --tag_set serve \
--signature_def x1_x2_to_y --inputs x1=/tmp/my_data1.npz[x] \
--input_exprs 'x2=np.ones((3,1))'
Result for output key y:
[[ 2]
[ 3]
[ 4]]
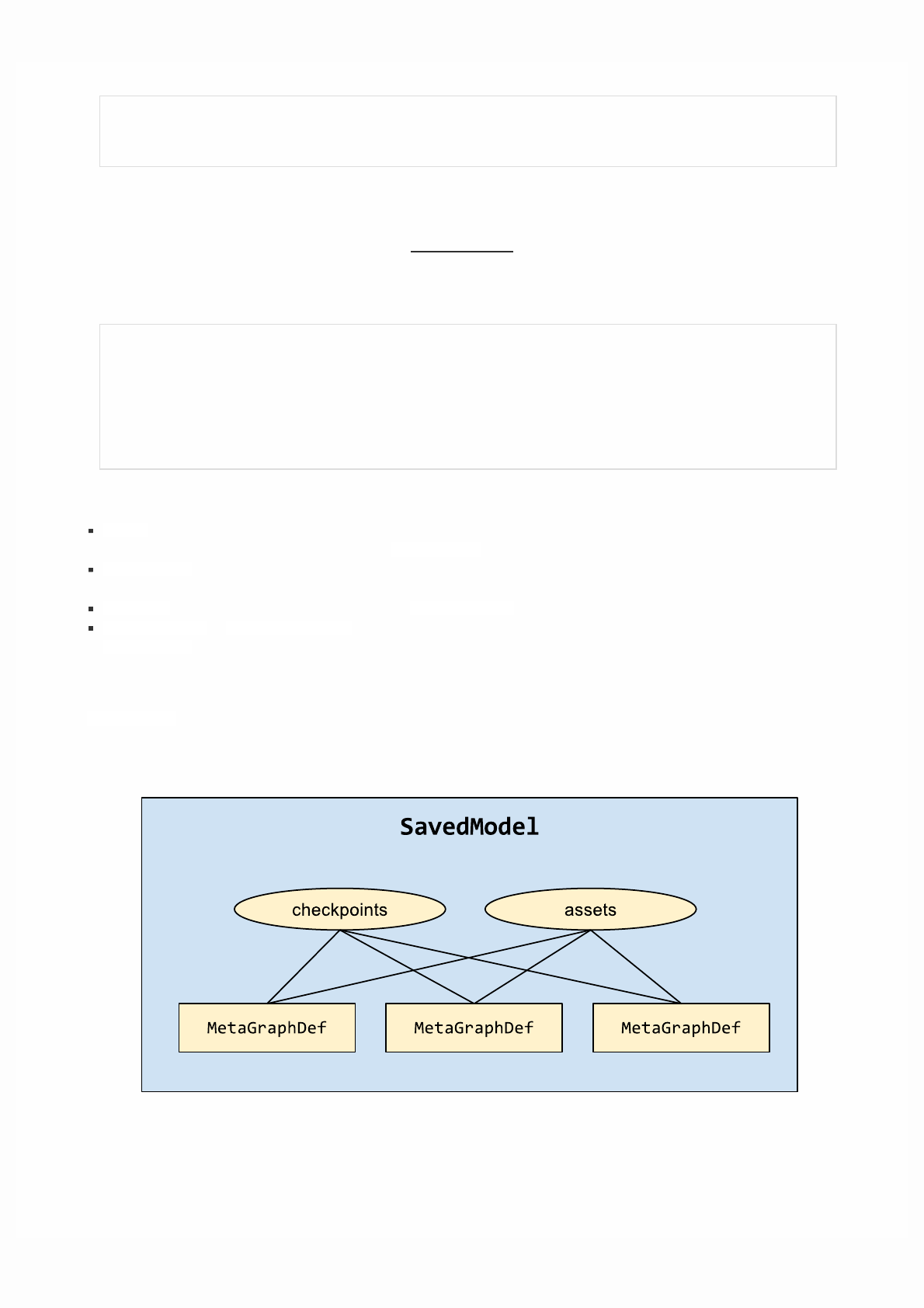
To run the model with the TensorFlow Debugger on, issue the following command:
$ saved_model_cli run --dir /tmp/saved_model_dir --tag_set serve \
--signature_def serving_default --inputs x=/tmp/data.npz[x] --tf_debug
Structure of a SavedModel directory
When you save a model in SavedModel format, TensorFlow creates a SavedModel directory consisting of the following
subdirectories and files:
assets/
assets.extra/
variables/
variables.data-?????-of-?????
variables.index
saved_model.pb|saved_model.pbtxt
where:
assets is a subfolder containing auxiliary (external) files, such as vocabularies. Assets are copied to the SavedModel
location and can be read when loading a specific MetaGraphDef.
assets.extra is a subfolder where higher-level libraries and users can add their own assets that co-exist with the model,
but are not loaded by the graph. This subfolder is not managed by the SavedModel libraries.
variables is a subfolder that includes output from tf.train.Saver.
saved_model.pb or saved_model.pbtxt is the SavedModel protocol buffer. It includes the graph definitions as
MetaGraphDef protocol buffers.
A single SavedModel can represent multiple graphs. In this case, all the graphs in the SavedModel share a singleset of
checkpoints (variables) and assets. For example, the following diagram shows one SavedModel containing three
MetaGraphDefs, all three of which share the same set of checkpoints and assets:
Each graph is associated with a specific set of tags, which enables identification during a load or restore operation.

Using GPUs
Supported devices
On a typical system, there are multiple computing devices. In TensorFlow, the supported device types are CPUand GPU. They
are represented as strings. For example:
"/cpu:0": The CPU of your machine.
"/device:GPU:0": The GPU of your machine, if you have one.
"/device:GPU:1": The second GPU of your machine, etc.
If a TensorFlow operation has both CPU and GPU implementations, the GPU devices will be given priority when the operation
is assigned to a device. For example, matmul has both CPU and GPU kernels. On a system with devices cpu:0 and gpu:0,
gpu:0 will be selected to run matmul.
Logging Device placement
To find out which devices your operations and tensors are assigned to, create the session with log_device_placement
configuration option set to True.
# Creates a graph.
a = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3], name='a')
b = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2], name='b')
c = tf.matmul(a, b)
# Creates a session with log_device_placement set to True.
sess = tf.Session(config=tf.ConfigProto(log_device_placement=True))
# Runs the op.
print(sess.run(c))
You should see the following output:
Device mapping:
/job:localhost/replica:0/task:0/device:GPU:0 -> device: 0, name: Tesla K40c, pci bus
id: 0000:05:00.0
b: /job:localhost/replica:0/task:0/device:GPU:0
a: /job:localhost/replica:0/task:0/device:GPU:0
MatMul: /job:localhost/replica:0/task:0/device:GPU:0
[[ 22. 28.]
[ 49. 64.]]
Manual device placement
If you would like a particular operation to run on a device of your choice instead of what's automatically selected for you, you
can use with tf.device to create a device context such that all the operations within that context will have the same device
assignment.

# Creates a graph.
with tf.device('/cpu:0'):
a = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3], name='a')
b = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2], name='b')
c = tf.matmul(a, b)
# Creates a session with log_device_placement set to True.
sess = tf.Session(config=tf.ConfigProto(log_device_placement=True))
# Runs the op.
print(sess.run(c))
You will see that now a and b are assigned to cpu:0. Since a device was not explicitly specified for the MatMul operation, the
TensorFlow runtime will choose one based on the operation and available devices (gpu:0 in this example) and automatically
copy tensors between devices if required.
Device mapping:
/job:localhost/replica:0/task:0/device:GPU:0 -> device: 0, name: Tesla K40c, pci bus
id: 0000:05:00.0
b: /job:localhost/replica:0/task:0/cpu:0
a: /job:localhost/replica:0/task:0/cpu:0
MatMul: /job:localhost/replica:0/task:0/device:GPU:0
[[ 22. 28.]
[ 49. 64.]]
Allowing GPU memory growth
By default, TensorFlow maps nearly all of the GPU memory of all GPUs (subject to CUDA_VISIBLE_DEVICES) visible to the
process. This is done to more efficiently use the relatively precious GPU memory resources on the devices by reducing memory
fragmentation.
In some cases it is desirable for the process to only allocate a subset of the available memory, or to only grow the memory
usage as is needed by the process. TensorFlow provides two Config options on the Session to control this.
The first is the allow_growth option, which attempts to allocate only as much GPU memory based on runtime allocations: it
starts out allocating very little memory, and as Sessions get run and more GPU memory is needed, we extend the GPU memory
region needed by the TensorFlow process. Note that we do not release memory, since that can lead to even worse memory
fragmentation. To turn this option on, set the option in the ConfigProto by:
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
session = tf.Session(config=config, ...)
The second method is the per_process_gpu_memory_fraction option, which determines the fraction of the overall amount
of memory that each visible GPU should be allocated. For example, you can tell TensorFlow to only allocate 40% of the total
memory of each GPU by:
config = tf.ConfigProto()
config.gpu_options.per_process_gpu_memory_fraction = 0.4
session = tf.Session(config=config, ...)
This is useful if you want to truly bound the amount of GPU memory available to the TensorFlow process.

Using a single GPU on a multi-GPU system
If you have more than one GPU in your system, the GPU with the lowest ID will be selected by default. If you would like to
run on a different GPU, you will need to specify the preference explicitly:
# Creates a graph.
with tf.device('/device:GPU:2'):
a = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3], name='a')
b = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2], name='b')
c = tf.matmul(a, b)
# Creates a session with log_device_placement set to True.
sess = tf.Session(config=tf.ConfigProto(log_device_placement=True))
# Runs the op.
print(sess.run(c))
If the device you have specified does not exist, you will get InvalidArgumentError:
InvalidArgumentError: Invalid argument: Cannot assign a device to node 'b':
Could not satisfy explicit device specification '/device:GPU:2'
[[Node: b = Const[dtype=DT_FLOAT, value=Tensor<type: float shape: [3,2]
values: 1 2 3...>, _device="/device:GPU:2"]()]]
If you would like TensorFlow to automatically choose an existing and supported device to run the operations in case the
specified one doesn't exist, you can set allow_soft_placement to True in the configuration option when creating the session.
# Creates a graph.
with tf.device('/device:GPU:2'):
a = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3], name='a')
b = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2], name='b')
c = tf.matmul(a, b)
# Creates a session with allow_soft_placement and log_device_placement set
# to True.
sess = tf.Session(config=tf.ConfigProto(
allow_soft_placement=True, log_device_placement=True))
# Runs the op.
print(sess.run(c))
Using multiple GPUs
If you would like to run TensorFlow on multiple GPUs, you can construct your model in a multi-tower fashion where each
tower is assigned to a different GPU. For example:

# Creates a graph.
c = []
for d in ['/device:GPU:2', '/device:GPU:3']:
with tf.device(d):
a = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[2, 3])
b = tf.constant([1.0, 2.0, 3.0, 4.0, 5.0, 6.0], shape=[3, 2])
c.append(tf.matmul(a, b))
with tf.device('/cpu:0'):
sum = tf.add_n(c)
# Creates a session with log_device_placement set to True.
sess = tf.Session(config=tf.ConfigProto(log_device_placement=True))
# Runs the op.
print(sess.run(sum))
You will see the following output.
Device mapping:
/job:localhost/replica:0/task:0/device:GPU:0 -> device: 0, name: Tesla K20m, pci bus
id: 0000:02:00.0
/job:localhost/replica:0/task:0/device:GPU:1 -> device: 1, name: Tesla K20m, pci bus
id: 0000:03:00.0
/job:localhost/replica:0/task:0/device:GPU:2 -> device: 2, name: Tesla K20m, pci bus
id: 0000:83:00.0
/job:localhost/replica:0/task:0/device:GPU:3 -> device: 3, name: Tesla K20m, pci bus
id: 0000:84:00.0
Const_3: /job:localhost/replica:0/task:0/device:GPU:3
Const_2: /job:localhost/replica:0/task:0/device:GPU:3
MatMul_1: /job:localhost/replica:0/task:0/device:GPU:3
Const_1: /job:localhost/replica:0/task:0/device:GPU:2
Const: /job:localhost/replica:0/task:0/device:GPU:2
MatMul: /job:localhost/replica:0/task:0/device:GPU:2
AddN: /job:localhost/replica:0/task:0/cpu:0
[[ 44. 56.]
[ 98. 128.]]
The cifar10 tutorial is a good example demonstrating how to do training with multiple GPUs.
Embeddings
This document introduces the concept of embeddings, gives a simple example of how to train an embedding in TensorFlow,
and explains how to view embeddings with the TensorBoard Embedding Projector (live example). The first two parts target
newcomers to machine learning or TensorFlow, and the Embedding Projector how-to is for users at all levels.
An embedding is a mapping from discrete objects, such as words, to vectors of real numbers. For example, a 300-dimensional
embedding for English words could include:
blue: (0.01359, 0.00075997, 0.24608, ..., -0.2524, 1.0048, 0.06259)
blues: (0.01396, 0.11887, -0.48963, ..., 0.033483, -0.10007, 0.1158)
orange: (-0.24776, -0.12359, 0.20986, ..., 0.079717, 0.23865, -0.014213)
oranges: (-0.35609, 0.21854, 0.080944, ..., -0.35413, 0.38511, -0.070976)
The individual dimensions in these vectors typically have no inherent meaning. Instead, it's the overall patterns of location and
distance between vectors that machine learning takes advantage of.

Embeddings are important for input to machine learning. Classifiers, and neural networks more generally, work on vectors of
real numbers. They train best on dense vectors, where all values contribute to define an object. However, many important
inputs to machine learning, such as words of text, do not have a natural vector representation. Embedding functions are the
standard and effective way to transform such discrete input objects into useful continuous vectors.
Embeddings are also valuable as outputs of machine learning. Because embeddings map objects to vectors, applications can
use similarity in vector space (for instance, Euclidean distance or the angle between vectors) as a robust and flexible measure of
object similarity. One common use is to find nearest neighbors. Using the same word embeddings as above, for instance, here
are the three nearest neighbors for each word and the corresponding angles:
blue: (red, 47.6°), (yellow, 51.9°), (purple, 52.4°)
blues: (jazz, 53.3°), (folk, 59.1°), (bluegrass, 60.6°)
orange: (yellow, 53.5°), (colored, 58.0°), (bright, 59.9°)
oranges: (apples, 45.3°), (lemons, 48.3°), (mangoes, 50.4°)
This would tell an application that apples and oranges are in some way more similar (45.3° apart) than lemons and oranges
(48.3° apart).
Embeddings in TensorFlow
To create word embeddings in TensorFlow, we first split the text into words and then assign an integer to every word in the
vocabulary. Let us assume that this has already been done, and that word_ids is a vector of these integers. For example, the
sentence “I have a cat.” could be split into [“I”, “have”, “a”, “cat”, “.”] and then the corresponding word_ids tensor
would have shape [5] and consist of 5 integers. To map these word ids to vectors, we need to create the embedding variable
and use the tf.nn.embedding_lookup function as follows:
word_embeddings = tf.get_variable(“word_embeddings”,
[vocabulary_size, embedding_size])
embedded_word_ids = tf.nn.embedding_lookup(word_embeddings, word_ids)
After this, the tensor embedded_word_ids will have shape [5, embedding_size] in our example and contain the
embeddings (dense vectors) for each of the 5 words. At the end of training, word_embeddings will contain the embeddings for
all words in the vocabulary.
Embeddings can be trained in many network types, and with various loss functions and data sets. For example, one could use
a recurrent neural network to predict the next word from the previous one given a large corpus of sentences, or one could train
two networks to do multi-lingual translation. These methods are described in the Vector Representations of Words tutorial.
Visualizing Embeddings
TensorBoard includes the Embedding Projector, a tool that lets you interactively visualize embeddings. This tool can read
embeddings from your model and render them in two or three dimensions.
The Embedding Projector has three panels:
Data panel on the top left, where you can choose the run, the embedding variable and data columns to color and label points
by.
Projections panel on the bottom left, where you can choose the type of projection.
Inspector panel on the right side, where you can search for particular points and see a list of nearest neighbors.
Projections
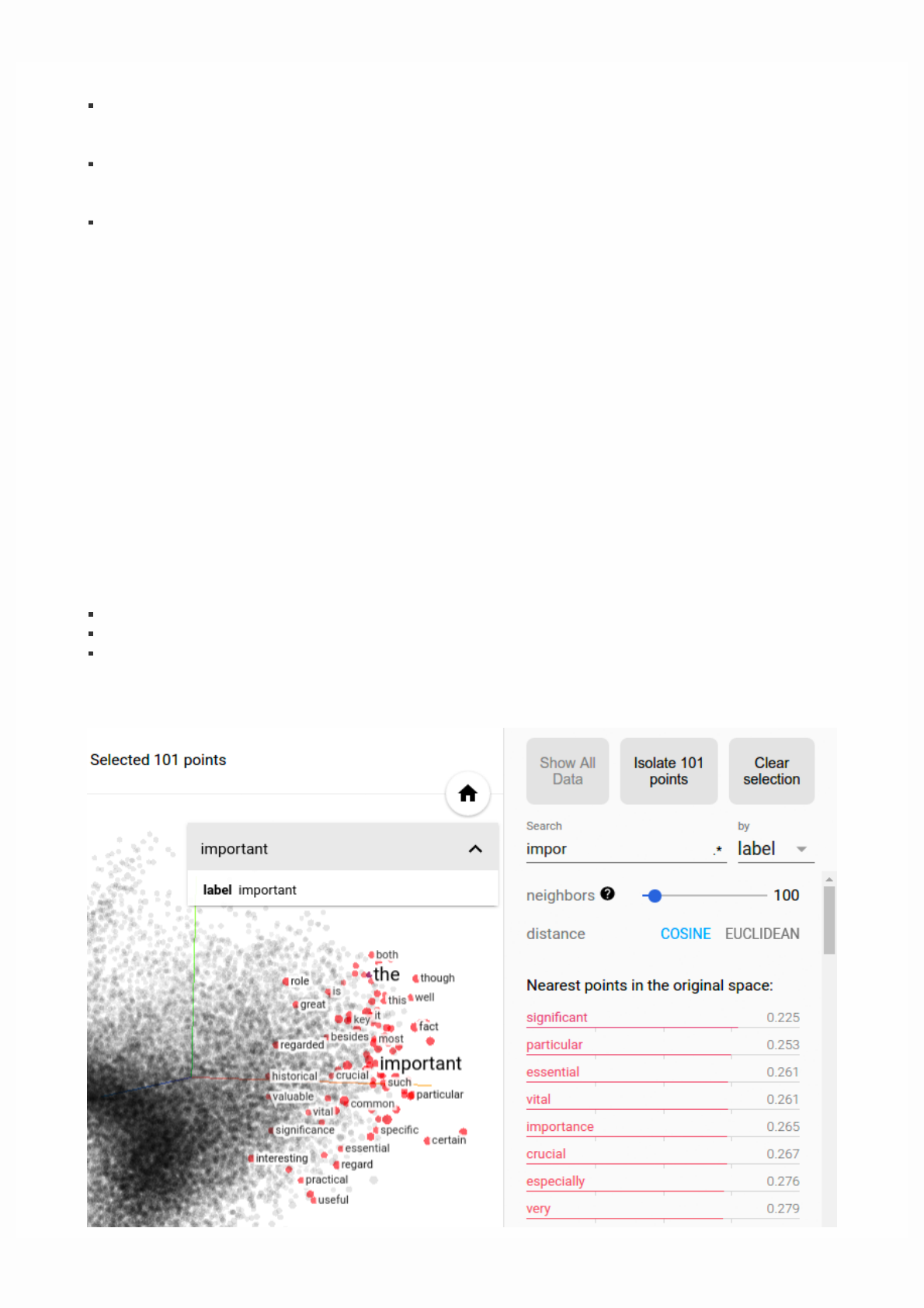
The Embedding Projector provides three ways to reduce the dimensionality of a data set.
t-SNE: a nonlinear nondeterministic algorithm (T-distributed stochastic neighbor embedding) that tries to preserve local
neighborhoods in the data, often at the expense of distorting global structure. You can choose whether to compute two- or
three-dimensional projections.
PCA: a linear deterministic algorithm (principal component analysis) that tries to capture as much of the data variability in
as few dimensions as possible. PCA tends to highlight large-scale structure in the data, but can distort local neighborhoods.
The Embedding Projector computes the top 10 principal components, from which you can choose two or three to view.
Custom: a linear projection onto horizontal and vertical axes that you specify using labels in the data. You define the
horizontal axis, for instance, by giving text patterns for "Left" and "Right". The Embedding Projector finds all points whose
label matches the "Left" pattern and computes the centroid of that set; similarly for "Right". The line passing through these
two centroids defines the horizontal axis. The vertical axis is likewise computed from the centroids for points matching the
"Up" and "Down" text patterns.
Further useful articles are How to Use t-SNE Effectively and Principal Component Analysis Explained Visually.
Exploration
You can explore visually by zooming, rotating, and panning using natural click-and-drag gestures. Hovering your mouse over
a point will show any metadata for that point. You can also inspect nearest-neighbor subsets. Clicking on a point causes the
right pane to list the nearest neighbors, along with distances to the current point. The nearest-neighbor points are also
highlighted in the projection.
It is sometimes useful to restrict the view to a subset of points and perform projections only on those points. To do so, you can
select points in multiple ways:
After clicking on a point, its nearest neighbors are also selected.
After a search, the points matching the query are selected.
Enabling selection, clicking on a point and dragging defines a selection sphere.
Then click the "Isolate nnn points" button at the top of the Inspector pane on the right hand side. The following image shows
101 points selected and ready for the user to click "Isolate 101 points":
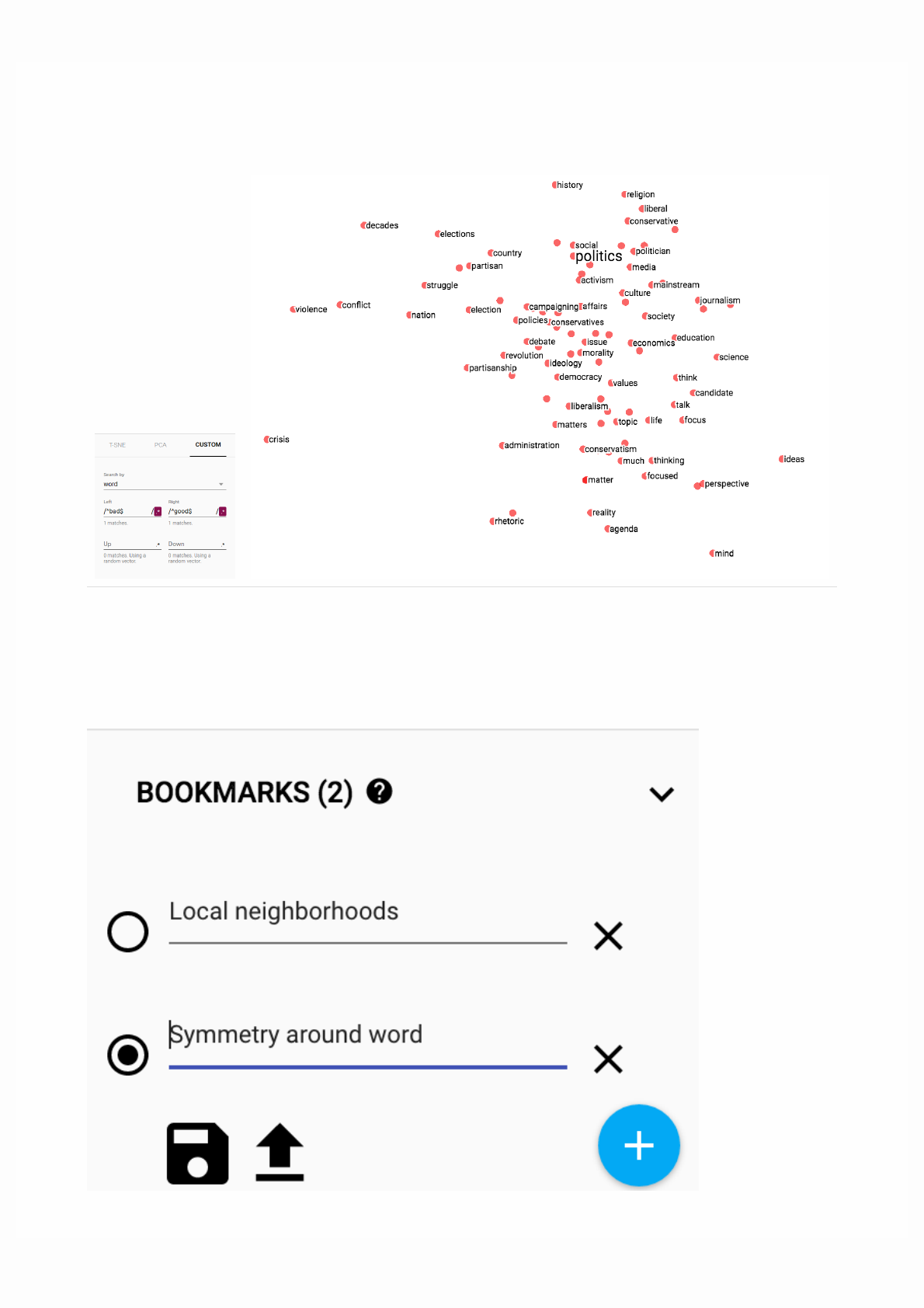
Custom projection
controls.
Custom projection of neighbors of "politics" onto "best" - "worst" vector.
Selection of the nearest neighbors of “important” in a word embedding dataset.
Advanced tip: filtering with custom projection can be powerful. Below, we filtered the 100 nearest neighbors of “politics” and
projected them onto the “worst” - “best” vector as an x axis. The y axis is random. As a result, one finds on the right side
“ideas”, “science”, “perspective”, “journalism” but on the left “crisis”, “violence” and “conflict”.
To share your findings, you can use the bookmark panel in the bottom right corner and save the current state (including
computed coordinates of any projection) as a small file. The Projector can then be pointed to a set of one or more of these files,
producing the panel below. Other users can then walk through a sequence of bookmarks.

0 1 2
3 4 5
6 7
Metadata
If you are working with an embedding, you'll probably want to attach labels/images to the data points. You can do this by
generating a metadata file containing the labels for each point and clicking "Load data" in the data panel of the Embedding
Projector.
The metadata can be either labels or images, which are stored in a separate file. For labels, the format should be a TSV file (tab
characters shown in red) whose first line contains column headers (shown in bold) and subsequent lines contain the metadata
values. For example:
**Word\tFrequency**Airplane\t345Car\t241...
The order of lines in the metadata file is assumed to match the order of vectors in the embedding variable, except for the
header. Consequently, the (i+1)-th line in the metadata file corresponds to the i-th row of the embedding variable. If the TSV
metadata file has only a single column, then we don’t expect a header row, and assume each row is the label of the embedding.
We include this exception because it matches the commonly-used "vocab file" format.
To use images as metadata, you must produce a single sprite image, consisting of small thumbnails, one for each vector in the
embedding. The sprite should store thumbnails in row-first order: the first data point placed in the top left and the last data
point in the bottom right, though the last row doesn't have to be filled, as shown below.
Follow this link to see a fun example of thumbnail images in the Embedding Projector.
Mini-FAQ
Is "embedding" an action or a thing? Both. People talk about embedding words in a vector space (action) and about producing
word embeddings (things). Common to both is the notion of embedding as a mapping from discrete objects to vectors.
Creating or applying that mapping is an action, but the mapping itself is a thing.
Are embeddings high-dimensional or low-dimensional? It depends. A 300-dimensional vector space of words and phrases,
for instance, is often called low-dimensional (and dense) when compared to the millions of words and phrases it can contain.
But mathematically it is high-dimensional, displaying many properties that are dramatically different from what our human
intuition has learned about 2- and 3-dimensional spaces.
Is an embedding the same as an embedding layer? No. An embedding layer is a part of neural network, but an embedding is a
more general concept.
Debugging TensorFlow Programs
TensorFlow debugger (tfdbg) is a specialized debugger for TensorFlow. It lets you view the internal structure and states of
running TensorFlow graphs during training and inference, which is difficult to debug with general-purpose debuggers such as
Python's pdb due to TensorFlow's computation-graph paradigm.
NOTE: TensorFlow debugger uses a curses-based text user interface. On Mac OS X, the ncurses library is required and
can be installed with brew install homebrew/dupes/ncurses. On Windows, curses isn't as well supported, so a
readline-based interface can be used with tfdbg by installing pyreadline with pip. If you use Anaconda3, you can install
it with a command such as "C:\Program Files\Anaconda3\Scripts\pip.exe" install pyreadline. Unofficial
Windows curses packages can be downloaded here, then subsequently installed using pip install

<your_version>.whl, however curses on Windows may not work as reliably as curses on Linux or Mac.
This tutorial demonstrates how to use the tfdbg command-line interface (CLI) to debug the appearance of nans and infs, a
frequently-encountered type of bug in TensorFlow model development. The following example is for users who use the low-
level Session API of TensorFlow. A later section of this document describes how to use tfdbg with a higher-level API, namely
tf-learn Estimators and Experiments. To observesuch an issue, run the following command without the debugger (the source
code can be found here):
python -m tensorflow.python.debug.examples.debug_mnist
This code trains a simple neural network for MNIST digit image recognition. Notice that the accuracy increases slightly after
the first training step, but then gets stuck at a low (near-chance) level:
Accuracy at step 0: 0.1113
Accuracy at step 1: 0.3183
Accuracy at step 2: 0.098
Accuracy at step 3: 0.098
Accuracy at step 4: 0.098
Wondering what might have gone wrong, you suspect that certain nodes in the training graph generated bad numeric values
such as infs and nans, because this is a common cause of this type of training failure. Let's use tfdbg to debug this issue and
pinpoint the exact graph node where this numeric problem first surfaced.
Wrapping TensorFlow Sessions with tfdbg
To add support for tfdbg in our example, all that is needed is to add the following lines of code and wrap the Session object
with a debugger wrapper. This code is already added in debug_mnist.py, so you can activate tfdbg CLI with the --debug flag
at the command line.
# Let your BUILD target depend on "//tensorflow/python/debug:debug_py"
# (You don't need to worry about the BUILD dependency if you are using a pip
# install of open-source TensorFlow.)
from tensorflow.python import debug as tf_debug
sess = tf_debug.LocalCLIDebugWrapperSession(sess)
This wrapper has the same interface as Session, so enabling debugging requires no other changes to the code. The wrapper
provides additional features, including:
Bringing up a CLI before and after Session.run() calls, to let you control the execution and inspect the graph's internal
state.
Allowing you to register special filters for tensor values, to facilitate the diagnosis of issues.
In this example, we have already registered a tensor filter called tfdbg.has_inf_or_nan, which simply determines if there are
any nan or inf values in any intermediate tensors (tensors that are neither inputs or outputs of the Session.run() call, but
are in the path leading from the inputs to the outputs). This filter is for nans and infs is a common enough use case that we
ship it with the debug_data module.
Note: You can also write your own custom filters. See the API documentation of DebugDumpDir.find() for additional
information.
Debugging Model Training with tfdbg
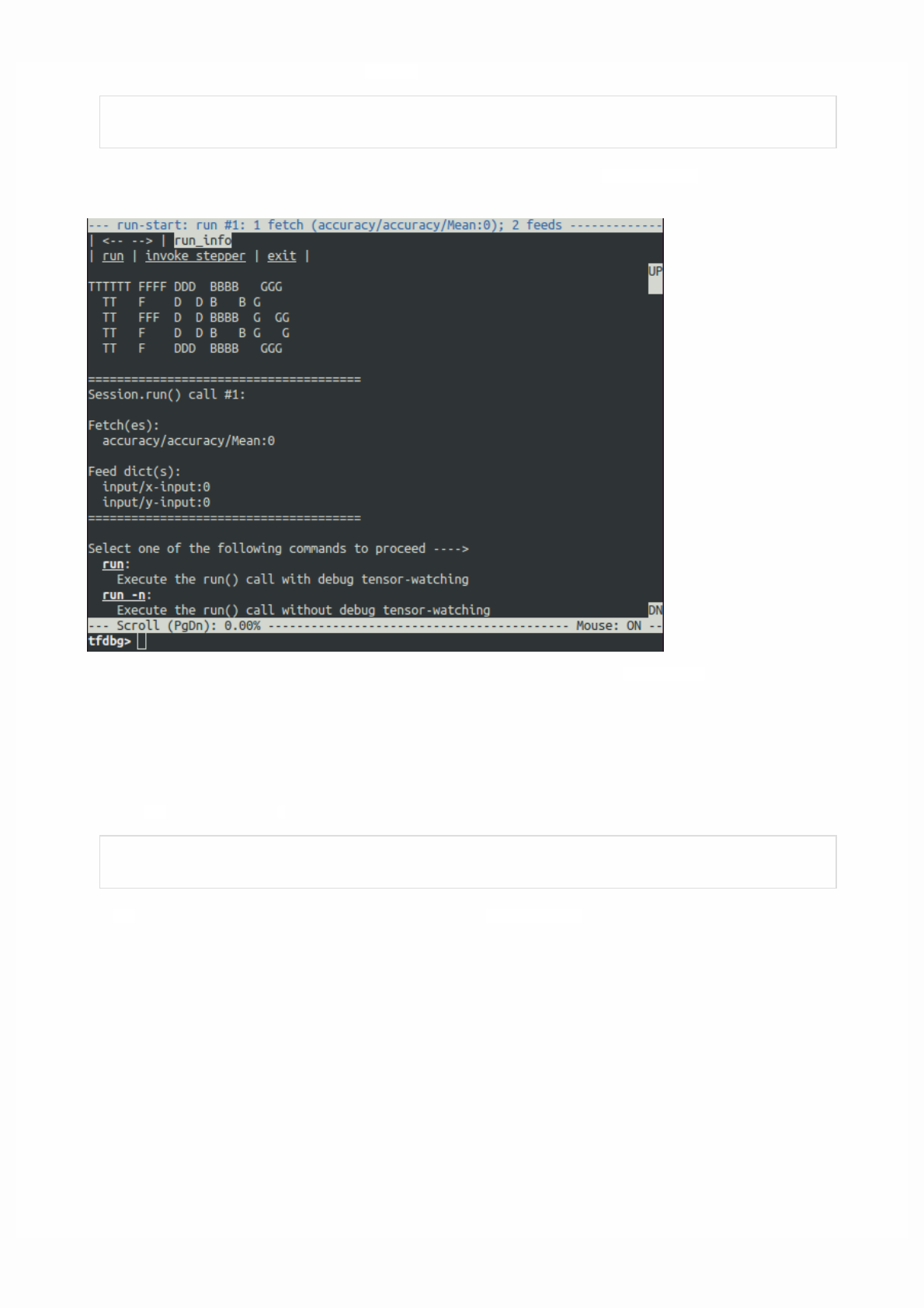
Let's try training the model again, but with the --debug flag added this time:
python -m tensorflow.python.debug.examples.debug_mnist --debug
The debug wrapper session will prompt you when it is about to execute the first Session.run() call, with information
regarding the fetched tensor and feed dictionaries displayed on the screen.
This is what we refer to as the run-start CLI. It lists the feeds and fetches to the current Session.run call, before executing
anything.
If the screen size is too small to display the content of the message in its entirety, you can resize it.
Use the PageUp / PageDown / Home / End keys to navigate the screen output. On most keyboards lacking those keys Fn +
Up / Fn + Down / Fn + Right / Fn + Left will work.
Enter the run command (or just r) at the command prompt:
tfdbg> run
The run command causes tfdbg to execute until the end of the next Session.run() call, which calculates the model's accuracy
using a test data set. tfdbg augments the runtime Graph to dump all intermediate tensors. After the run ends, tfdbg displays all
the dumped tensors values in the run-end CLI. For example:
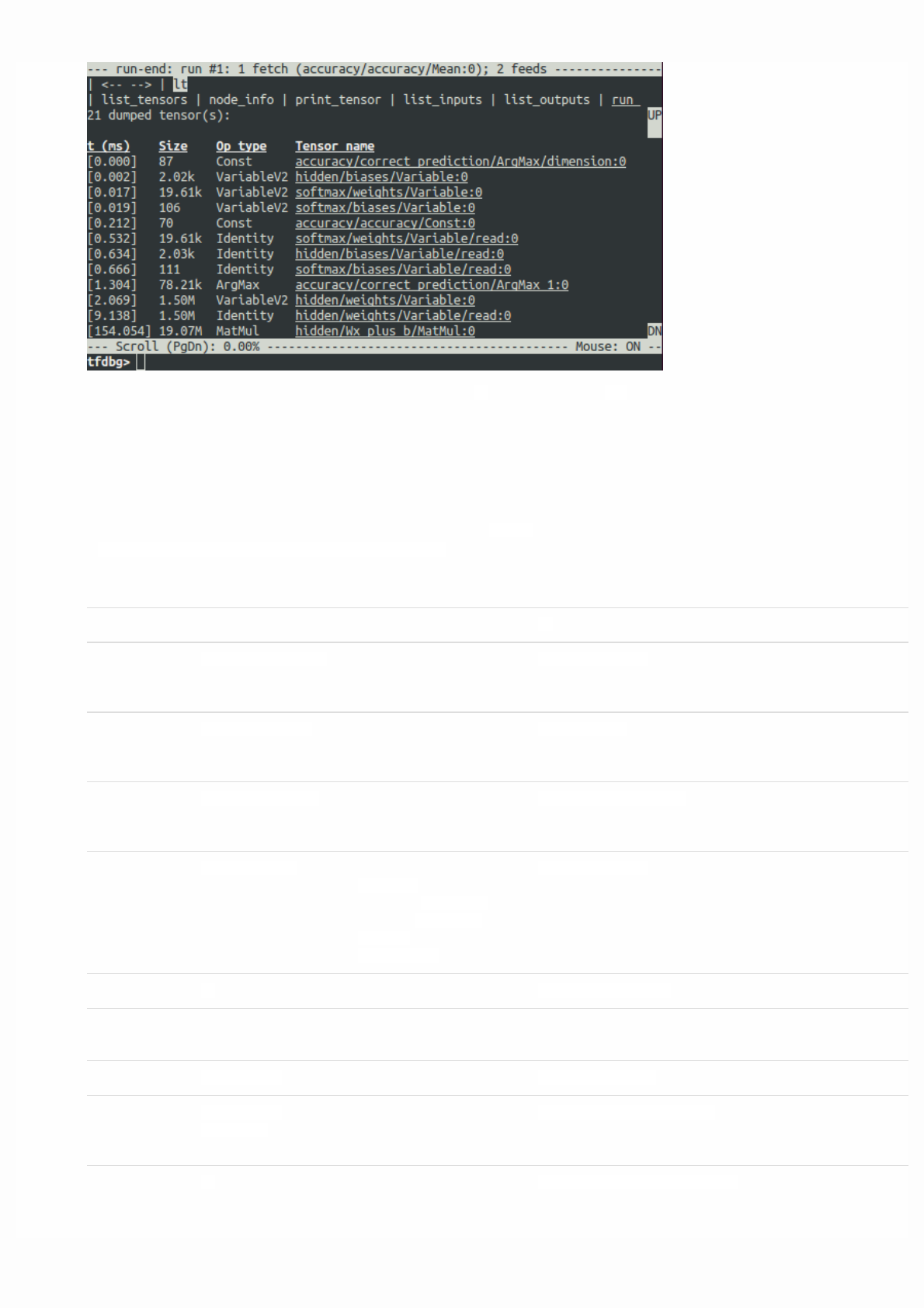
Command Syntax or Option Explanation Example
lt List dumped tensors. lt
-n <name_pattern> List dumped tensors with
names matching given
regular-expression pattern.
lt -n Softmax.*
-t <op_pattern> List dumped tensors with
op types matching given
regular-expression pattern.
lt -t MatMul
-f <filter_name> List only the tensors that
pass a registered tensor
filter.
lt -f has_inf_or_nan
-s <sort_key> Sort the output by given
sort_key, whose possible
values are timestamp
(default), dump_size,
op_type and
tensor_name.
lt -s dump_size
-r Sort in reverse order. lt -r -s dump_size
pt Print value of a dumped
tensor.
pt <tensor> Print tensor value. pt hidden/Relu:0
pt <tensor>
[slicing]
Print a subarray of tensor,
using numpy-style array
slicing.
pt hidden/Relu:0[0:50,:]
-a Print the entirety of a large
tensor, without using
ellipses. (May take a long
pt -a hidden/Relu:0[0:50,:]
This list of tensors can also be obtained by running the command lt after you executed run.
tfdbg CLI Frequently-Used Commands
Try the following commands at the tfdbg> prompt (referencing the code
attensorflow/python/debug/examples/debug_mnist.py):
time for large tensors.)
-r <range> Highlight elements falling
into specified numerical
range. Multiple ranges can
be used in conjunction.
pt hidden/Relu:0 -a -r [[-inf,-1],[1,inf]]
-n <number> Print dump corresponding
to specified 0-based dump
number. Required for
tensors with multiple
dumps.
pt -n 0 hidden/Relu:0
-s Include a summary of the
numeric values of the
tensor (applicable only to
non-empty tensors with
Boolean and numeric types
such as int* and float*.)
pt -s hidden/Relu:0[0:50,:]
@[coordinates] Navigate to specified
element in pt output.
@[10,0] or @10,0
/regex less-style search for given
regular expression.
/inf
/Scroll to the next line with
matches to the searched
regex (if any).
/
pf Print a value in the
feed_dict to Session.run.
pf
<feed_tensor_name>
Print the value of the feed.
Also note that the pf
command has the -a, -r
and -sflags (not listed
below), which have the
same syntax and semantics
as the identically-named
flags of pt.
pf input_xs:0
eval Evaluate arbitrary Python
and numpy expression.
eval <expression> Evaluate a Python /
numpy expression, with
numpy available as np and
debug tensor names
enclosed in backticks.
eval
"np.matmul((output/Identity:0/Softmax:0).T,Softmax:0)"
-a Print a large-sized
evaluation result in its
entirety, i.e., without using
ellipses.
eval -a 'np.sum(Softmax:0, axis=1)'
ni Display node information.
-a Include node attributes in
the output.
ni -a hidden/Relu
-d List the debug dumps
available from the node.
ni -d hidden/Relu
-t Display the Python stack
trace of the node's creation.
ni -t hidden/Relu
li List inputs to node
-r List the inputs to node,
recursively (the input tree.)
li -r hidden/Relu:0
-d <max_depth> Limit recursion depth
under the -r mode.
li -r -d 3 hidden/Relu:0
-c Include control inputs. li -c -r hidden/Relu:0
-t Show op types of input
nodes.
li -t -r hidden/Relu:0
lo List output recipients of
node
-r List the output recipients of
node, recursively (the
output tree.)
lo -r hidden/Relu:0
-d <max_depth> Limit recursion depth
under the -r mode.
lo -r -d 3 hidden/Relu:0
-c Include recipients via
control edges.
lo -c -r hidden/Relu:0
-t Show op types of recipient
nodes.
lo -t -r hidden/Relu:0
ls List Python source files
involved in node creation.
-p <path_pattern> Limit output to source files
matching given regular-
expression path pattern.
ls -p .*debug_mnist.*
-n Limit output to node
names matching given
regular-expression pattern.
ls -n Softmax.*
ps Print Python source file.
ps <file_path> Print given Python source
file source.py, with the
lines annotated with the
nodes created at each of
them (if any).
ps /path/to/source.py
-t Perform annotation with
respect to Tensors, instead
of the default, nodes.
ps -t /path/to/source.py
-b <line_number> Annotate source.py
beginning at given line.
ps -b 30 /path/to/source.py
-m <max_elements> Limit the number of
elements in the annotation
for each line.
ps -m 100 /path/to/source.py
run Proceed to the next
Session.run()
run
-n Execute through the next
Session.runwithout
debugging, and drop to
CLI right before the run
after that.
run -n

-t <T> Execute Session.run T -
1 times without
debugging, followed by a
run with debugging. Then
drop to CLI right after the
debugged run.
run -t 10
-f <filter_name> Continue executing
Session.run until any
intermediate tensor
triggers the specified
Tensor filter (causes the
filter to return True).
run -f has_inf_or_nan
--node_name_filter
<pattern>
Execute the next
Session.run, watching
only nodes with names
matching the given
regular-expression pattern.
run --node_name_filter Softmax.*
--op_type_filter
<pattern>
Execute the next
Session.run, watching
only nodes with op types
matching the given
regular-expression pattern.
run --op_type_filter Variable.*
--
tensor_dtype_filter
<pattern>
Execute the next
Session.run, dumping
only Tensors with data
types (dtypes) matching
the given regular-
expression pattern.
run --tensor_dtype_filter int.*
-p Execute the next
Session.run call in
profiling mode.
run -p
ri Display information about
the run the current run,
including fetches and
feeds.
ri
config Set or show persistent
TFDBG UI configuration.
set Set the value of a config
item:
{graph_recursion_depth,
mouse_mode}.
config set graph_recursion_depth 3
show Show current persistent UI
configuration.
config show
help Print general help
information
help
help <command> Print help for given
command.
help lt
Note that each time you enter a command, a new screen output will appear. This is somewhat analogous to web pages in a
browser. You can navigate between these screens by clicking the <-- and --> text arrows near the top-left corner of the CLI.

Other Features of the tfdbg CLI
In addition to the commands listed above, the tfdbg CLI provides the following addditional features:
To navigate through previous tfdbg commands, type in a few characters followed by the Up or Down arrow keys. tfdbg will
show you the history of commands that started with those characters.
To navigate through the history of screen outputs, do either of the following:
Use the prev and next commands.
Click underlined <-- and --> links near the top left corner of the screen.
Tab completion of commands and some command arguments.
To redirect the screen output to a file instead of the screen, end the command with bash-style redirection. For example, the
following command redirects the output of the pt command to the /tmp/xent_value_slices.txt file:
tfdbg> pt cross_entropy/Log:0[:, 0:10] > /tmp/xent_value_slices.txt
Finding nans and infs
In this first Session.run() call, there happen to be no problematic numerical values. You can move on to the next run by
using the command run or its shorthand r.
TIP: If you enter run or r repeatedly, you will be able to move through the Session.run() calls in a sequential manner.
You can also use the -t flag to move ahead a number of Session.run() calls at a time, for example:
tfdbg> run -t 10
Instead of entering run repeatedly and manually searching for nans and infs in the run-end UI after every Session.run()
call (for example, by using the pt command shown in the table above) , you can use the following command to let the
debugger repeatedly execute Session.run() calls without stopping at the run-start or run-end prompt, until the first nan or
inf value shows up in the graph. This is analogous to conditional breakpoints in some procedural-language debuggers:
tfdbg> run -f has_inf_or_nan
NOTE: The preceding command works properly because a tensor filter called has_inf_or_nan has been registered for
you when the wrapped session is created. This filter detects nans and infs (as explained previously). If you have
registered any other filters, you can use "run -f" to have tfdbg run until any tensor triggers that filter (cause the filter to
return True).
def my_filter_callable(datum, tensor):
# A filter that detects zero-valued scalars.
return len(tensor.shape) == 0 and tensor == 0.0
sess.add_tensor_filter('my_filter', my_filter_callable)
Then at the tfdbg run-start prompt run until your filter is triggered:
tfdbg> run -f my_filter
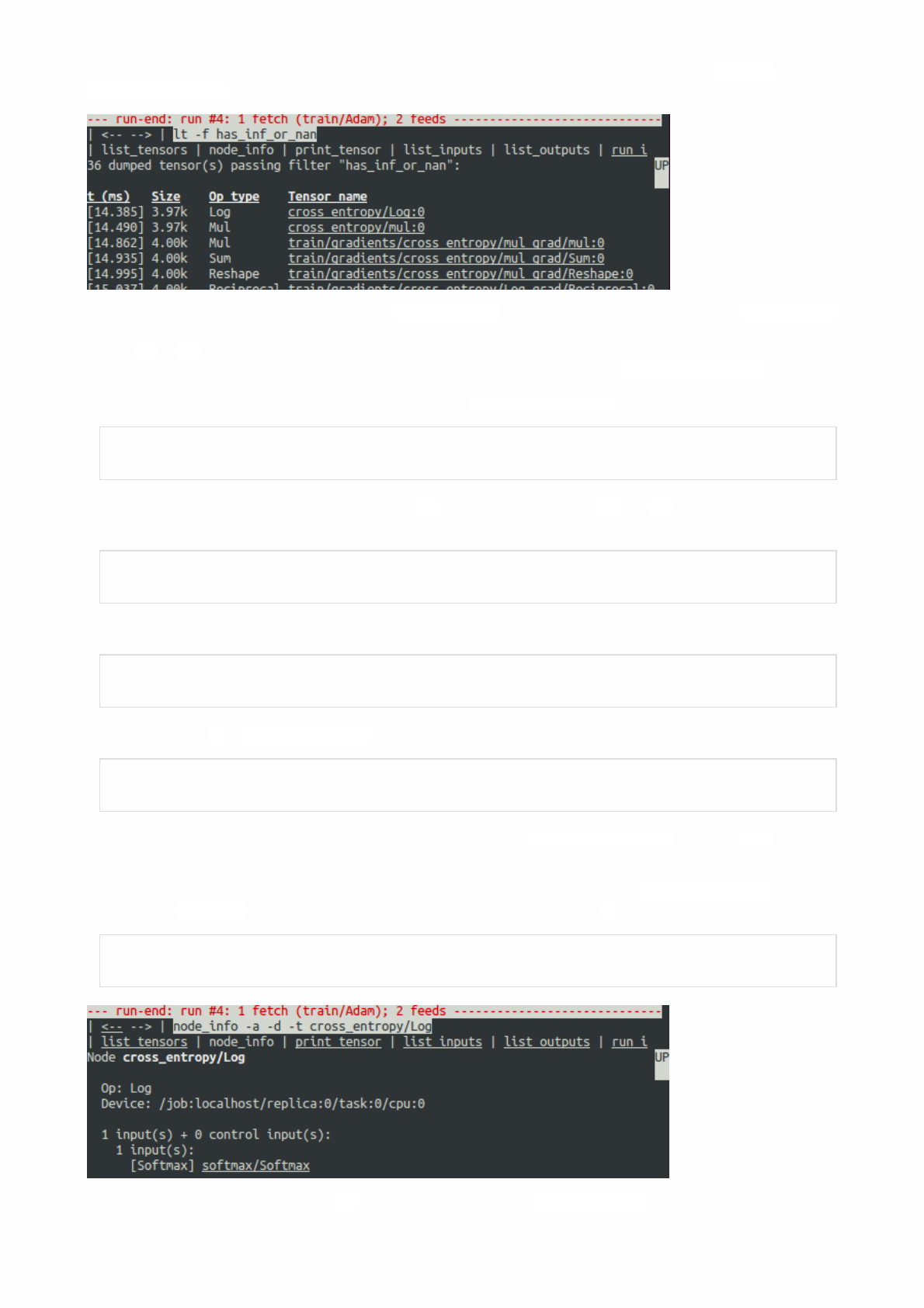
See this API document for more information on the expected signature and return value of the predicateCallable used with
add_tensor_filter().
As the screen display indicates on the first line, the has_inf_or_nan filter is first triggered during the fourth Session.run()
call: an Adam optimizer forward-backward training pass on the graph. In this run, 36 (out of the total 95) intermediate tensors
contain nan or inf values. These tensors are listed in chronological order, with their timestamps displayed on the left. At the
top of the list, you can see the first tensor in which the bad numerical values first surfaced: cross_entropy/Log:0.
To view the value of the tensor, click the underlined tensor name cross_entropy/Log:0 or enter the equivalent command:
tfdbg> pt cross_entropy/Log:0
Scroll down a little and you will notice some scattered inf values. If the instances of inf and nan are difficult to spot by eye,
you can use the following command to perform a regex search and highlight the output:
tfdbg> /inf
Or, alternatively:
tfdbg> /(inf|nan)
You can also use the -s or --numeric_summary command to get a quick summary of the types of numeric values in the tensor:
tfdbg> pt -s cross_entropy/Log:0
From the summary, you can see that several of the 1000 elements of the cross_entropy/Log:0 tensor are -infs (negative
infinities).
Why did these infinities appear? To further debug, display more information about the node cross_entropy/Log by clicking
the underlined node_info menu item on the top or entering the equivalent node_info (ni) command:
tfdbg> ni cross_entropy/Log
You can see that this node has the op type Log and that its input is the node softmax/Softmax. Run the following command to
take a closer look at the input tensor:
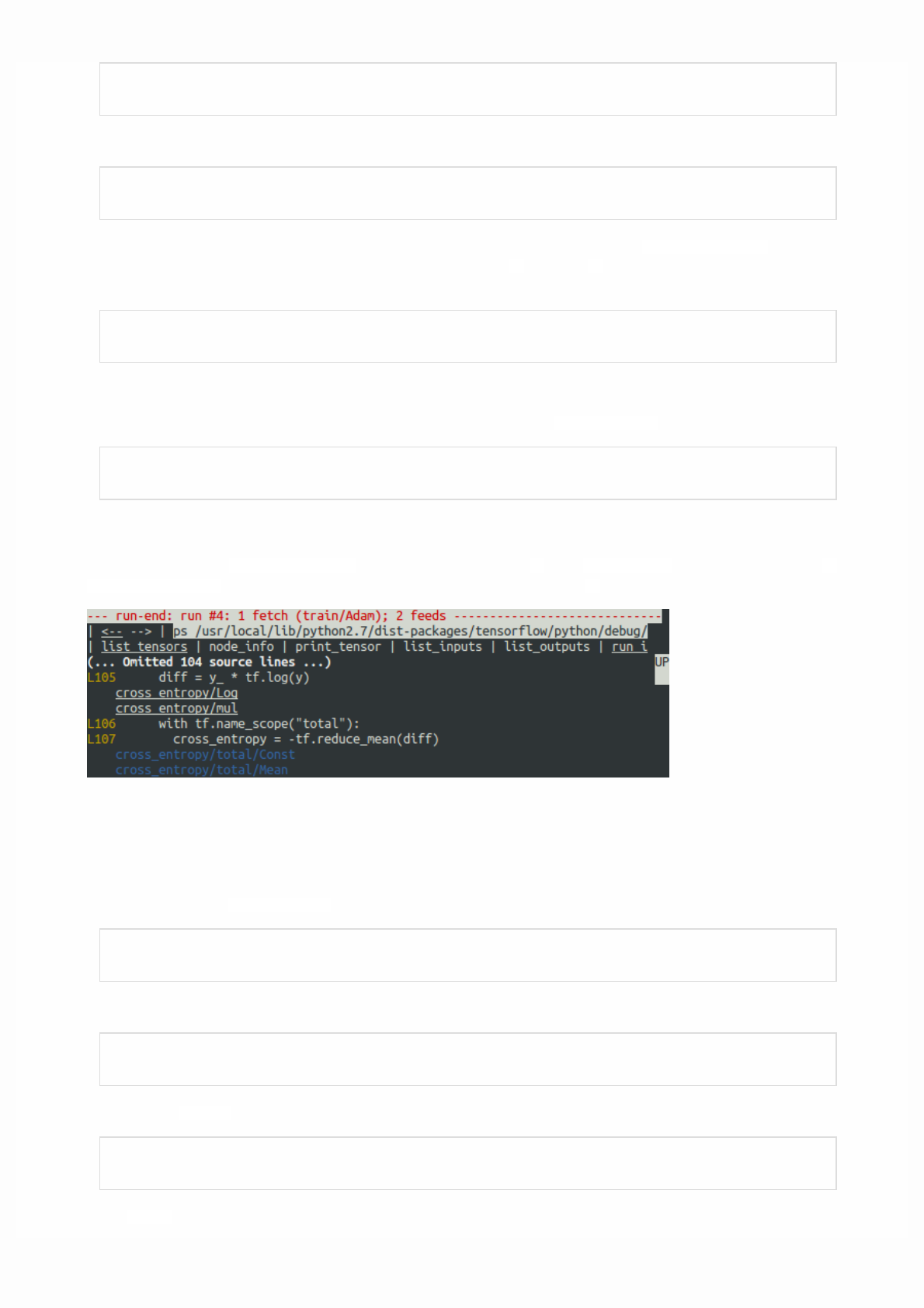
tfdbg> pt softmax/Softmax:0
Examine the values in the input tensor, searching for zeros:
tfdbg> /0\.000
Indeed, there are zeros. Now it is clear that the origin of the bad numerical values is the node cross_entropy/Log taking logs
of zeros. To find out the culprit line in the Python source code, use the -tflag of the ni command to show the traceback of the
node's construction:
tfdbg> ni -t cross_entropy/Log
If you click "node_info" at the top of the screen, tfdbg automatically shows the traceback of the node's construction.
From the traceback, you can see that the op is constructed at the following line: debug_mnist.py:
diff = y_ * tf.log(y)
tfdbg has a feature that makes it easy to trace Tensors and ops back to lines in Python source files. It can annotate lines of a
Python file with the ops or Tensors created by them. To use this feature, simply click the underlined line numbers in the stack
trace output of the ni -t <op_name> commands, or use the ps (or print_source) command such as: ps
/path/to/source.py. For example, the following screenshot shows the output of a ps command.
Fixing the problem
To fix the problem, edit debug_mnist.py, changing the original line:
diff = -(y_ * tf.log(y))
to the built-in, numerically-stable implementation of softmax cross-entropy:
diff = tf.losses.sparse_softmax_cross_entropy(labels=y_, logits=logits)
Rerun with the --debug flag as follows:
python -m tensorflow.python.debug.examples.debug_mnist --debug
At the tfdbg> prompt, enter the following command:

run -f has_inf_or_nan`
Confirm that no tensors are flagged as containing nan or inf values, and accuracy now continues to rise rather than getting
stuck. Success!
Debugging tf-learn Estimators and Experiments
This section explains how to debug TensorFlow programs that use the Estimator and Experiment APIs. Part of the
convenience provided by these APIs is that they manage Sessions internally. This makes the
LocalCLIDebugWrapperSession described in the preceding sections inapplicable. Fortunately, you can still debug them by
using special hooks provided by tfdbg.
Debugging tf.contrib.learn Estimators
Currently, tfdbg can debug the fit() evaluate() methods of tf-learn Estimators. To debug Estimator.fit(), create a
LocalCLIDebugHook and supply it in the monitors argument. For example:
# First, let your BUILD target depend on "//tensorflow/python/debug:debug_py"
# (You don't need to worry about the BUILD dependency if you are using a pip
# install of open-source TensorFlow.)
from tensorflow.python import debug as tf_debug
# Create a LocalCLIDebugHook and use it as a monitor when calling fit().
hooks = [tf_debug.LocalCLIDebugHook()]
classifier.fit(x=training_set.data,
y=training_set.target,
steps=1000,
monitors=hooks)
To debug Estimator.evaluate(), assign hooks to the hooks parameter, as in the following example:
accuracy_score = classifier.evaluate(x=test_set.data,
y=test_set.target,
hooks=hooks)["accuracy"]
debug_tflearn_iris.py, based on {$tflearn$tf-learn's iris tutorial}, contains a full example of how to use the tfdbg with
Estimators. To run this example, do:
python -m tensorflow.python.debug.examples.debug_tflearn_iris --debug
Debugging tf.contrib.learn Experiments
Experiment is a construct in tf.contrib.learn at a higher level than Estimator. It provides a single interface for training
and evaluating a model. To debug the train() and evaluate() calls to an Experimentobject, you can use the keyword
arguments train_monitors and eval_hooks, respectively, when calling its constructor. For example:

# First, let your BUILD target depend on "//tensorflow/python/debug:debug_py"
# (You don't need to worry about the BUILD dependency if you are using a pip
# install of open-source TensorFlow.)
from tensorflow.python import debug as tf_debug
hooks = [tf_debug.LocalCLIDebugHook()]
ex = experiment.Experiment(classifier,
train_input_fn=iris_input_fn,
eval_input_fn=iris_input_fn,
train_steps=FLAGS.train_steps,
eval_delay_secs=0,
eval_steps=1,
train_monitors=hooks,
eval_hooks=hooks)
ex.train()
accuracy_score = ex.evaluate()["accuracy"]
To build and run the debug_tflearn_iris example in the Experiment mode, do:
python -m tensorflow.python.debug.examples.debug_tflearn_iris \
--use_experiment --debug
The LocalCLIDebugHook also allows you to configure a watch_fn that can be used to flexibly specify what Tensors to watch
on different Session.run() calls, as a function of the fetches and feed_dict and other states. See this API doc for more
details.
Debugging Keras Models with TFDBG
To use TFDBG with Keras, let the Keras backend use a TFDBG-wrapped Session object. For example, to use the CLI wrapper:
import tensorflow as tf
from keras import backend as keras_backend
from tensorflow.python import debug as tf_debug
keras_backend.set_session(tf_debug.LocalCLIDebugWrapperSession(tf.Session()))
# Define your keras model, called "model".
model.fit(...) # This will break into the TFDBG CLI.
Debugging tf-slim with TFDBG
TFDBG supports debugging of training and evaluation with tf-slim. As detailed below, training and evaluation require slightly
different debugging workflows.
Debugging training in tf-slim

To debug the training process, provide LocalCLIDebugWrapperSession to the session_wrapper argument of
slim.learning.train(). For example:
import tensorflow as tf
from tensorflow.python import debug as tf_debug
# ... Code that creates the graph and the train_op ...
tf.contrib.slim.learning.train(
train_op,
logdir,
number_of_steps=10,
session_wrapper=tf_debug.LocalCLIDebugWrapperSession)
Debugging evaluation in tf-slim
To debug the evaluation process, provide LocalCLIDebugHook to the hooks argument of
slim.evaluation.evaluate_once(). For example:
import tensorflow as tf
from tensorflow.python import debug as tf_debug
# ... Code that creates the graph and the eval and final ops ...
tf.contrib.slim.evaluation.evaluate_once(
'',
checkpoint_path,
logdir,
eval_op=my_eval_op,
final_op=my_value_op,
hooks=[tf_debug.LocalCLIDebugHook()])
Offline Debugging of Remotely-Running Sessions
Often, your model is running on a remote machine or a process that you don't have terminal access to. To perform model
debugging in such cases, you can use the offline_analyzer binary of tfdbg (described below). It operates on dumped data
directories. This can be done to both the lower-level Session API and the higher-level Estimator and Experiment APIs.
Debugging Remote tf.Sessions
If you interact directly with the tf.Session API in python, you can configure the RunOptions proto that you call your
Session.run() method with, by using the method tfdbg.watch_graph. This will cause the intermediate tensors and
runtime graphs to be dumped to a shared storage location of your choice when the Session.run() call occurs (at the cost of
slower performance). For example:

from tensorflow.python import debug as tf_debug
# ... Code where your session and graph are set up...
run_options = tf.RunOptions()
tf_debug.watch_graph(
run_options,
session.graph,
debug_urls=["file:///shared/storage/location/tfdbg_dumps_1"])
# Be sure to specify different directories for different run() calls.
session.run(fetches, feed_dict=feeds, options=run_options)
Later, in an environment that you have terminal access to (for example, a local computer that can access the shared storage
location specified in the code above), you can load and inspect the data in the dump directory on the shared storage by using
the offline_analyzer binary of tfdbg. For example:
python -m tensorflow.python.debug.cli.offline_analyzer \
--dump_dir=/shared/storage/location/tfdbg_dumps_1
The Session wrapper DumpingDebugWrapperSession offers an easier and more flexible way to generate file-system dumps
that can be analyzed offline. To use it, simply wrap your session in a tf_debug.DumpingDebugWrapperSession. For example:
# Let your BUILD target depend on "//tensorflow/python/debug:debug_py
# (You don't need to worry about the BUILD dependency if you are using a pip
# install of open-source TensorFlow.)
from tensorflow.python import debug as tf_debug
sess = tf_debug.DumpingDebugWrapperSession(
sess, "/shared/storage/location/tfdbg_dumps_1/", watch_fn=my_watch_fn)
The watch_fn argument accepts a Callable that allows you to configure what tensors to watch on different Session.run()
calls, as a function of the fetches and feed_dict to the run() call and other states.
C++ and other languages
If your model code is written in C++ or other languages, you can also modify the debug_options field of RunOptions to
generate debug dumps that can be inspected offline. See the proto definition for more details.
Debugging Remotely-Running tf-learn Estimators and Experiments
If your remote TensorFlow server runs Estimators, you can use the non-interactive DumpingDebugHook. For example:
# Let your BUILD target depend on "//tensorflow/python/debug:debug_py
# (You don't need to worry about the BUILD dependency if you are using a pip
# install of open-source TensorFlow.)
from tensorflow.python import debug as tf_debug
hooks = [tf_debug.DumpingDebugHook("/shared/storage/location/tfdbg_dumps_1")]

Then this hook can be used in the same way as the LocalCLIDebugHook examples described earlier in this document. As the
training and/or evalution of Estimator or Experiment happens, tfdbg creates directories having the following name
pattern:/shared/storage/location/tfdbg_dumps_1/run_<epoch_timestamp_microsec>_<uuid>. Each directory
corresponds to a Session.run() call that underlies the fit() or evaluate() call. You can load these directories and inspect
them in a command-line interface in an offline manner using the offline_analyzeroffered by tfdbg. For example:
python -m tensorflow.python.debug.cli.offline_analyzer \
--dump_dir="/shared/storage/location/tfdbg_dumps_1/run_<epoch_timestamp_microsec>_<uuid>"
Frequently Asked Questions
Q: Do the timestamps on the left side of the lt output reflect actual performance in a non-debugging session?
A: No. The debugger inserts additional special-purpose debug nodes to the graph to record the values of intermediate tensors.
These nodes slow down the graph execution. If you are interested in profiling your model, check out
1. The profiling mode of tfdbg: tfdbg> run -p.
2. tfprof and other profiling tools for TensorFlow.
Q: How do I link tfdbg against my Session in Bazel? Why do I see an error such as "ImportError: cannot import name debug"?
A: In your BUILD rule, declare dependencies: "//tensorflow:tensorflow_py" and
"//tensorflow/python/debug:debug_py". The first is the dependency that you include to use TensorFlow even without
debugger support; the second enables the debugger. Then, In your Python file, add:
from tensorflow.python import debug as tf_debug
# Then wrap your TensorFlow Session with the local-CLI wrapper.
sess = tf_debug.LocalCLIDebugWrapperSession(sess)
Q: Does tfdbg help debug runtime errors such as shape mismatches?
A: Yes. tfdbg intercepts errors generated by ops during runtime and presents the errors with some debug instructions to the
user in the CLI. See examples:
# Debugging shape mismatch during matrix multiplication.
python -m tensorflow.python.debug.examples.debug_errors \
--error shape_mismatch --debug
# Debugging uninitialized variable.
python -m tensorflow.python.debug.examples.debug_errors \
--error uninitialized_variable --debug
Q: How can I let my tfdbg-wrapped Sessions or Hooks run the debug mode only from the main thread?
A: This is a common use case, in which the Session object is used from multiple threads concurrently. Typically, the child
threads take care of background tasks such as running enqueue operations. Often, you want to debug only the main thread (or
less frequently, only one of the child threads). You can use thethread_name_filter keyword argument of
LocalCLIDebugWrapperSession to achieve this type of thread-selective debugging. For example, to debug from the main
thread only, construct a wrapped Session as follows:
sess = tf_debug.LocalCLIDebugWrapperSession(sess, thread_name_filter="MainThread$")
The above example relies on the fact that main threads in Python have the default name MainThread.

Q: The model I am debugging is very large. The data dumped by tfdbg fills up the free space of my disk. What can I do?
A: You might encounter this problem in any of the following situations:
models with many intermediate tensors
very large intermediate tensors
many tf.while_loop iterations
There are three possible workarounds or solutions:
The constructors of LocalCLIDebugWrapperSession and LocalCLIDebugHook provide a keyword argument, dump_root,
to specify the path to which tfdbg dumps the debug data. You can use it to let tfdbg dump the debug data on a disk with
larger free space. For example:
``` python # For LocalCLIDebugWrapperSession sess =
tf_debug.LocalCLIDebugWrapperSession(dump_root="/with/lots/of/space")
# For LocalCLIDebugHook hooks = [tf_debug.LocalCLIDebugHook(dump_root="/with/lots/of/space")] `` Make sure
that the directory pointed to by dump_root is empty or nonexistent. tfdbg cleans up the dump
directories before exiting. * Reduce the batch size used during the runs. * Use the filtering options
of tfdbg'srun` command to watch only specific nodes in the graph. For example:
tfdbg> run --node_name_filter .*hidden.* tfdbg> run --op_type_filter Variable.* tfdbg> run --
tensor_dtype_filter int.*
The first command above watches only nodes whose name match the regular-expression pattern .*hidden.*. The second
command watches only operations whose name match the pattern Variable.*. The third one watches only the tensors whose
dtype match the pattern int.* (e.g., int32).
Q: Why can't I select text in the tfdbg CLI?
A: This is because the tfdbg CLI enables mouse events in the terminal by default. This mouse-mask mode overrides default
terminal interactions, including text selection. You can re-enable text selection by using the command mouse off or m off.
Q: Why does the tfdbg CLI show no dumped tensors when I debug code like the following?
a = tf.ones([10], name="a")
b = tf.add(a, a, name="b")
sess = tf.Session()
sess = tf_debug.LocalCLIDebugWrapperSession(sess)
sess.run(b)
A: The reason why you see no data dumped is because every node in the executed TensorFlow graph is constant-folded by the
TensorFlow runtime. In this exapmle, a is a constant tensor; therefore, the fetched tensor b is effectively also a constant tensor.
TensorFlow's graph optimization folds the graph that contains aand b into a single node to speed up future runs of the graph,
which is why tfdbg does not generate any intermediate tensor dumps. However, if a were a tf.Variable, as in the following
example:
import numpy as np
a = tf.Variable(np.ones[10], name="a")
b = tf.add(a, a, name="b")
sess = tf.Session()
sess.run(tf.global_variables_initializer())
sess = tf_debug.LocalCLIDebugWrapperSession(sess)
sess.run(b)
the constant-folding would not occur and tfdbg should show the intermediate tensor dumps.
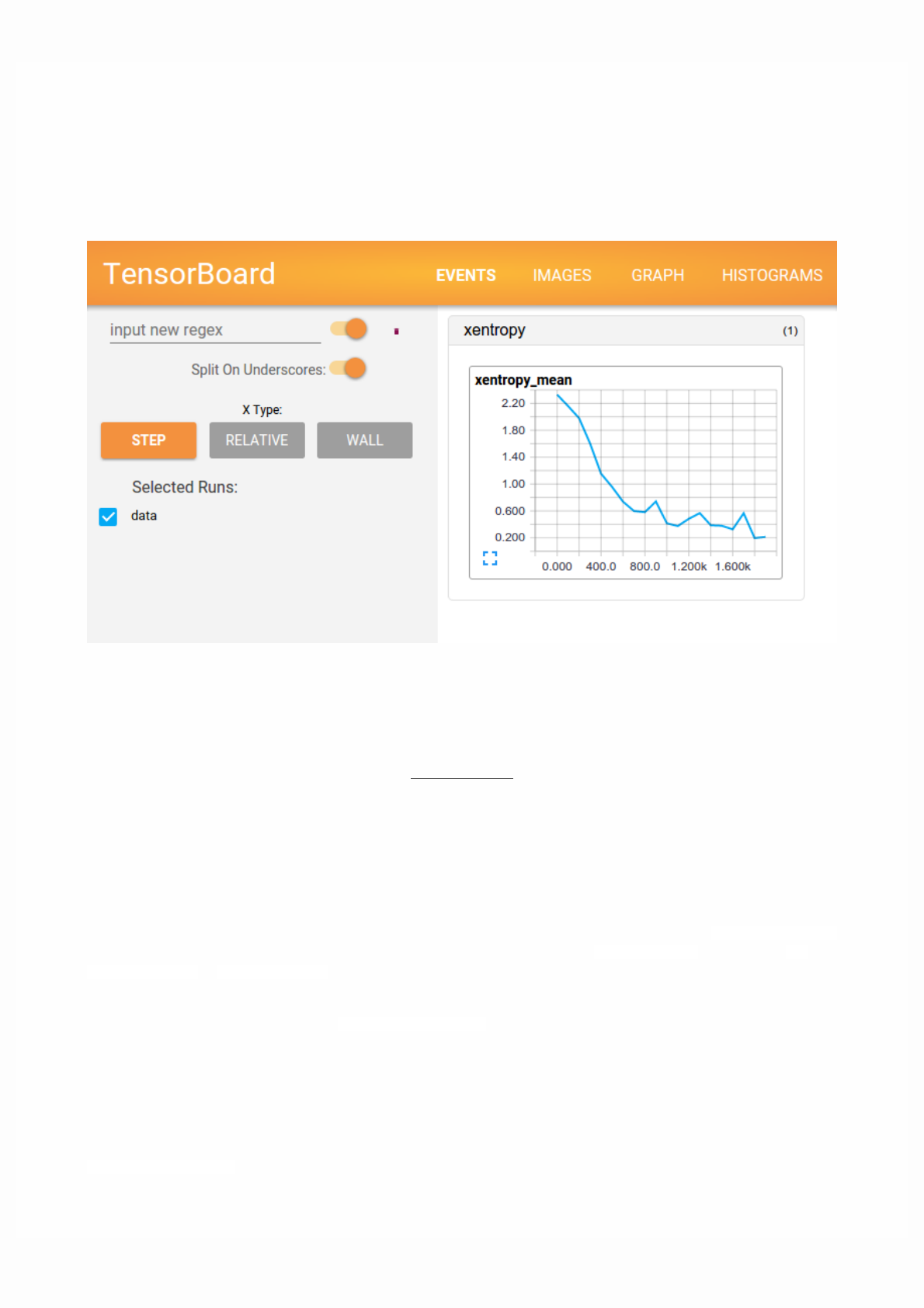
TensorBoard: Visualizing Learning
The computations you'll use TensorFlow for - like training a massive deep neural network - can be complex and confusing. To
make it easier to understand, debug, and optimize TensorFlow programs, we've included a suite of visualization tools called
TensorBoard. You can use TensorBoard to visualize your TensorFlow graph, plot quantitative metrics about the execution of
your graph, and show additional data like images that pass through it. When TensorBoard is fully configured, it looks like this:
This tutorial is intended to get you started with simple TensorBoard usage. There are other resources available as well! The
TensorBoard's GitHub has a lot more information on TensorBoard usage, including tips & tricks, and debugging information.
Serializing the data
TensorBoard operates by reading TensorFlow events files, which contain summary data that you can generate when running
TensorFlow. Here's the general lifecycle for summary data within TensorBoard.
First, create the TensorFlow graph that you'd like to collect summary data from, and decide which nodes you would like to
annotate with summary operations.
For example, suppose you are training a convolutional neural network for recognizing MNIST digits. You'd like to record how
the learning rate varies over time, and how the objective function is changing. Collect these by attaching tf.summary.scalar
ops to the nodes that output the learning rate and loss respectively. Then, give each scalar_summary a meaningful tag, like
'learning rate' or 'loss function'.
Perhaps you'd also like to visualize the distributions of activations coming off a particular layer, or the distribution of gradients
or weights. Collect this data by attaching tf.summary.histogram ops to the gradient outputs and to the variable that holds
your weights, respectively.
For details on all of the summary operations available, check out the docs on summary operations.
Operations in TensorFlow don't do anything until you run them, or an op that depends on their output. And the summary
nodes that we've just created are peripheral to your graph: none of the ops you are currently running depend on them. So, to
generate summaries, we need to run all of these summary nodes. Managing them by hand would be tedious, so use
tf.summary.merge_all to combine them into a single op that generates all the summary data.

Then, you can just run the merged summary op, which will generate a serialized Summary protobuf object with all of your
summary data at a given step. Finally, to write this summary data to disk, pass the summary protobuf to a
tf.summary.FileWriter.
The FileWriter takes a logdir in its constructor - this logdir is quite important, it's the directory where all of the events will be
written out. Also, the FileWriter can optionally take a Graph in its constructor. If it receives a Graph object, then TensorBoard
will visualize your graph along with tensor shape information. This will give you a much better sense of what flows through
the graph: see Tensor shape information.
Now that you've modified your graph and have a FileWriter, you're ready to start running your network! If you want, you
could run the merged summary op every single step, and record a ton of training data. That's likely to be more data than you
need, though. Instead, consider running the merged summary op every n steps.
The code example below is a modification of the simple MNIST tutorial, in which we have added some summary ops, and run
them every ten steps. If you run this and then launch tensorboard --logdir=/tmp/tensorflow/mnist, you'll be able to
visualize statistics, such as how the weights or accuracy varied during training. The code below is an excerpt; full source is
here.
def variable_summaries(var):
"""Attach a lot of summaries to a Tensor (for TensorBoard visualization)."""
with tf.name_scope('summaries'):
mean = tf.reduce_mean(var)
tf.summary.scalar('mean', mean)
with tf.name_scope('stddev'):
stddev = tf.sqrt(tf.reduce_mean(tf.square(var - mean)))
tf.summary.scalar('stddev', stddev)
tf.summary.scalar('max', tf.reduce_max(var))
tf.summary.scalar('min', tf.reduce_min(var))
tf.summary.histogram('histogram', var)
def nn_layer(input_tensor, input_dim, output_dim, layer_name, act=tf.nn.relu):
"""Reusable code for making a simple neural net layer.
It does a matrix multiply, bias add, and then uses relu to nonlinearize.
It also sets up name scoping so that the resultant graph is easy to read,
and adds a number of summary ops.
"""
# Adding a name scope ensures logical grouping of the layers in the graph.
with tf.name_scope(layer_name):
# This Variable will hold the state of the weights for the layer
with tf.name_scope('weights'):
weights = weight_variable([input_dim, output_dim])
variable_summaries(weights)
with tf.name_scope('biases'):
biases = bias_variable([output_dim])
variable_summaries(biases)
with tf.name_scope('Wx_plus_b'):
preactivate = tf.matmul(input_tensor, weights) + biases
tf.summary.histogram('pre_activations', preactivate)
activations = act(preactivate, name='activation')
tf.summary.histogram('activations', activations)
return activations
hidden1 = nn_layer(x, 784, 500, 'layer1')
with tf.name_scope('dropout'):
keep_prob = tf.placeholder(tf.float32)
tf.summary.scalar('dropout_keep_probability', keep_prob)
dropped = tf.nn.dropout(hidden1, keep_prob)
# Do not apply softmax activation yet, see below.
y = nn_layer(dropped, 500, 10, 'layer2', act=tf.identity)

with tf.name_scope('cross_entropy'):
# The raw formulation of cross-entropy,
#
# tf.reduce_mean(-tf.reduce_sum(y_ * tf.log(tf.softmax(y)),
# reduction_indices=[1]))
#
# can be numerically unstable.
#
# So here we use tf.losses.sparse_softmax_cross_entropy on the
# raw logit outputs of the nn_layer above.
with tf.name_scope('total'):
cross_entropy = tf.losses.sparse_softmax_cross_entropy(labels=y_, logits=y)
tf.summary.scalar('cross_entropy', cross_entropy)
with tf.name_scope('train'):
train_step = tf.train.AdamOptimizer(FLAGS.learning_rate).minimize(
cross_entropy)
with tf.name_scope('accuracy'):
with tf.name_scope('correct_prediction'):
correct_prediction = tf.equal(tf.argmax(y, 1), tf.argmax(y_, 1))
with tf.name_scope('accuracy'):
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
tf.summary.scalar('accuracy', accuracy)
# Merge all the summaries and write them out to /tmp/mnist_logs (by default)
merged = tf.summary.merge_all()
train_writer = tf.summary.FileWriter(FLAGS.summaries_dir + '/train',
sess.graph)
test_writer = tf.summary.FileWriter(FLAGS.summaries_dir + '/test')
tf.global_variables_initializer().run()
After we've initialized the FileWriters, we have to add summaries to the FileWriters as we train and test the model.
# Train the model, and also write summaries.
# Every 10th step, measure test-set accuracy, and write test summaries
# All other steps, run train_step on training data, & add training summaries
def feed_dict(train):
"""Make a TensorFlow feed_dict: maps data onto Tensor placeholders."""
if train or FLAGS.fake_data:
xs, ys = mnist.train.next_batch(100, fake_data=FLAGS.fake_data)
k = FLAGS.dropout
else:
xs, ys = mnist.test.images, mnist.test.labels
k = 1.0
return {x: xs, y_: ys, keep_prob: k}
for i in range(FLAGS.max_steps):
if i % 10 == 0: # Record summaries and test-set accuracy
summary, acc = sess.run([merged, accuracy], feed_dict=feed_dict(False))
test_writer.add_summary(summary, i)
print('Accuracy at step %s: %s' % (i, acc))
else: # Record train set summaries, and train
summary, _ = sess.run([merged, train_step], feed_dict=feed_dict(True))
train_writer.add_summary(summary, i)
You're now all set to visualize this data using TensorBoard.

Launching TensorBoard
To run TensorBoard, use the following command (alternatively python -m tensorboard.main)
tensorboard --logdir=path/to/log-directory
where logdir points to the directory where the FileWriter serialized its data. If this logdir directory contains subdirectories
which contain serialized data from separate runs, then TensorBoard will visualize the data from all of those runs. Once
TensorBoard is running, navigate your web browser to localhost:6006 to view the TensorBoard.
When looking at TensorBoard, you will see the navigation tabs in the top right corner. Each tab represents a set of serialized
data that can be visualized.
For in depth information on how to use the graph tab to visualize your graph, see TensorBoard: Graph Visualization.
For more usage information on TensorBoard in general, see the TensorBoard's GitHub.
TensorBoard: Graph Visualization
TensorFlow computation graphs are powerful but complicated. The graph visualization can help you understand and debug
them. Here's an example of the visualization at work.
Visualization of a TensorFlow graph.
To see your own graph, run TensorBoard pointing it to the log directory of the job, click on the graph tab on the top pane and
select the appropriate run using the menu at the upper left corner. For in depth information on how to run TensorBoard and
make sure you are logging all the necessary information, see TensorBoard: Visualizing Learning.
Name scoping and nodes
Typical TensorFlow graphs can have many thousands of nodes--far too many to see easily all at once, or even to lay out using
standard graph tools. To simplify, variable names can be scoped and the visualization uses this information to define a
hierarchy on the nodes in the graph. By default, only the top of this hierarchy is shown. Here is an example that defines three
operations under the hidden name scope using tf.name_scope:
import tensorflow as tf
with tf.name_scope('hidden') as scope:
a = tf.constant(5, name='alpha')
W = tf.Variable(tf.random_uniform([1, 2], -1.0, 1.0), name='weights')
b = tf.Variable(tf.zeros([1]), name='biases')
This results in the following three op names:
hidden/alpha
hidden/weights
hidden/biases
By default, the visualization will collapse all three into a node labeled hidden. The extra detail isn't lost. You can double-click,
or click on the orange + sign in the top right to expand the node, and then you'll see three subnodes for alpha, weights and
biases.
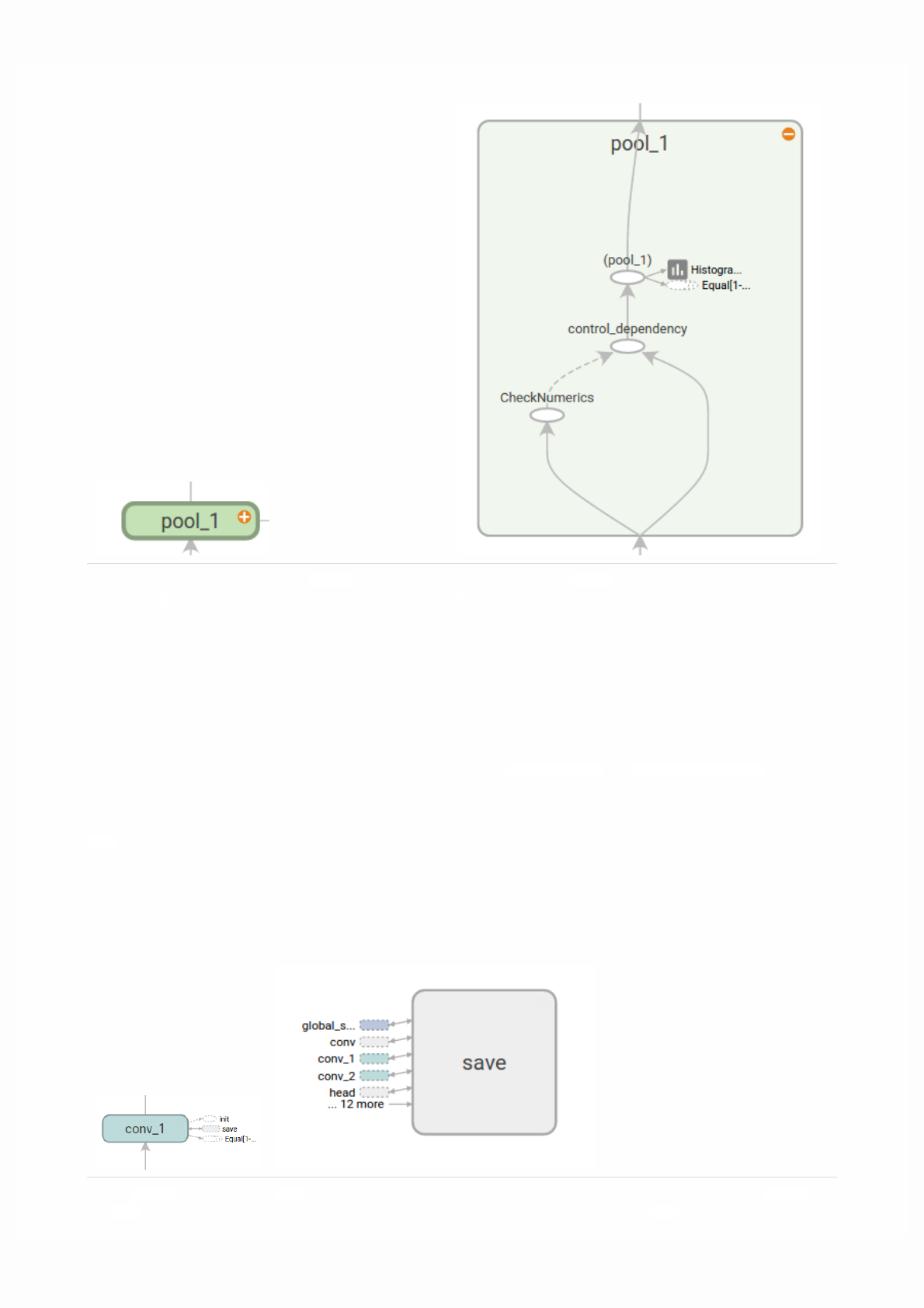
Initial view of top-level name scope pool_1. Clicking on
the orange + button on the top right or double-clicking on
the node itself will expand it.
Expanded view of pool_1 name scope. Clicking on the orange
- button on the top right or double-clicking on the node itself
will collapse the name scope.
Node conv_1 is connected
to save. Note the little
save has a high degree, and will appear as an auxiliary node. The connection with conv_1 is
shown as a node icon on its left. To further reduce clutter, since save has a lot of connections,
Here's a real-life example of a more complicated node in its initial and expanded states.
Grouping nodes by name scopes is critical to making a legible graph. If you're building a model, name scopes give you control
over the resulting visualization. The better your name scopes, the better your visualization.
The figure above illustrates a second aspect of the visualization. TensorFlow graphs have two kinds of connections: data
dependencies and control dependencies. Data dependencies show the flow of tensors between two ops and are shown as solid
arrows, while control dependencies use dotted lines. In the expanded view (right side of the figure above) all the connections
are data dependencies with the exception of the dotted line connecting CheckNumerics and control_dependency.
There's a second trick to simplifying the layout. Most TensorFlow graphs have a few nodes with many connections to other
nodes. For example, many nodes might have a control dependency on an initialization step. Drawing all edges between the
init node and its dependencies would create a very cluttered view.
To reduce clutter, the visualization separates out all high-degree nodes to an auxiliary area on the right and doesn't draw lines
to represent their edges. Instead of lines, we draw small node icons to indicate the connections. Separating out the auxiliary
nodes typically doesn't remove critical information since these nodes are usually related to bookkeeping functions. See
Interaction for how to move nodes between the main graph and the auxiliary area.
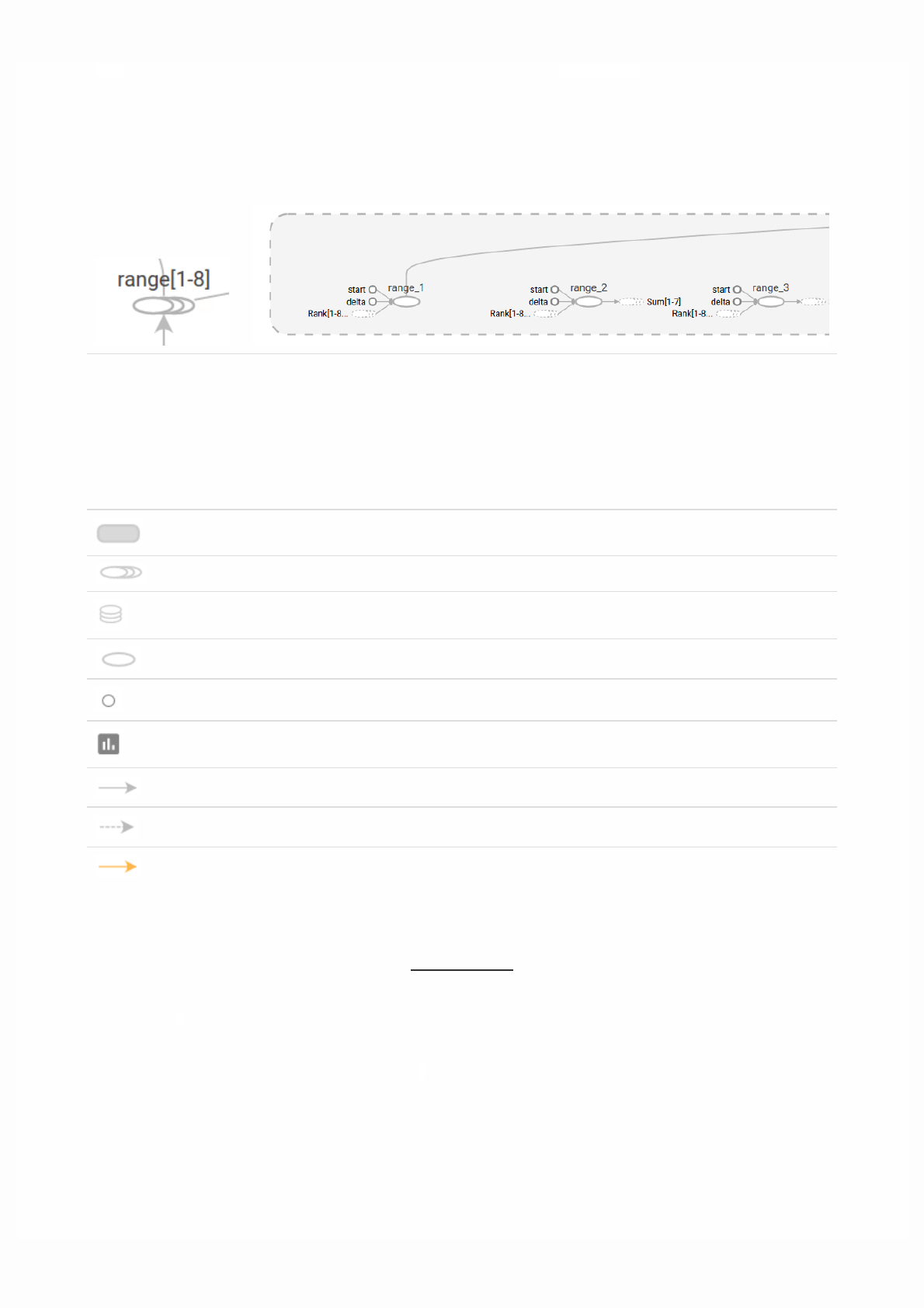
savenode icon on its right. we show the first 5 and abbreviate the others as ... 12 more.
A collapsed view of a
node sequence.
A small piece of the expanded view, after double-click.
Symbol Meaning
High-level node representing a name scope. Double-click to expand a high-level node.
Sequence of numbered nodes that are not connected to each other.
Sequence of numbered nodes that are connected to each other.
An individual operation node.
A constant.
A summary node.
Edge showing the data flow between operations.
Edge showing the control dependency between operations.
A reference edge showing that the outgoing operation node can mutate the incoming tensor.
One last structural simplification is series collapsing. Sequential motifs--that is, nodes whose names differ by a number at the
end and have isomorphic structures--are collapsed into a single stack of nodes, as shown below. For networks with long
sequences, this greatly simplifies the view. As with hierarchical nodes, double-clicking expands the series. See Interaction for
how to disable/enable series collapsing for a specific set of nodes.
Finally, as one last aid to legibility, the visualization uses special icons for constants and summary nodes. To summarize, here's
a table of node symbols:
Interaction
Navigate the graph by panning and zooming. Click and drag to pan, and use a scroll gesture to zoom. Double-click on a node,
or click on its + button, to expand a name scope that represents a group of operations. To easily keep track of the current
viewpoint when zooming and panning, there is a minimap in the bottom right corner.
To close an open node, double-click it again or click its - button. You can also click once to select a node. It will turn a darker
color, and details about it and the nodes it connects to will appear in the info card at upper right corner of the visualization.

Info card showing detailed information for the conv2 name
scope. The inputs and outputs are combined from the inputs and
outputs of the operation nodes inside the name scope. For name
scopes no attributes are shown.
Info card showing detailed information for the
DecodeRawoperation node. In addition to inputs and
outputs, the card shows the device and the attributes
associated with the current operation.
TensorBoard provides several ways to change the visual layout of the graph. This doesn't change the graph's computational
semantics, but it can bring some clarity to the network's structure. By right clicking on a node or pressing buttons on the
bottom of that node's info card, you can make the following changes to its layout:
Nodes can be moved between the main graph and the auxiliary area.
A series of nodes can be ungrouped so that the nodes in the series do not appear grouped together. Ungrouped series can
likewise be regrouped.
Selection can also be helpful in understanding high-degree nodes. Select any high-degree node, and the corresponding node
icons for its other connections will be selected as well. This makes it easy, for example, to see which nodes are being saved--and
which aren't.
Clicking on a node name in the info card will select it. If necessary, the viewpoint will automatically pan so that the node is
visible.
Finally, you can choose two color schemes for your graph, using the color menu above the legend. The default Structure View
shows structure: when two high-level nodes have the same structure, they appear in the same color of the rainbow. Uniquely
structured nodes are gray. There's a second view, which shows what device the different operations run on. Name scopes are
colored proportionally to the fraction of devices for the operations inside them.
The images below give an illustration for a piece of a real-life graph.
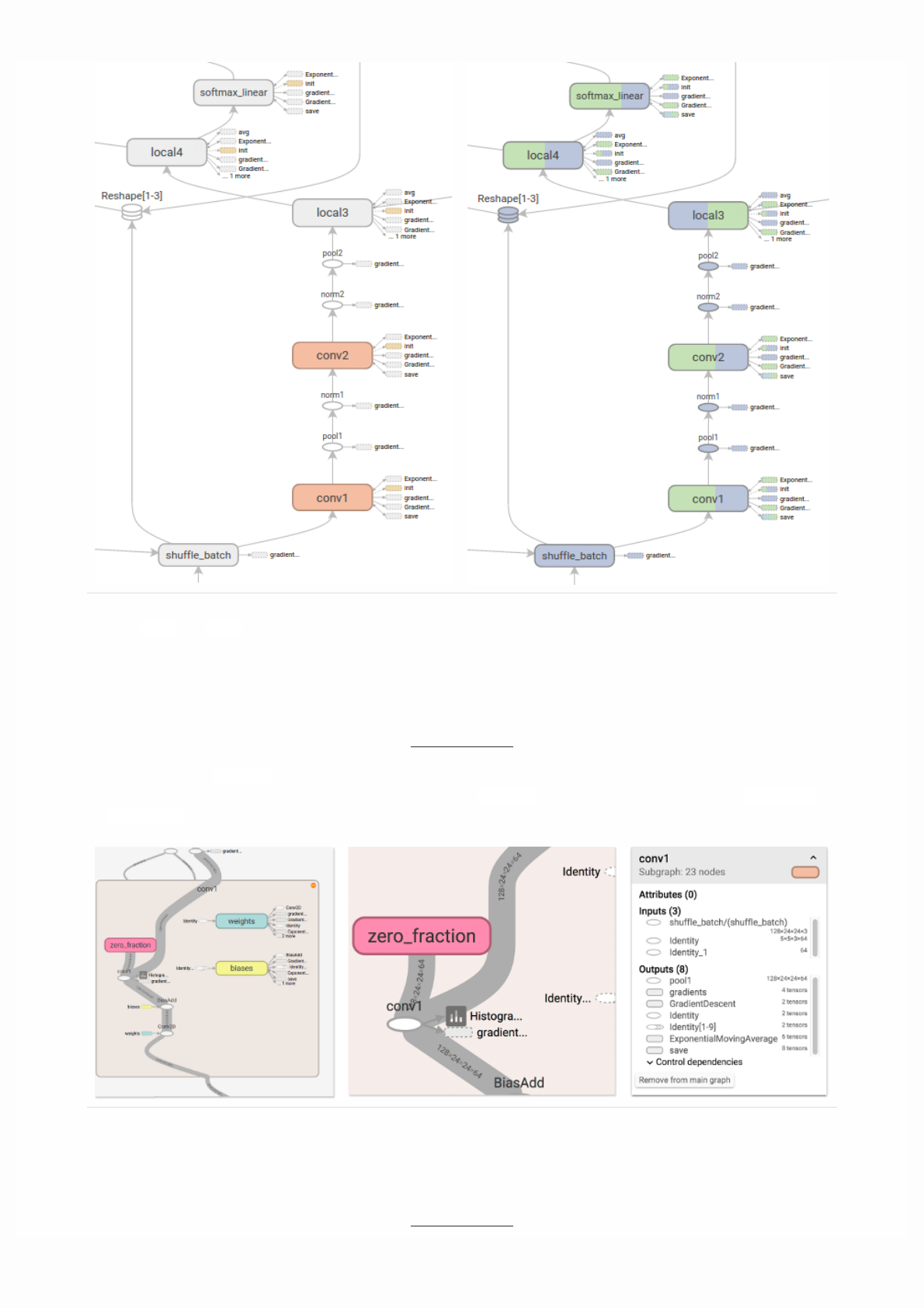
Structure view: The gray nodes have unique structure. The
orange conv1 and conv2 nodes have the same structure,
and analogously for nodes with other colors.
Device view: Name scopes are colored proportionally to the
fraction of devices of the operation nodes inside them. Here,
purple means GPU and the green is CPU.
CIFAR-10 model with tensor shape information.
Tensor shape information
When the serialized GraphDef includes tensor shapes, the graph visualizer labels edges with tensor dimensions, and edge
thickness reflects total tensor size. To include tensor shapes in the GraphDef pass the actual graph object (as in sess.graph) to
the FileWriter when serializing the graph. The images below show the CIFAR-10 model with tensor shape information:
Runtime statistics

Often it is useful to collect runtime metadata for a run, such as total memory usage, total compute time, and tensor shapes for
nodes. The code example below is a snippet from the train and test section of a modification of the simple MNIST tutorial, in
which we have recorded summaries and runtime statistics. See the Summaries Tutorial for details on how to record summaries.
Full source is here.
# Train the model, and also write summaries.
# Every 10th step, measure test-set accuracy, and write test summaries
# All other steps, run train_step on training data, & add training summaries
def feed_dict(train):
"""Make a TensorFlow feed_dict: maps data onto Tensor placeholders."""
if train or FLAGS.fake_data:
xs, ys = mnist.train.next_batch(100, fake_data=FLAGS.fake_data)
k = FLAGS.dropout
else:
xs, ys = mnist.test.images, mnist.test.labels
k = 1.0
return {x: xs, y_: ys, keep_prob: k}
for i in range(FLAGS.max_steps):
if i % 10 == 0: # Record summaries and test-set accuracy
summary, acc = sess.run([merged, accuracy], feed_dict=feed_dict(False))
test_writer.add_summary(summary, i)
print('Accuracy at step %s: %s' % (i, acc))
else: # Record train set summaries, and train
if i % 100 == 99: # Record execution stats
run_options = tf.RunOptions(trace_level=tf.RunOptions.FULL_TRACE)
run_metadata = tf.RunMetadata()
summary, _ = sess.run([merged, train_step],
feed_dict=feed_dict(True),
options=run_options,
run_metadata=run_metadata)
train_writer.add_run_metadata(run_metadata, 'step%d' % i)
train_writer.add_summary(summary, i)
print('Adding run metadata for', i)
else: # Record a summary
summary, _ = sess.run([merged, train_step], feed_dict=feed_dict(True))
train_writer.add_summary(summary, i)
This code will emit runtime statistics for every 100th step starting at step99.
When you launch tensorboard and go to the Graph tab, you will now see options under "Session runs" which correspond to the
steps where run metadata was added. Selecting one of these runs will show you the snapshot of the network at that step,
fading out unused nodes. In the controls on the left hand side, you will be able to color the nodes by total memory or total
compute time. Additionally, clicking on a node will display the exact total memory, compute time, and tensor output sizes.